bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: eggV2
2,483,276 sequences; 915,453,621 total letters
Query= Eten_1902_orf2
Length=130
Score E
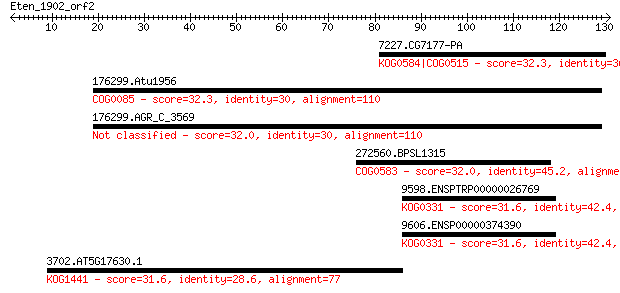
Sequences producing significant alignments: (Bits) Value
7227.CG7177-PA 32.3 3.8
176299.Atu1956 32.3 4.6
176299.AGR_C_3569 32.0 4.9
272560.BPSL1315 32.0 5.7
9598.ENSPTRP00000026769 31.6 6.6
9606.ENSP00000374390 31.6 6.9
3702.AT5G17630.1 31.6 8.0
> 7227.CG7177-PA
Length=2308
Score = 32.3 bits (72), Expect = 3.8, Method: Composition-based stats.
Identities = 18/49 (36%), Positives = 23/49 (46%), Gaps = 3/49 (6%)
Query 81 AAPLASAAAAAAGAPSTAGDLAAERPKNAAGPSAPRAGCNVQEGEVISA 129
A+P S+ A +G P T +P N AGPS P G N + SA
Sbjct 1675 ASPAMSSLKATSGQPQTQ---EITKPNNGAGPSVPSVGQNTPTAALTSA 1720
> 176299.Atu1956
Length=1378
Score = 32.3 bits (72), Expect = 4.6, Method: Composition-based stats.
Identities = 33/135 (24%), Positives = 58/135 (42%), Gaps = 28/135 (20%)
Query 19 RGSLFLLLLDAPF-------------ASELSSSEEGALDEVSATAYAMVLTSKETKTGKR 65
R ++ LL +APF + +++ G +D+V AT ++ +++ GK
Sbjct 697 RQAVPLLRAEAPFVGTGMEPIVARDSGAAIAARRGGVVDQVDATRI-VIRATEDLDAGKS 755
Query 66 SVKLYRRK------------LRPLLAVAAPLASAAAAAAGAPSTAGDLAAERPKNAAGPS 113
V +YR + RPL++V ++ A G + GDLA R NA
Sbjct 756 GVDIYRLQKFQRSNQNTCVNQRPLVSVGDAISKGDIIADGPSTDLGDLALGR--NALVAF 813
Query 114 APRAGCNVQEGEVIS 128
P G N ++ ++S
Sbjct 814 MPWNGYNYEDSILMS 828
> 176299.AGR_C_3569
Length=1411
Score = 32.0 bits (71), Expect = 4.9, Method: Composition-based stats.
Identities = 33/135 (24%), Positives = 58/135 (42%), Gaps = 28/135 (20%)
Query 19 RGSLFLLLLDAPF-------------ASELSSSEEGALDEVSATAYAMVLTSKETKTGKR 65
R ++ LL +APF + +++ G +D+V AT ++ +++ GK
Sbjct 730 RQAVPLLRAEAPFVGTGMEPIVARDSGAAIAARRGGVVDQVDATRI-VIRATEDLDAGKS 788
Query 66 SVKLYRRK------------LRPLLAVAAPLASAAAAAAGAPSTAGDLAAERPKNAAGPS 113
V +YR + RPL++V ++ A G + GDLA R NA
Sbjct 789 GVDIYRLQKFQRSNQNTCVNQRPLVSVGDAISKGDIIADGPSTDLGDLALGR--NALVAF 846
Query 114 APRAGCNVQEGEVIS 128
P G N ++ ++S
Sbjct 847 MPWNGYNYEDSILMS 861
> 272560.BPSL1315
Length=299
Score = 32.0 bits (71), Expect = 5.7, Method: Compositional matrix adjust.
Identities = 19/42 (45%), Positives = 21/42 (50%), Gaps = 3/42 (7%)
Query 76 PLLAVAAPLASAAAAAAGAPSTAGDLAAERPKNAAGPSAPRA 117
PL+ VA+P A A G P T DL RP N A PS R
Sbjct 162 PLINVASP---AYLARHGVPRTPADLERHRPVNYASPSNGRV 200
> 9598.ENSPTRP00000026769
Length=730
Score = 31.6 bits (70), Expect = 6.6, Method: Composition-based stats.
Identities = 14/33 (42%), Positives = 20/33 (60%), Gaps = 0/33 (0%)
Query 86 SAAAAAAGAPSTAGDLAAERPKNAAGPSAPRAG 118
S A+ G P TAG + + P++A GPS R+G
Sbjct 233 SRGASIPGTPPTAGRVVSLSPEDAPGPSLRRSG 265
> 9606.ENSP00000374390
Length=730
Score = 31.6 bits (70), Expect = 6.9, Method: Composition-based stats.
Identities = 14/33 (42%), Positives = 20/33 (60%), Gaps = 0/33 (0%)
Query 86 SAAAAAAGAPSTAGDLAAERPKNAAGPSAPRAG 118
S A+ G P TAG + + P++A GPS R+G
Sbjct 233 SRGASIPGTPPTAGRVVSLSPEDAPGPSLRRSG 265
> 3702.AT5G17630.1
Length=417
Score = 31.6 bits (70), Expect = 8.0, Method: Composition-based stats.
Identities = 22/81 (27%), Positives = 39/81 (48%), Gaps = 4/81 (4%)
Query 9 LPGWGGCFGARGS----LFLLLLDAPFASELSSSEEGALDEVSATAYAMVLTSKETKTGK 64
+PG+ + G+ F +LL F + S ALDE+S +++ T K
Sbjct 312 VPGYHKAIASVGTPSTFYFWVLLSGVFYHLYNQSSYQALDEISPLTFSVGNTMKRVVVII 371
Query 65 RSVKLYRRKLRPLLAVAAPLA 85
+V ++R +RPL A+ + +A
Sbjct 372 STVLVFRNPVRPLNALGSAIA 392
Lambda K H
0.311 0.127 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 23020010250
Database: eggV2
Posted date: Dec 15, 2009 4:47 PM
Number of letters in database: 915,453,621
Number of sequences in database: 2,483,276
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40