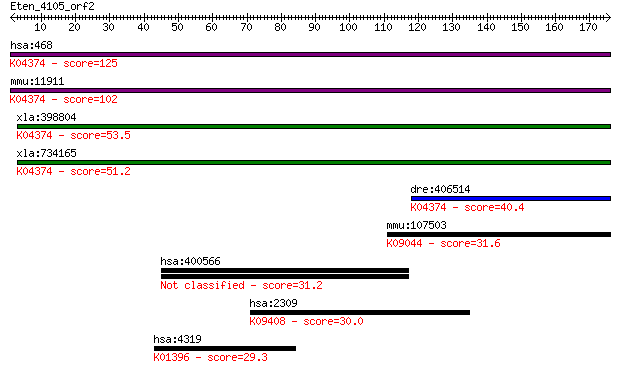
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
164,496 sequences; 82,071,388 total letters
Query= Eten_4105_orf2
Length=175
Score E
Sequences producing significant alignments: (Bits) Value
hsa:468 ATF4, CREB-2, CREB2, TAXREB67, TXREB; activating trans... 125 9e-29
mmu:11911 Atf4, Atf-4, C/ATF, CREB2, MGC96460, TAXREB67; activ... 102 5e-22
xla:398804 atf4-a, MGC68539, atf-4, atf4, creb-2, creb2, taxre... 53.5 3e-07
xla:734165 atf4-b, MGC196486, MGC196490, atf-4, atf4-ii, creb-... 51.2 2e-06
dre:406514 atf4b1, atf4, wu:fj01c08, wu:fk30b04, zgc:85951; ac... 40.4 0.003
mmu:107503 Atf5, AFTA, Atf7, Atfx, MGC102397, ODA-10; activati... 31.6 1.4
hsa:400566 C17orf97; chromosome 17 open reading frame 97 31.2
hsa:2309 FOXO3, AF6q21, DKFZp781A0677, FKHRL1, FKHRL1P2, FOXO2... 30.0 4.2
hsa:4319 MMP10, SL-2, STMY2; matrix metallopeptidase 10 (strom... 29.3 7.5
> hsa:468 ATF4, CREB-2, CREB2, TAXREB67, TXREB; activating transcription
factor 4 (tax-responsive enhancer element B67); K04374
activating transcription factor 4
Length=351
Score = 125 bits (313), Expect = 9e-29, Method: Compositional matrix adjust.
Identities = 75/177 (42%), Positives = 94/177 (53%), Gaps = 3/177 (1%)
Query 1 FFKLFPLSPGTLFSPPDLSFILELSREGVFPEGDSNPD--SPLLGFPQGIKGGGPPPKNN 58
F + PLSPG L S PD SF LEL E EGD PD + + PQ IK P N+
Sbjct 159 FLQPLPLSPGVLSSTPDHSFSLELGSEVDITEGDRKPDYTAYVAMIPQCIKEEDTPSDND 218
Query 59 RGFFLTPDSFWGFPRDSPSPPGGFPNKTLFFPGALRGFSPPKPLALFGGKMFPPKFRGGK 118
G ++P+S+ G P+ SPS G PN++L PG L G + PKP G KM K +G K
Sbjct 219 SGICMSPESYLGSPQHSPSTRGS-PNRSLPSPGVLCGSARPKPYDPPGEKMVAAKVKGEK 277
Query 119 LPKNLKKMGQTKPPVFCFPQKKGGEQGAPPGEFKGLEKKKWGFKRKGNFFAQGVPFF 175
L K LKKM Q K + QKK EQ A GE K LEKK K + + A+ + +
Sbjct 278 LDKKLKKMEQNKTAATRYRQKKRAEQEALTGECKELEKKNEALKERADSLAKEIQYL 334
> mmu:11911 Atf4, Atf-4, C/ATF, CREB2, MGC96460, TAXREB67; activating
transcription factor 4; K04374 activating transcription
factor 4
Length=349
Score = 102 bits (255), Expect = 5e-22, Method: Compositional matrix adjust.
Identities = 70/183 (38%), Positives = 88/183 (48%), Gaps = 16/183 (8%)
Query 1 FFKLFPLSPGTLFSPPDLSFILELSREGVFPEGDSNPDSP--LLGFPQGIKGGGPPPKNN 58
F + FP SPG L S P+ SF LEL E EGD PDS + P +K P N+
Sbjct 158 FLQPFPCSPGVLSSTPEHSFSLELGSEVDISEGDRKPDSAAYITLIPPCVKEEDTPSDND 217
Query 59 RGFFLTPDSFWGFPRDSPS----PPGGFPNKTLFFPGALRGFSPPKPLALFG--GKMFPP 112
G ++P+S+ G P+ SPS PP P+ PG RG PKP + G
Sbjct 218 SGICMSPESYLGSPQHSPSTSRAPPDNLPS-----PGGSRGSPRPKP---YDPPGVSLTA 269
Query 113 KFRGGKLPKNLKKMGQTKPPVFCFPQKKGGEQGAPPGEFKGLEKKKWGFKRKGNFFAQGV 172
K + KL K LKKM Q K + QKK EQ A GE K LEKK K K + A+ +
Sbjct 270 KVKTEKLDKKLKKMEQNKTAATRYRQKKRAEQEALTGECKELEKKNEALKEKADSLAKEI 329
Query 173 PFF 175
+
Sbjct 330 QYL 332
> xla:398804 atf4-a, MGC68539, atf-4, atf4, creb-2, creb2, taxreb67,
txreb; activating transcription factor 4 (tax-responsive
enhancer element B67); K04374 activating transcription factor
4
Length=336
Score = 53.5 bits (127), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 55/183 (30%), Positives = 76/183 (41%), Gaps = 16/183 (8%)
Query 3 KLFPLSPG---TLFSPPDLSFILELSREGVFPEGDSNPDSPLLGFPQGIKGGGPPPKNNR 59
++ P+SP +L S PD SFIL+L E E D S N+
Sbjct 142 QVAPVSPDLSESLSSSPDHSFILDLGSEVDVAE-DEKKSSATYTLKMLKCEKEDSSDNDS 200
Query 60 GFFLTPDSFWGFPRDSPSPPGGF--PNKTLFFPGALRGFSPPKPLAL---FGGKMFPPKF 114
G ++P S+ P+ SPS + P +L P + R PKP L M K
Sbjct 201 GISMSP-SYLESPQQSPSVAECYSSPQSSLASPVSER----PKPYDLPCKDRDVMAKVKV 255
Query 115 RGG--KLPKNLKKMGQTKPPVFCFPQKKGGEQGAPPGEFKGLEKKKWGFKRKGNFFAQGV 172
GG ++ K KKM Q K + QKK EQ A GE + LE K K K + + +
Sbjct 256 AGGQPRVDKKKKKMEQNKTAATRYRQKKRAEQEAISGECRELEGKNDSLKEKVDSLTKEI 315
Query 173 PFF 175
+
Sbjct 316 QYL 318
> xla:734165 atf4-b, MGC196486, MGC196490, atf-4, atf4-ii, creb-2,
creb2, taxreb67, txreb; activating transcription factor
4 (tax-responsive enhancer element B67); K04374 activating
transcription factor 4
Length=342
Score = 51.2 bits (121), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 54/183 (29%), Positives = 79/183 (43%), Gaps = 16/183 (8%)
Query 3 KLFPLSPG---TLFSPPDLSFILELSREGVFPEGDSNPDSPLLGFPQGIKGGGPPPKNNR 59
++ P+SP +L PD SFIL++ E V GD SP F N+
Sbjct 148 QVAPVSPDLTESLSLGPDHSFILDVGSE-VDVSGDEKKSSPTYTFEMLKCEKEDSSDNDS 206
Query 60 GFFLTPDSFWGFPRDSPSPPGGF--PNKTLFFPGALRGFSPPKPLAL---FGGKMFPPKF 114
G ++P S+ G P+ SPS + P +L P + R PKP L + K
Sbjct 207 GISMSP-SYMGSPQQSPSIVECYSSPQSSLESPVSER----PKPYDLPCKDRDVIAKVKV 261
Query 115 RGG--KLPKNLKKMGQTKPPVFCFPQKKGGEQGAPPGEFKGLEKKKWGFKRKGNFFAQGV 172
GG ++ K KKM Q K + QKK EQ GE + LE+K + K + + +
Sbjct 262 AGGQPRVDKKKKKMEQNKTAATRYRQKKRVEQELISGECRELEEKNDSLREKVDSLTKEI 321
Query 173 PFF 175
+
Sbjct 322 QYL 324
> dre:406514 atf4b1, atf4, wu:fj01c08, wu:fk30b04, zgc:85951;
activating transcription factor 4b1 (tax-responsive enhancer
element B67); K04374 activating transcription factor 4
Length=339
Score = 40.4 bits (93), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 19/58 (32%), Positives = 27/58 (46%), Gaps = 0/58 (0%)
Query 118 KLPKNLKKMGQTKPPVFCFPQKKGGEQGAPPGEFKGLEKKKWGFKRKGNFFAQGVPFF 175
K+ K LKKM Q K + QKK EQ + E LEKK K + ++ + +
Sbjct 265 KVEKKLKKMEQNKTAATRYRQKKRVEQESLNSECSELEKKNRELSEKADSLSREIQYL 322
> mmu:107503 Atf5, AFTA, Atf7, Atfx, MGC102397, ODA-10; activating
transcription factor 5; K09044 activating transcription
factor 5
Length=283
Score = 31.6 bits (70), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 16/65 (24%), Positives = 26/65 (40%), Gaps = 3/65 (4%)
Query 111 PPKFRGGKLPKNLKKMGQTKPPVFCFPQKKGGEQGAPPGEFKGLEKKKWGFKRKGNFFAQ 170
P RG + KK Q K + Q+K E A GE +GLE + + + +
Sbjct 204 PASTRGDR---KQKKRDQNKSAALRYRQRKRAEGEALEGECQGLEARNRELRERAESVER 260
Query 171 GVPFF 175
+ +
Sbjct 261 EIQYV 265
> hsa:400566 C17orf97; chromosome 17 open reading frame 97
Length=423
Score = 31.2 bits (69), Expect = 1.8, Method: Compositional matrix adjust.
Identities = 25/72 (34%), Positives = 35/72 (48%), Gaps = 6/72 (8%)
Query 45 PQGIKGGGPPPKNNRGFFLTPDSFWGFPRDSPSPPGGFPNKTLFFPGALRGFSPPKPLAL 104
P+ +KG P P+ +GF P++ GF D + G P+ P AL+GF P P AL
Sbjct 259 PEALKGFHPDPEALKGFHTDPEALKGFHIDPEALKGFHPD-----PKALKGFH-PDPKAL 312
Query 105 FGGKMFPPKFRG 116
G P +G
Sbjct 313 KGFHTDPEALKG 324
Score = 29.6 bits (65), Expect = 5.4, Method: Compositional matrix adjust.
Identities = 26/72 (36%), Positives = 35/72 (48%), Gaps = 6/72 (8%)
Query 45 PQGIKGGGPPPKNNRGFFLTPDSFWGFPRDSPSPPGGFPNKTLFFPGALRGFSPPKPLAL 104
P+ +KG P PK +GF P++ GF D + G P+ P AL+GF P P AL
Sbjct 299 PKALKGFHPDPKALKGFHTDPEALKGFHPDPKALKGFHPD-----PEALKGFH-PDPEAL 352
Query 105 FGGKMFPPKFRG 116
G P +G
Sbjct 353 KGFHPDPEALKG 364
> hsa:2309 FOXO3, AF6q21, DKFZp781A0677, FKHRL1, FKHRL1P2, FOXO2,
FOXO3A, MGC12739, MGC31925; forkhead box O3; K09408 forkhead
box protein O3
Length=673
Score = 30.0 bits (66), Expect = 4.2, Method: Composition-based stats.
Identities = 16/64 (25%), Positives = 25/64 (39%), Gaps = 0/64 (0%)
Query 71 FPRDSPSPPGGFPNKTLFFPGALRGFSPPKPLALFGGKMFPPKFRGGKLPKNLKKMGQTK 130
P PSP GG ++ FP +G P + F +F P ++ + + K
Sbjct 396 LPPSQPSPTGGLMQRSSSFPYTTKGSGLGSPTSSFNSTVFGPSSLNSLRQSPMQTIQENK 455
Query 131 PPVF 134
P F
Sbjct 456 PATF 459
> hsa:4319 MMP10, SL-2, STMY2; matrix metallopeptidase 10 (stromelysin
2) (EC:3.4.24.22); K01396 matrix metalloproteinase-10
(stromelysin 2) [EC:3.4.24.22]
Length=476
Score = 29.3 bits (64), Expect = 7.5, Method: Compositional matrix adjust.
Identities = 16/53 (30%), Positives = 23/53 (43%), Gaps = 12/53 (22%)
Query 43 GFPQGIKGGGPPP------------KNNRGFFLTPDSFWGFPRDSPSPPGGFP 83
G+P+GI G PP + + +F D +W F +S S GFP
Sbjct 370 GYPRGIHTLGFPPTIRKIDAAVSDKEKKKTYFFAADKYWRFDENSQSMEQGFP 422
Lambda K H
0.321 0.149 0.495
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 4535951560
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
Posted date: Sep 17, 2011 2:57 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40