bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_5217_orf2
Length=110
Score E
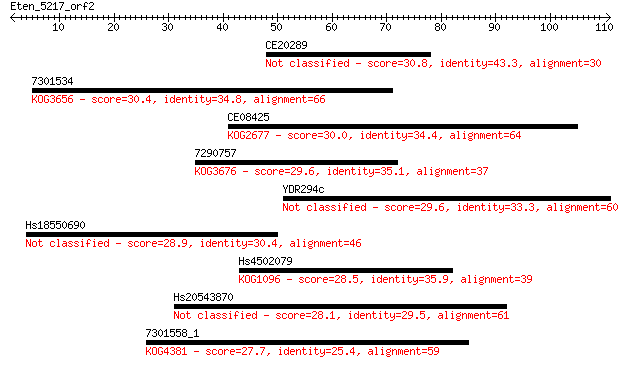
Sequences producing significant alignments: (Bits) Value
CE20289 30.8 0.66
7301534 30.4 0.75
CE08425 30.0 1.1
7290757 29.6 1.3
YDR294c 29.6 1.4
Hs18550690 28.9 2.6
Hs4502079 28.5 3.3
Hs20543870 28.1 3.7
7301558_1 27.7 5.5
> CE20289
Length=354
Score = 30.8 bits (68), Expect = 0.66, Method: Composition-based stats.
Identities = 13/30 (43%), Positives = 16/30 (53%), Gaps = 0/30 (0%)
Query 48 QQQRSSSARRPRLPVAPLSPGATRLRPPPP 77
Q++ RRPR AP SP + RP PP
Sbjct 135 QERVHQPERRPRRSAAPASPAKPQYRPQPP 164
> 7301534
Length=452
Score = 30.4 bits (67), Expect = 0.75, Method: Composition-based stats.
Identities = 23/70 (32%), Positives = 29/70 (41%), Gaps = 4/70 (5%)
Query 5 RSGNSKCTDRTEGLNCFCCLRGVGFGLGGVVA-GQLQDQQQQQQQQQRSSSARRPR---L 60
RSG + R GL +CCLR VG + G + S+ R+PR L
Sbjct 379 RSGFVQLMHRMPGLRRWCCLRSVGDRMNATSGTGPALPLNRMNTSTTYISARRKPRATSL 438
Query 61 PVAPLSPGAT 70
PLS G T
Sbjct 439 RANPLSCGET 448
> CE08425
Length=1657
Score = 30.0 bits (66), Expect = 1.1, Method: Composition-based stats.
Identities = 22/65 (33%), Positives = 30/65 (46%), Gaps = 10/65 (15%)
Query 41 DQQQQQQQQQRSSSARRPRLPVAPLSPGATRLRPPPPHSHPCSRVEGCWTAAANTGD-PG 99
+ Q Q Q + SA PR PVAP+ P + PP P W ++ +TGD P
Sbjct 613 ENSQYQVDQHPAESAAPPR-PVAPMEP---LFKQAPPQPDPFG-----WDSSGHTGDAPT 663
Query 100 VSKGV 104
S+ V
Sbjct 664 ASEPV 668
> 7290757
Length=1123
Score = 29.6 bits (65), Expect = 1.3, Method: Composition-based stats.
Identities = 13/37 (35%), Positives = 21/37 (56%), Gaps = 0/37 (0%)
Query 35 VAGQLQDQQQQQQQQQRSSSARRPRLPVAPLSPGATR 71
V+ + D ++ Q+ RS+ RR + P PL P +TR
Sbjct 1075 VSPEQSDDPDERSQRGRSAYTRRTQSPPDPLEPWSTR 1111
> YDR294c
Length=589
Score = 29.6 bits (65), Expect = 1.4, Method: Composition-based stats.
Identities = 20/64 (31%), Positives = 26/64 (40%), Gaps = 4/64 (6%)
Query 51 RSSSARRPRLPVAPLSPGATRLRPPPPHSHPCSRVEGCWTAAANTGDPGVSKG----VGA 106
R+S R + V P G P S P + V GCW N G+ G + VGA
Sbjct 394 RNSDLRMHQYYVNPAWTGGLYGSPTLAGSRPGAIVVGCWATMVNMGENGYIESCQEIVGA 453
Query 107 FLSF 110
+ F
Sbjct 454 AMKF 457
> Hs18550690
Length=630
Score = 28.9 bits (63), Expect = 2.6, Method: Compositional matrix adjust.
Identities = 14/46 (30%), Positives = 23/46 (50%), Gaps = 0/46 (0%)
Query 4 GRSGNSKCTDRTEGLNCFCCLRGVGFGLGGVVAGQLQDQQQQQQQQ 49
RS + +C C +G+G+GL V+ L+ QQ Q+ Q+
Sbjct 576 ARSQKENLNYKHLEDHCSQCSQGMGYGLPRVLGSALEKQQAQEPQK 621
> Hs4502079
Length=776
Score = 28.5 bits (62), Expect = 3.3, Method: Composition-based stats.
Identities = 14/40 (35%), Positives = 19/40 (47%), Gaps = 1/40 (2%)
Query 43 QQQQQQQQRSSSARRPRL-PVAPLSPGATRLRPPPPHSHP 81
Q QQ + A P + P P+ GAT L P P++ P
Sbjct 98 QMPPQQDWKGPPAASPAMSPTTPVVTGATSLPTPAPYAMP 137
> Hs20543870
Length=612
Score = 28.1 bits (61), Expect = 3.7, Method: Compositional matrix adjust.
Identities = 18/65 (27%), Positives = 29/65 (44%), Gaps = 7/65 (10%)
Query 31 LGGVVAGQLQDQQQQQQQQQRSSSARRPRLPVAP----LSPGATRLRPPPPHSHPCSRVE 86
L G+ G+ QD + + +Q + +LPV L R ++HPC ++E
Sbjct 507 LSGMREGRKQDFWEGKAKQAFPQKGKESKLPVMEKDKRLQNNTLR---SSAYNHPCCKLE 563
Query 87 GCWTA 91
CW A
Sbjct 564 SCWKA 568
> 7301558_1
Length=951
Score = 27.7 bits (60), Expect = 5.5, Method: Composition-based stats.
Identities = 15/59 (25%), Positives = 26/59 (44%), Gaps = 0/59 (0%)
Query 26 GVGFGLGGVVAGQLQDQQQQQQQQQRSSSARRPRLPVAPLSPGATRLRPPPPHSHPCSR 84
G+ + + ++QDQ+++Q+Q S+ +P A SPG H C R
Sbjct 816 GIQLSVSKLKVSEMQDQERRQRQLLSGSAQSLQAMPEAVGSPGIWAPDSIATHCTACER 874
Lambda K H
0.318 0.136 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1195973986
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40