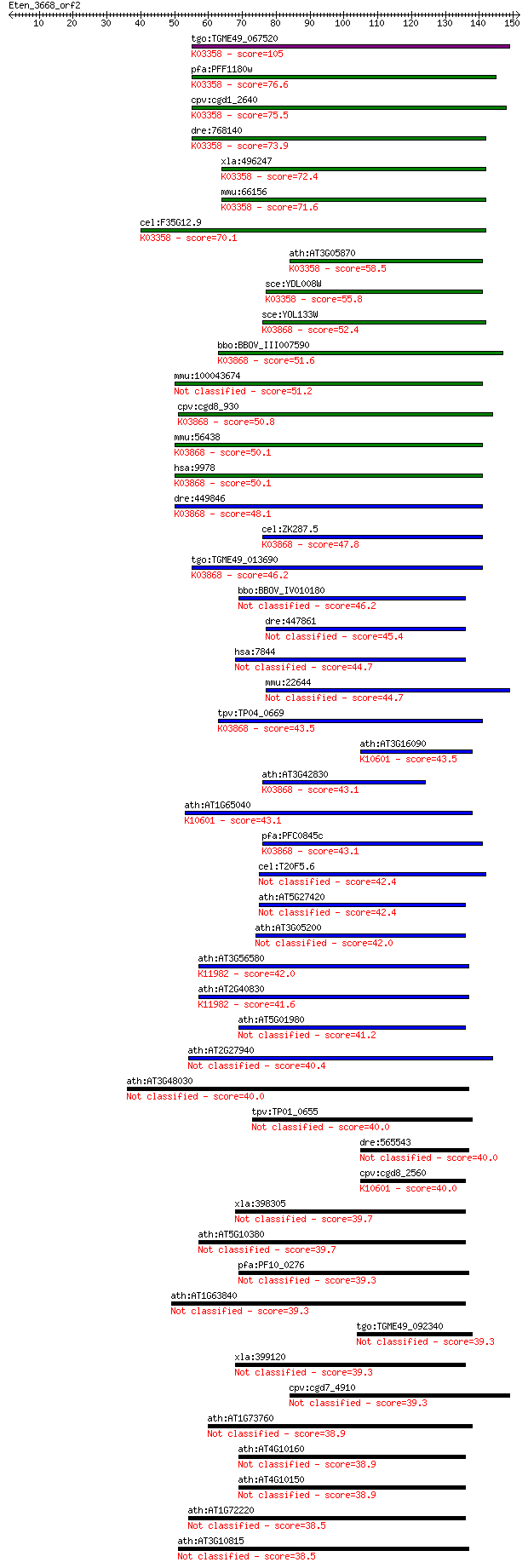
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
164,496 sequences; 82,071,388 total letters
Query= Eten_3668_orf2
Length=151
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_067520 hypothetical protein ; K03358 anaphase-promo... 105 5e-23
pfa:PFF1180w anaphase-promoting complex subunit, putative; K03... 76.6 3e-14
cpv:cgd1_2640 hypothetical protein ; K03358 anaphase-promoting... 75.5 6e-14
dre:768140 anapc11, si:dkey-237i9.4, zgc:153671; APC11 anaphas... 73.9 2e-13
xla:496247 anapc11; anaphase promoting complex subunit 11; K03... 72.4 5e-13
mmu:66156 Anapc11, 1110011I19Rik, R75218; anaphase promoting c... 71.6 1e-12
cel:F35G12.9 apc-11; Anaphase Promoting Complex; see also mat ... 70.1 3e-12
ath:AT3G05870 APC11; APC11; protein binding / ubiquitin-protei... 58.5 8e-09
sce:YDL008W APC11; Apc11p (EC:6.3.2.-); K03358 anaphase-promot... 55.8 5e-08
sce:YOL133W HRT1, HRT2, RBX1, ROC1; Hrt1p; K03868 RING-box pro... 52.4 6e-07
bbo:BBOV_III007590 17.m07664; hypothetical protein; K03868 RIN... 51.6 1e-06
mmu:100043674 Gm9840; predicted gene 9840 51.2
cpv:cgd8_930 RBX1 ortholog, RING finger ; K03868 RING-box prot... 50.8 2e-06
mmu:56438 Rbx1, 1500002P15Rik, AA517855, ROC1; ring-box 1; K03... 50.1 3e-06
hsa:9978 RBX1, BA554C12.1, FLJ60363, MGC13357, MGC1481, RNF75,... 50.1 3e-06
dre:449846 rbx1, im:7137515, zgc:136444; ring-box 1; K03868 RI... 48.1 1e-05
cel:ZK287.5 rbx-1; yeast RBX (ring finger protein) homolog fam... 47.8 1e-05
tgo:TGME49_013690 hypothetical protein ; K03868 RING-box prote... 46.2 4e-05
bbo:BBOV_IV010180 23.m06036; zinc finger, C3HC4 type domain co... 46.2 4e-05
dre:447861 rnf103, zgc:92077; ring finger protein 103 45.4
hsa:7844 RNF103, HKF-1, KF-1, KF1, MGC102815, MGC41857, ZFP-10... 44.7 1e-04
mmu:22644 Rnf103, AW146237, AW212918, Zfp103, kf-1; ring finge... 44.7 1e-04
tpv:TP04_0669 hypothetical protein; K03868 RING-box protein 1 43.5
ath:AT3G16090 zinc finger (C3HC4-type RING finger) family prot... 43.5 3e-04
ath:AT3G42830 ring-box protein Roc1/Rbx1/Hrt1, putative; K0386... 43.1 3e-04
ath:AT1G65040 protein binding / zinc ion binding; K10601 E3 ub... 43.1 3e-04
pfa:PFC0845c ubiquitin-protein ligase, putative (EC:6.3.2.19);... 43.1 4e-04
cel:T20F5.6 hypothetical protein 42.4 6e-04
ath:AT5G27420 zinc finger (C3HC4-type RING finger) family protein 42.4 6e-04
ath:AT3G05200 ATL6; ATL6; protein binding / zinc ion binding 42.0 7e-04
ath:AT3G56580 zinc finger (C3HC4-type RING finger) family prot... 42.0 8e-04
ath:AT2G40830 RHC1A; RHC1A; protein binding / zinc ion binding... 41.6 0.001
ath:AT5G01980 zinc finger (C3HC4-type RING finger) family protein 41.2 0.001
ath:AT2G27940 zinc finger (C3HC4-type RING finger) family protein 40.4 0.002
ath:AT3G48030 hypoxia-responsive family protein / zinc finger ... 40.0 0.003
tpv:TP01_0655 hypothetical protein 40.0 0.003
dre:565543 ring finger protein 6-like 40.0 0.003
cpv:cgd8_2560 HRD1 like membrane associated RING finger contai... 40.0 0.003
xla:398305 rnf103-a, kf1, rnf103, x-kf-1a, zfp103; ring finger... 39.7 0.004
ath:AT5G10380 RING1; RING1; protein binding / ubiquitin-protei... 39.7 0.004
pfa:PF10_0276 Zinc finger, C3HC4 type, putative 39.3 0.004
ath:AT1G63840 zinc finger (C3HC4-type RING finger) family protein 39.3 0.005
tgo:TGME49_092340 zinc finger (C3HC4 RING finger) protein, put... 39.3 0.005
xla:399120 rnf103-b, KF-1a, MGC99268, Rnf103, kf1, x-kf-1a, x-... 39.3 0.005
cpv:cgd7_4910 conserved 3 transmembrane domain membrane associ... 39.3 0.005
ath:AT1G73760 zinc finger (C3HC4-type RING finger) family protein 38.9 0.006
ath:AT4G10160 zinc finger (C3HC4-type RING finger) family protein 38.9 0.006
ath:AT4G10150 zinc finger (C3HC4-type RING finger) family protein 38.9 0.007
ath:AT1G72220 zinc finger (C3HC4-type RING finger) family protein 38.5 0.007
ath:AT3G10815 zinc finger (C3HC4-type RING finger) family protein 38.5 0.007
> tgo:TGME49_067520 hypothetical protein ; K03358 anaphase-promoting
complex subunit 11
Length=144
Score = 105 bits (263), Expect = 5e-23, Method: Compositional matrix adjust.
Identities = 49/94 (52%), Positives = 58/94 (61%), Gaps = 0/94 (0%)
Query 55 LEVLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCI 114
+ V VH VG RW VP D C IC N FE+ C+ACD PG+ CPPAFGACGH+FHLHCI
Sbjct 36 VRVRHVHFVGGWRWAGVPPDEKCGICSNAFEVHCSACDRPGEACPPAFGACGHVFHLHCI 95
Query 115 AAWLGGRGLRGAAQDTCPMCRQLWRFAAQQQHLS 148
WL +Q CP+CR+ WRF + S
Sbjct 96 GTWLSREDALRESQANCPLCRRPWRFKTANEASS 129
> pfa:PFF1180w anaphase-promoting complex subunit, putative; K03358
anaphase-promoting complex subunit 11
Length=89
Score = 76.6 bits (187), Expect = 3e-14, Method: Compositional matrix adjust.
Identities = 37/90 (41%), Positives = 46/90 (51%), Gaps = 6/90 (6%)
Query 55 LEVLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCI 114
+ V +H V +W D C IC++ E C C PG+ CPPAFG CGH FHLHC+
Sbjct 4 ITVKRIHAVARWKWIGSTIDSVCAICNSSLENTCTTCMRPGNGCPPAFGKCGHHFHLHCM 63
Query 115 AAWLGGRGLRGAAQDTCPMCRQLWRFAAQQ 144
W+ L TCP CR W + QQ
Sbjct 64 EKWIKQNKL------TCPCCRADWYYETQQ 87
> cpv:cgd1_2640 hypothetical protein ; K03358 anaphase-promoting
complex subunit 11
Length=117
Score = 75.5 bits (184), Expect = 6e-14, Method: Compositional matrix adjust.
Identities = 40/94 (42%), Positives = 50/94 (53%), Gaps = 7/94 (7%)
Query 55 LEVLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCI 114
+++ + +G W + D C IC FEL C C PGD CPPAFG CGH FHLHCI
Sbjct 25 IKINKLRMIGY--WNWLCGDDLCSICSESFELTCPQCFKPGDSCPPAFGECGHSFHLHCI 82
Query 115 AAWLG-GRGLRGAAQDTCPMCRQLWRFAAQQQHL 147
WL R G CPMCR+ ++F + L
Sbjct 83 HEWLSRARNDSGM----CPMCRREFKFQSNANIL 112
> dre:768140 anapc11, si:dkey-237i9.4, zgc:153671; APC11 anaphase
promoting complex subunit 11 homolog (yeast); K03358 anaphase-promoting
complex subunit 11
Length=88
Score = 73.9 bits (180), Expect = 2e-13, Method: Compositional matrix adjust.
Identities = 36/87 (41%), Positives = 45/87 (51%), Gaps = 4/87 (4%)
Query 55 LEVLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCI 114
++V GV W V D +C IC F C C PGD+CP +G C H FH+HCI
Sbjct 1 MKVKIKQWNGVASWLWVANDENCGICRASFNGCCPDCKVPGDDCPLVWGQCSHCFHMHCI 60
Query 115 AAWLGGRGLRGAAQDTCPMCRQLWRFA 141
WL + + Q CPMCRQ W+F
Sbjct 61 LKWLNSQQV----QQQCPMCRQEWKFK 83
> xla:496247 anapc11; anaphase promoting complex subunit 11; K03358
anaphase-promoting complex subunit 11
Length=84
Score = 72.4 bits (176), Expect = 5e-13, Method: Compositional matrix adjust.
Identities = 35/78 (44%), Positives = 43/78 (55%), Gaps = 4/78 (5%)
Query 64 GVCRWKEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGL 123
GV W+ V D +C IC F C C PGD+CP +G C H FH+HCI WL + +
Sbjct 10 GVASWQWVANDDNCGICRMAFNGCCPECKIPGDDCPLVWGHCSHCFHMHCILKWLNSQQV 69
Query 124 RGAAQDTCPMCRQLWRFA 141
Q CPMCRQ W+F
Sbjct 70 ----QQHCPMCRQEWKFK 83
> mmu:66156 Anapc11, 1110011I19Rik, R75218; anaphase promoting
complex subunit 11; K03358 anaphase-promoting complex subunit
11
Length=84
Score = 71.6 bits (174), Expect = 1e-12, Method: Compositional matrix adjust.
Identities = 35/78 (44%), Positives = 42/78 (53%), Gaps = 4/78 (5%)
Query 64 GVCRWKEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGL 123
GV W V D +C IC F C C PGD+CP +G C H FH+HCI WL + +
Sbjct 10 GVATWLWVANDENCGICRMAFNGCCPDCKVPGDDCPLVWGQCSHCFHMHCILKWLNAQQV 69
Query 124 RGAAQDTCPMCRQLWRFA 141
Q CPMCRQ W+F
Sbjct 70 ----QQHCPMCRQEWKFK 83
> cel:F35G12.9 apc-11; Anaphase Promoting Complex; see also mat
family member (apc-11); K03358 anaphase-promoting complex
subunit 11
Length=135
Score = 70.1 bits (170), Expect = 3e-12, Method: Compositional matrix adjust.
Identities = 38/102 (37%), Positives = 49/102 (48%), Gaps = 3/102 (2%)
Query 40 DDLKYFILQMGGPEALEVLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAACDTPGDECP 99
DD +L+ ++ V +H G +W + ED C IC FE C C PGD+CP
Sbjct 36 DDHPGILLKTNTRMSITVKKLHVCGEWKWLQGGEDT-CGICRMEFESACNMCKFPGDDCP 94
Query 100 PAFGACGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQLWRFA 141
G C H FH HCI W+ + AQ CP+CRQ W
Sbjct 95 LVLGICRHAFHRHCIDKWIAAPTNQPRAQ--CPLCRQDWTIV 134
> ath:AT3G05870 APC11; APC11; protein binding / ubiquitin-protein
ligase/ zinc ion binding; K03358 anaphase-promoting complex
subunit 11
Length=57
Score = 58.5 bits (140), Expect = 8e-09, Method: Compositional matrix adjust.
Identities = 27/57 (47%), Positives = 35/57 (61%), Gaps = 4/57 (7%)
Query 84 FELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQLWRF 140
F+ C C PGD+CP +GAC H FHLHCI W+ + +Q CPMCR+ W+F
Sbjct 3 FDGCCPDCKLPGDDCPLIWGACNHAFHLHCILKWVNSQ----TSQAHCPMCRREWQF 55
> sce:YDL008W APC11; Apc11p (EC:6.3.2.-); K03358 anaphase-promoting
complex subunit 11
Length=165
Score = 55.8 bits (133), Expect = 5e-08, Method: Compositional matrix adjust.
Identities = 27/64 (42%), Positives = 33/64 (51%), Gaps = 4/64 (6%)
Query 77 CVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQ 136
C IC + C +C PGD+CP G C H FH HCI WL +G CPMCRQ
Sbjct 41 CGICRASYNGTCPSCKFPGDQCPLVIGLCHHNFHDHCIYRWLDTPTSKGL----CPMCRQ 96
Query 137 LWRF 140
++
Sbjct 97 TFQL 100
> sce:YOL133W HRT1, HRT2, RBX1, ROC1; Hrt1p; K03868 RING-box protein
1
Length=121
Score = 52.4 bits (124), Expect = 6e-07, Method: Compositional matrix adjust.
Identities = 27/71 (38%), Positives = 33/71 (46%), Gaps = 12/71 (16%)
Query 76 HCVICDNPFELPCAACD-----TPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQDT 130
+C IC N PC C +EC A+G C H FHLHCI W+ R D
Sbjct 54 NCAICRNHIMEPCIECQPKAMTDTDNECVAAWGVCNHAFHLHCINKWIKTR-------DA 106
Query 131 CPMCRQLWRFA 141
CP+ Q W+ A
Sbjct 107 CPLDNQPWQLA 117
> bbo:BBOV_III007590 17.m07664; hypothetical protein; K03868 RING-box
protein 1
Length=100
Score = 51.6 bits (122), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 31/92 (33%), Positives = 41/92 (44%), Gaps = 17/92 (18%)
Query 63 VGVCRWKEVPEDLHCVICDNPFELPCAACDTPGDE--------CPPAFGACGHIFHLHCI 114
V + WK ++ C IC N C C T E C A+GAC H FHLHCI
Sbjct 18 VALWNWKIAVDN--CAICRNHIMDLCIECQTAETEEQTSNEEQCSVAWGACSHAFHLHCI 75
Query 115 AAWLGGRGLRGAAQDTCPMCRQLWRFAAQQQH 146
+ WL R + CP+ W FA ++ +
Sbjct 76 SKWLKTRHV-------CPLDNAQWNFATKETN 100
> mmu:100043674 Gm9840; predicted gene 9840
Length=140
Score = 51.2 bits (121), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 30/96 (31%), Positives = 41/96 (42%), Gaps = 14/96 (14%)
Query 50 GGPEALEVLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAAC-----DTPGDECPPAFGA 104
G + EV + V + W V ++ C IC N C C +EC A+G
Sbjct 49 AGKKRFEVKKWNAVALWAWNIVVDN--CAICRNHIMDLCIECQANQASATSEECTVAWGV 106
Query 105 CGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQLWRF 140
C H FH HCI+ WL R + CP+ + W F
Sbjct 107 CNHAFHFHCISRWLKKRQV-------CPLDNREWEF 135
> cpv:cgd8_930 RBX1 ortholog, RING finger ; K03868 RING-box protein
1
Length=124
Score = 50.8 bits (120), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 30/99 (30%), Positives = 41/99 (41%), Gaps = 15/99 (15%)
Query 51 GPEALEVLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAACDTP------GDECPPAFGA 104
G + E+ + V + W V ++ C IC N C C +EC +G
Sbjct 33 GQKRFEIKKWNAVALWSWDIVVDN--CAICRNHIMDLCIECQANTGSAQRSEECTVTWGQ 90
Query 105 CGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQLWRFAAQ 143
C H FHLHCI+ WL R + CP+ W F Q
Sbjct 91 CNHAFHLHCISRWLKTRNV-------CPLDNTEWVFQKQ 122
> mmu:56438 Rbx1, 1500002P15Rik, AA517855, ROC1; ring-box 1; K03868
RING-box protein 1
Length=108
Score = 50.1 bits (118), Expect = 3e-06, Method: Compositional matrix adjust.
Identities = 30/96 (31%), Positives = 41/96 (42%), Gaps = 14/96 (14%)
Query 50 GGPEALEVLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAAC-----DTPGDECPPAFGA 104
G + EV + V + W V ++ C IC N C C +EC A+G
Sbjct 17 AGKKRFEVKKWNAVALWAWDIVVDN--CAICRNHIMDLCIECQANQASATSEECTVAWGV 74
Query 105 CGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQLWRF 140
C H FH HCI+ WL R + CP+ + W F
Sbjct 75 CNHAFHFHCISRWLKTRQV-------CPLDNREWEF 103
> hsa:9978 RBX1, BA554C12.1, FLJ60363, MGC13357, MGC1481, RNF75,
ROC1; ring-box 1, E3 ubiquitin protein ligase; K03868 RING-box
protein 1
Length=108
Score = 50.1 bits (118), Expect = 3e-06, Method: Compositional matrix adjust.
Identities = 30/96 (31%), Positives = 41/96 (42%), Gaps = 14/96 (14%)
Query 50 GGPEALEVLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAAC-----DTPGDECPPAFGA 104
G + EV + V + W V ++ C IC N C C +EC A+G
Sbjct 17 AGKKRFEVKKWNAVALWAWDIVVDN--CAICRNHIMDLCIECQANQASATSEECTVAWGV 74
Query 105 CGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQLWRF 140
C H FH HCI+ WL R + CP+ + W F
Sbjct 75 CNHAFHFHCISRWLKTRQV-------CPLDNREWEF 103
> dre:449846 rbx1, im:7137515, zgc:136444; ring-box 1; K03868
RING-box protein 1
Length=108
Score = 48.1 bits (113), Expect = 1e-05, Method: Compositional matrix adjust.
Identities = 29/96 (30%), Positives = 40/96 (41%), Gaps = 14/96 (14%)
Query 50 GGPEALEVLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAAC-----DTPGDECPPAFGA 104
+ EV + V + W V ++ C IC N C C +EC A+G
Sbjct 17 ASKKRFEVKKWNAVALWAWDIVVDN--CAICRNHIMDLCIECQANQASATSEECTVAWGV 74
Query 105 CGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQLWRF 140
C H FH HCI+ WL R + CP+ + W F
Sbjct 75 CNHAFHFHCISRWLKTRQV-------CPLDNREWEF 103
> cel:ZK287.5 rbx-1; yeast RBX (ring finger protein) homolog family
member (rbx-1); K03868 RING-box protein 1
Length=110
Score = 47.8 bits (112), Expect = 1e-05, Method: Compositional matrix adjust.
Identities = 25/70 (35%), Positives = 31/70 (44%), Gaps = 12/70 (17%)
Query 76 HCVICDNPFELPCAACDTP-----GDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQDT 130
+C IC N C C DEC A+G C H FH HCI+ WL R +
Sbjct 43 NCAICRNHIMDLCIECQANQAAGLKDECTVAWGNCNHAFHFHCISRWLKTRQV------- 95
Query 131 CPMCRQLWRF 140
CP+ + W F
Sbjct 96 CPLDNREWEF 105
> tgo:TGME49_013690 hypothetical protein ; K03868 RING-box protein
1
Length=128
Score = 46.2 bits (108), Expect = 4e-05, Method: Compositional matrix adjust.
Identities = 29/91 (31%), Positives = 38/91 (41%), Gaps = 14/91 (15%)
Query 55 LEVLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAACD-----TPGDECPPAFGACGHIF 109
EV V + W V ++ C IC N C C + +EC A+G C H F
Sbjct 42 FEVKKWSAVALWSWDIVVDN--CAICRNHIMDLCIECQASQGGSSSEECTVAWGVCNHAF 99
Query 110 HLHCIAAWLGGRGLRGAAQDTCPMCRQLWRF 140
H HCI+ WL R + CP+ W F
Sbjct 100 HFHCISRWLKTRQV-------CPLDNADWEF 123
> bbo:BBOV_IV010180 23.m06036; zinc finger, C3HC4 type domain
containing protein
Length=1151
Score = 46.2 bits (108), Expect = 4e-05, Method: Composition-based stats.
Identities = 23/67 (34%), Positives = 29/67 (43%), Gaps = 19/67 (28%)
Query 69 KEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQ 128
+E D C+IC + F+ C D CGHIFHL C+ +WL
Sbjct 294 EEKERDGTCIICRDEFDDDCRKID------------CGHIFHLSCLKSWL-------FQH 334
Query 129 DTCPMCR 135
TCP CR
Sbjct 335 STCPTCR 341
> dre:447861 rnf103, zgc:92077; ring finger protein 103
Length=656
Score = 45.4 bits (106), Expect = 7e-05, Method: Composition-based stats.
Identities = 21/59 (35%), Positives = 25/59 (42%), Gaps = 16/59 (27%)
Query 77 CVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQDTCPMCR 135
CV+C FE C P CGH+FH CI WL G + CP+CR
Sbjct 594 CVVCLENFETDCLVMGLP----------CGHVFHQQCIVVWLAG------GRHCCPVCR 636
> hsa:7844 RNF103, HKF-1, KF-1, KF1, MGC102815, MGC41857, ZFP-103,
ZFP103; ring finger protein 103
Length=681
Score = 44.7 bits (104), Expect = 1e-04, Method: Composition-based stats.
Identities = 24/72 (33%), Positives = 30/72 (41%), Gaps = 20/72 (27%)
Query 68 WKEVPEDL----HCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGL 123
W P D+ CV+C FE C P CGH+FH +CI WL G
Sbjct 604 WLTWPADMLHCTECVVCLENFENGCLLMGLP----------CGHVFHQNCIVMWLAG--- 650
Query 124 RGAAQDTCPMCR 135
+ CP+CR
Sbjct 651 ---GRHCCPVCR 659
> mmu:22644 Rnf103, AW146237, AW212918, Zfp103, kf-1; ring finger
protein 103
Length=683
Score = 44.7 bits (104), Expect = 1e-04, Method: Composition-based stats.
Identities = 28/78 (35%), Positives = 34/78 (43%), Gaps = 24/78 (30%)
Query 77 CVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQ 136
CV+C FE C P CGH+FH +CI WL G + CP+CR
Sbjct 619 CVVCLENFENGCLLMGLP----------CGHVFHQNCIVMWLAG------GRHCCPVCR- 661
Query 137 LW------RFAAQQQHLS 148
W + AQQQ LS
Sbjct 662 -WPSYKKKQPYAQQQPLS 678
> tpv:TP04_0669 hypothetical protein; K03868 RING-box protein
1
Length=101
Score = 43.5 bits (101), Expect = 2e-04, Method: Compositional matrix adjust.
Identities = 27/88 (30%), Positives = 38/88 (43%), Gaps = 19/88 (21%)
Query 63 VGVCRWKEVPEDLHCVICDNPFELPCAACDTPGDE----------CPPAFGACGHIFHLH 112
V V W+ ++ C IC N C C ++ C ++GACGH FHLH
Sbjct 18 VAVWSWRMSVDN--CAICRNHIMDHCLDCQAKHNKESQDITEQPGCTISWGACGHAFHLH 75
Query 113 CIAAWLGGRGLRGAAQDTCPMCRQLWRF 140
CI+ WL R + CP+ W +
Sbjct 76 CISTWLKTRRV-------CPLDNTQWDY 96
> ath:AT3G16090 zinc finger (C3HC4-type RING finger) family protein;
K10601 E3 ubiquitin-protein ligase synoviolin [EC:6.3.2.19]
Length=492
Score = 43.5 bits (101), Expect = 3e-04, Method: Composition-based stats.
Identities = 16/33 (48%), Positives = 20/33 (60%), Gaps = 7/33 (21%)
Query 105 CGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQL 137
CGH+FH+HC+ +WL Q TCP CR L
Sbjct 307 CGHLFHVHCLRSWL-------ERQQTCPTCRAL 332
> ath:AT3G42830 ring-box protein Roc1/Rbx1/Hrt1, putative; K03868
RING-box protein 1
Length=115
Score = 43.1 bits (100), Expect = 3e-04, Method: Compositional matrix adjust.
Identities = 20/53 (37%), Positives = 25/53 (47%), Gaps = 5/53 (9%)
Query 76 HCVICDNPFELPCAAC-----DTPGDECPPAFGACGHIFHLHCIAAWLGGRGL 123
+C IC N C C +EC A+G C H FH HCI+ WL R +
Sbjct 48 NCAICRNHIMDLCIECLANQASATSEECTVAWGVCNHAFHFHCISRWLKTRQV 100
> ath:AT1G65040 protein binding / zinc ion binding; K10601 E3
ubiquitin-protein ligase synoviolin [EC:6.3.2.19]
Length=389
Score = 43.1 bits (100), Expect = 3e-04, Method: Compositional matrix adjust.
Identities = 26/89 (29%), Positives = 40/89 (44%), Gaps = 11/89 (12%)
Query 53 EALEVLAVHTVGVCRWKEVPEDLHCVICD-NPFELPC--AACDTPGDECPPAFG-ACGHI 108
E + R++++ +++ D P EL A C +E A CGH+
Sbjct 180 ETFRNFKIRVTDYLRYRKITSNMNDRFPDATPEELSSNDATCIICREEMTSAKKLVCGHL 239
Query 109 FHLHCIAAWLGGRGLRGAAQDTCPMCRQL 137
FH+HC+ +WL Q+TCP CR L
Sbjct 240 FHVHCLRSWL-------ERQNTCPTCRAL 261
> pfa:PFC0845c ubiquitin-protein ligase, putative (EC:6.3.2.19);
K03868 RING-box protein 1
Length=107
Score = 43.1 bits (100), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 25/78 (32%), Positives = 32/78 (41%), Gaps = 20/78 (25%)
Query 76 HCVICDNPFELPCAAC-----DTPGDE--------CPPAFGACGHIFHLHCIAAWLGGRG 122
+C IC N C C D D+ C A+G C H FHLHCI+ W+ R
Sbjct 32 NCAICRNHIMDLCIECQAKTTDHENDKDKKIDKEGCTVAWGVCNHAFHLHCISRWIKARQ 91
Query 123 LRGAAQDTCPMCRQLWRF 140
+ CP+ W F
Sbjct 92 V-------CPLDNTTWEF 102
> cel:T20F5.6 hypothetical protein
Length=794
Score = 42.4 bits (98), Expect = 6e-04, Method: Composition-based stats.
Identities = 24/67 (35%), Positives = 27/67 (40%), Gaps = 9/67 (13%)
Query 75 LHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQDTCPMC 134
L C IC +PF G P F CGH L CI L R G CP C
Sbjct 114 LSCGICYDPF--------NTGKRIPKVF-PCGHTICLQCIKKLLNTRTFLGGNTVICPSC 164
Query 135 RQLWRFA 141
RQ R++
Sbjct 165 RQNTRYS 171
> ath:AT5G27420 zinc finger (C3HC4-type RING finger) family protein
Length=368
Score = 42.4 bits (98), Expect = 6e-04, Method: Compositional matrix adjust.
Identities = 25/61 (40%), Positives = 27/61 (44%), Gaps = 16/61 (26%)
Query 75 LHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQDTCPMC 134
L C IC N FE DE C H+FH HCI AWL G TCP+C
Sbjct 122 LECAICLNEFE---------DDETLRLLPKCDHVFHPHCIGAWLQG-------HVTCPVC 165
Query 135 R 135
R
Sbjct 166 R 166
> ath:AT3G05200 ATL6; ATL6; protein binding / zinc ion binding
Length=398
Score = 42.0 bits (97), Expect = 7e-04, Method: Compositional matrix adjust.
Identities = 25/62 (40%), Positives = 28/62 (45%), Gaps = 16/62 (25%)
Query 74 DLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQDTCPM 133
+L C IC N FE DE C H+FH HCI AWL A TCP+
Sbjct 125 ELECAICLNEFE---------DDETLRLLPKCDHVFHPHCIDAWL-------EAHVTCPV 168
Query 134 CR 135
CR
Sbjct 169 CR 170
> ath:AT3G56580 zinc finger (C3HC4-type RING finger) family protein;
K11982 E3 ubiquitin-protein ligase RNF115/126 [EC:6.3.2.19]
Length=320
Score = 42.0 bits (97), Expect = 8e-04, Method: Compositional matrix adjust.
Identities = 23/80 (28%), Positives = 35/80 (43%), Gaps = 17/80 (21%)
Query 57 VLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAA 116
+ A+ T+ + + D HC +C + FEL A P C HI+H CI
Sbjct 166 IDALPTIKITQKHLKSSDSHCPVCKDEFELKSEAKQMP----------CHHIYHSDCIVP 215
Query 117 WLGGRGLRGAAQDTCPMCRQ 136
WL ++CP+CR+
Sbjct 216 WL-------VQHNSCPVCRK 228
> ath:AT2G40830 RHC1A; RHC1A; protein binding / zinc ion binding;
K11982 E3 ubiquitin-protein ligase RNF115/126 [EC:6.3.2.19]
Length=328
Score = 41.6 bits (96), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 23/80 (28%), Positives = 35/80 (43%), Gaps = 17/80 (21%)
Query 57 VLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAA 116
+ A+ T+ + + D +C +C + FEL A P C HI+H CI
Sbjct 170 IDALPTIKIAQRHLRSSDSNCPVCKDEFELGSEAKQMP----------CNHIYHSDCIVP 219
Query 117 WLGGRGLRGAAQDTCPMCRQ 136
WL ++CP+CRQ
Sbjct 220 WL-------VQHNSCPVCRQ 232
> ath:AT5G01980 zinc finger (C3HC4-type RING finger) family protein
Length=493
Score = 41.2 bits (95), Expect = 0.001, Method: Composition-based stats.
Identities = 21/67 (31%), Positives = 30/67 (44%), Gaps = 17/67 (25%)
Query 69 KEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQ 128
+ V + L C IC F L P C H++H HCI WL +A+
Sbjct 342 EHVMKGLVCAICKELFSLRNETTQLP----------CLHLYHAHCIVPWL-------SAR 384
Query 129 DTCPMCR 135
++CP+CR
Sbjct 385 NSCPLCR 391
> ath:AT2G27940 zinc finger (C3HC4-type RING finger) family protein
Length=237
Score = 40.4 bits (93), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 29/94 (30%), Positives = 43/94 (45%), Gaps = 22/94 (23%)
Query 54 ALEVLAVHTVGVCRW----KEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIF 109
L+ AV ++ V R+ K+ ED CVIC + FE E CGH+F
Sbjct 115 GLDSQAVRSLPVYRYTKAAKQRNED--CVICLSDFE---------EGETVKVIPHCGHVF 163
Query 110 HLHCIAAWLGGRGLRGAAQDTCPMCRQLWRFAAQ 143
H+ C+ WL ++ TCP+CR F+ +
Sbjct 164 HVDCVDTWL-------SSYVTCPLCRSNQLFSDK 190
> ath:AT3G48030 hypoxia-responsive family protein / zinc finger
(C3HC4-type RING finger) family protein
Length=349
Score = 40.0 bits (92), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 35/104 (33%), Positives = 43/104 (41%), Gaps = 21/104 (20%)
Query 36 FRAPDDLKYFILQMGGPE--ALEVLAVHTVG-VCRWKEVPEDLHCVICDNPFELPCAACD 92
F +P F L G + A++ L V G V E P D C +C N F D
Sbjct 165 FSSPQLQHLFFLHDSGLDQTAIDALPVFLYGNVTISLEQPFD--CAVCLNEFS------D 216
Query 93 TPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQ 136
T P C H FHLHCI WL + TCP+CR+
Sbjct 217 TDKLRLLPV---CSHAFHLHCIDTWL-------LSNSTCPLCRR 250
> tpv:TP01_0655 hypothetical protein
Length=618
Score = 40.0 bits (92), Expect = 0.003, Method: Composition-based stats.
Identities = 18/65 (27%), Positives = 31/65 (47%), Gaps = 19/65 (29%)
Query 73 EDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQDTCP 132
++L+C+IC + + + CGH+FHL+C+ +WL + CP
Sbjct 292 KNLNCIICRDVITVNSRKLE------------CGHVFHLNCLKSWL-------FQHNNCP 332
Query 133 MCRQL 137
CR+L
Sbjct 333 SCRKL 337
> dre:565543 ring finger protein 6-like
Length=750
Score = 40.0 bits (92), Expect = 0.003, Method: Composition-based stats.
Identities = 15/32 (46%), Positives = 19/32 (59%), Gaps = 7/32 (21%)
Query 105 CGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQ 136
C H FH+HCI WL + +TCP+CRQ
Sbjct 718 CAHEFHIHCIDRWL-------SENNTCPICRQ 742
> cpv:cgd8_2560 HRD1 like membrane associated RING finger containing
protein signal peptide plus 6 transmembrane domains ;
K10601 E3 ubiquitin-protein ligase synoviolin [EC:6.3.2.19]
Length=637
Score = 40.0 bits (92), Expect = 0.003, Method: Composition-based stats.
Identities = 14/31 (45%), Positives = 18/31 (58%), Gaps = 7/31 (22%)
Query 105 CGHIFHLHCIAAWLGGRGLRGAAQDTCPMCR 135
CGH+FHL C+ +W Q TCP+CR
Sbjct 353 CGHVFHLDCLKSWF-------IQQQTCPICR 376
> xla:398305 rnf103-a, kf1, rnf103, x-kf-1a, zfp103; ring finger
protein 103
Length=678
Score = 39.7 bits (91), Expect = 0.004, Method: Composition-based stats.
Identities = 22/72 (30%), Positives = 29/72 (40%), Gaps = 20/72 (27%)
Query 68 WKEVPEDL----HCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGL 123
W P ++ CV+C FE P CGH+FH +CI WL G
Sbjct 602 WATWPGNMLHCTECVVCLENFENRSLLMGLP----------CGHVFHQNCIVMWLAG--- 648
Query 124 RGAAQDTCPMCR 135
+ CP+CR
Sbjct 649 ---GRHCCPVCR 657
> ath:AT5G10380 RING1; RING1; protein binding / ubiquitin-protein
ligase/ zinc ion binding
Length=301
Score = 39.7 bits (91), Expect = 0.004, Method: Compositional matrix adjust.
Identities = 22/79 (27%), Positives = 33/79 (41%), Gaps = 16/79 (20%)
Query 57 VLAVHTVGVCRWKEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAA 116
+ ++ VG + + + + C +C N FE DE C H FHL+CI
Sbjct 115 INSITVVGFKKGEGIIDGTECSVCLNEFE---------EDESLRLLPKCSHAFHLNCIDT 165
Query 117 WLGGRGLRGAAQDTCPMCR 135
WL + CP+CR
Sbjct 166 WL-------LSHKNCPLCR 177
> pfa:PF10_0276 Zinc finger, C3HC4 type, putative
Length=274
Score = 39.3 bits (90), Expect = 0.004, Method: Compositional matrix adjust.
Identities = 23/68 (33%), Positives = 29/68 (42%), Gaps = 16/68 (23%)
Query 69 KEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQ 128
K + + C IC N F++ DEC C H FH CI WL +R A
Sbjct 212 KNISNESKCSICLNDFQI---------DECVRTLLLCNHTFHKSCIDLWL----IRSA-- 256
Query 129 DTCPMCRQ 136
TCP C+
Sbjct 257 -TCPNCKS 263
> ath:AT1G63840 zinc finger (C3HC4-type RING finger) family protein
Length=166
Score = 39.3 bits (90), Expect = 0.005, Method: Compositional matrix adjust.
Identities = 26/89 (29%), Positives = 39/89 (43%), Gaps = 16/89 (17%)
Query 49 MGGPEALEVLAVHTVGVCRWKEV--PEDLHCVICDNPFELPCAACDTPGDECPPAFGACG 106
+ P++ +LA + V R+ ++ PE C +C FE D+ C
Sbjct 59 LTKPDSAAILAGEMLPVVRFSDINRPESECCAVCLYDFE---------NDDEIRRLTNCR 109
Query 107 HIFHLHCIAAWLGGRGLRGAAQDTCPMCR 135
HIFH C+ W+ G Q TCP+CR
Sbjct 110 HIFHRGCLDRWMMGYN-----QMTCPLCR 133
> tgo:TGME49_092340 zinc finger (C3HC4 RING finger) protein, putative
Length=988
Score = 39.3 bits (90), Expect = 0.005, Method: Compositional matrix adjust.
Identities = 17/34 (50%), Positives = 21/34 (61%), Gaps = 5/34 (14%)
Query 104 ACGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQL 137
+CGH FH HC+ WL R ++TCPMC QL
Sbjct 953 SCGHKFHRHCVDVWLFKR-----QKETCPMCGQL 981
> xla:399120 rnf103-b, KF-1a, MGC99268, Rnf103, kf1, x-kf-1a,
x-kf-1b, zfp103; ring finger protein 103
Length=670
Score = 39.3 bits (90), Expect = 0.005, Method: Composition-based stats.
Identities = 22/72 (30%), Positives = 29/72 (40%), Gaps = 20/72 (27%)
Query 68 WKEVPEDL----HCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLGGRGL 123
W P ++ CV+C FE P CGH+FH +CI WL G
Sbjct 594 WATWPGNMLHCTECVVCLENFENGSLLMGLP----------CGHVFHQNCIVMWLAG--- 640
Query 124 RGAAQDTCPMCR 135
+ CP+CR
Sbjct 641 ---GRHCCPVCR 649
> cpv:cgd7_4910 conserved 3 transmembrane domain membrane associated
RING finger domain (shared by plants and apicomplexans)
Length=311
Score = 39.3 bits (90), Expect = 0.005, Method: Compositional matrix adjust.
Identities = 26/67 (38%), Positives = 31/67 (46%), Gaps = 9/67 (13%)
Query 84 FELPCAAC--DTPGDECPPAFGACGHIFHLHCIAAWLGGRGLRGAAQDTCPMCRQLWRFA 141
F PC+ C D +E ACGH FH CI AWL LR A CP C+ L R+
Sbjct 208 FNDPCSICIEDFKLNEEIRTLPACGHSFHKGCIDAWL----LRNA---ICPNCKTLVRYG 260
Query 142 AQQQHLS 148
+ S
Sbjct 261 KTETGTS 267
> ath:AT1G73760 zinc finger (C3HC4-type RING finger) family protein
Length=367
Score = 38.9 bits (89), Expect = 0.006, Method: Compositional matrix adjust.
Identities = 22/78 (28%), Positives = 35/78 (44%), Gaps = 17/78 (21%)
Query 60 VHTVGVCRWKEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFHLHCIAAWLG 119
+ V CR D C+IC + +E A D G+ CGH FH+ C+ WL
Sbjct 302 LRKVKPCRQDTTVADRKCIICQDEYE----AKDEVGEL------RCGHRFHIDCVNQWL- 350
Query 120 GRGLRGAAQDTCPMCRQL 137
+++CP+C+ +
Sbjct 351 ------VRKNSCPVCKTM 362
> ath:AT4G10160 zinc finger (C3HC4-type RING finger) family protein
Length=225
Score = 38.9 bits (89), Expect = 0.006, Method: Compositional matrix adjust.
Identities = 24/72 (33%), Positives = 33/72 (45%), Gaps = 12/72 (16%)
Query 69 KEVPEDLHCVICDNPF---ELPCAAC--DTPGDECPPAFGACGHIFHLHCIAAWLGGRGL 123
K++ E L VI F + C+ C D +E +CGH FH+ CI WL
Sbjct 75 KDIREMLPIVIYKESFTVNDTQCSVCLGDYQAEEKLQQMPSCGHTFHMECIDLWL----- 129
Query 124 RGAAQDTCPMCR 135
+ TCP+CR
Sbjct 130 --TSHTTCPLCR 139
> ath:AT4G10150 zinc finger (C3HC4-type RING finger) family protein
Length=236
Score = 38.9 bits (89), Expect = 0.007, Method: Compositional matrix adjust.
Identities = 24/72 (33%), Positives = 33/72 (45%), Gaps = 12/72 (16%)
Query 69 KEVPEDLHCVICDNPF---ELPCAAC--DTPGDECPPAFGACGHIFHLHCIAAWLGGRGL 123
K++ E L VI F + C+ C D +E +CGH FH+ CI WL
Sbjct 89 KDIREMLPVVIYKESFIVKDSQCSVCLGDYQAEEKLQQMPSCGHTFHMECIDLWL----- 143
Query 124 RGAAQDTCPMCR 135
+ TCP+CR
Sbjct 144 --TSHTTCPLCR 153
> ath:AT1G72220 zinc finger (C3HC4-type RING finger) family protein
Length=413
Score = 38.5 bits (88), Expect = 0.007, Method: Compositional matrix adjust.
Identities = 23/85 (27%), Positives = 36/85 (42%), Gaps = 19/85 (22%)
Query 54 ALEVLAVHTVGVCRWKE---VPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHIFH 110
L+ ++++ +C +K + E C +C N FE DE C H FH
Sbjct 151 GLQQSIINSITICNYKRGDGLIERTDCPVCLNEFEE---------DESLRLLPKCNHAFH 201
Query 111 LHCIAAWLGGRGLRGAAQDTCPMCR 135
+ CI WL ++ CP+CR
Sbjct 202 ISCIDTWL-------SSHTNCPLCR 219
> ath:AT3G10815 zinc finger (C3HC4-type RING finger) family protein
Length=199
Score = 38.5 bits (88), Expect = 0.007, Method: Compositional matrix adjust.
Identities = 26/88 (29%), Positives = 40/88 (45%), Gaps = 19/88 (21%)
Query 51 GPEALEVLAVHTVGVC--RWKEVPEDLHCVICDNPFELPCAACDTPGDECPPAFGACGHI 108
GP + A++++ R K + D +C +C + FE+ A P C HI
Sbjct 93 GPPPASLAAINSLQKIKIRQKHLGLDPYCPVCQDQFEIGSDARKMP----------CKHI 142
Query 109 FHLHCIAAWLGGRGLRGAAQDTCPMCRQ 136
+H CI WL R +TCP+CR+
Sbjct 143 YHSECILPWLVQR-------NTCPVCRK 163
Lambda K H
0.331 0.145 0.525
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3199347004
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
Posted date: Sep 17, 2011 2:57 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40