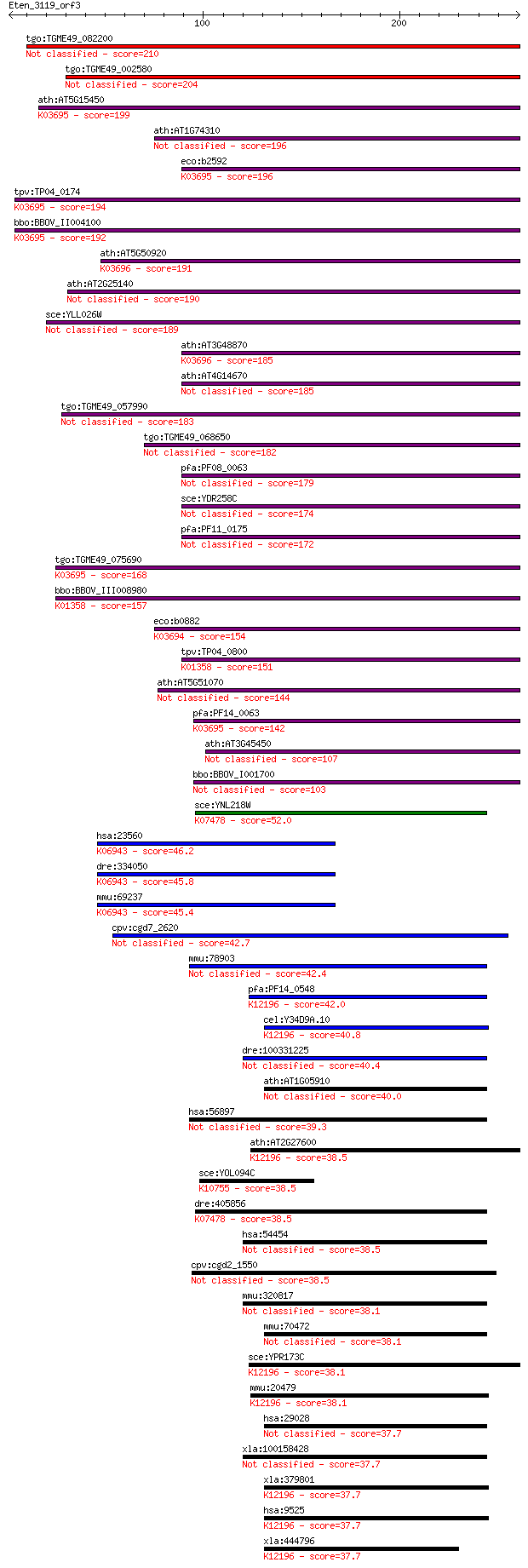
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
164,496 sequences; 82,071,388 total letters
Query= Eten_3119_orf3
Length=260
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_082200 clpB protein, putative 210 4e-54
tgo:TGME49_002580 heat shock protein, putative (EC:3.4.21.53) 204 3e-52
ath:AT5G15450 CLPB3; CLPB3 (CASEIN LYTIC PROTEINASE B3); ATP b... 199 1e-50
ath:AT1G74310 ATHSP101 (ARABIDOPSIS THALIANA HEAT SHOCK PROTEI... 196 5e-50
eco:b2592 clpB, ECK2590, htpM, JW2573; protein disaggregation ... 196 1e-49
tpv:TP04_0174 hypothetical protein; K03695 ATP-dependent Clp p... 194 3e-49
bbo:BBOV_II004100 18.m06340; ClpB; K03695 ATP-dependent Clp pr... 192 1e-48
ath:AT5G50920 CLPC1; CLPC1; ATP binding / ATP-dependent peptid... 191 3e-48
ath:AT2G25140 CLPB4; CLPB4 (CASEIN LYTIC PROTEINASE B4); ATP b... 190 5e-48
sce:YLL026W HSP104; Heat shock protein that cooperates with Yd... 189 8e-48
ath:AT3G48870 HSP93-III; ATP binding / ATPase/ DNA binding / n... 185 2e-46
ath:AT4G14670 CLPB2; CLPB2; ATP binding / nucleoside-triphosph... 185 2e-46
tgo:TGME49_057990 heat shock protein, putative (EC:3.4.21.53) 183 4e-46
tgo:TGME49_068650 clp ATP-binding chain B1, putative (EC:3.4.2... 182 2e-45
pfa:PF08_0063 ClpB protein, putative 179 7e-45
sce:YDR258C HSP78; Hsp78p 174 2e-43
pfa:PF11_0175 heat shock protein 101, putative 172 1e-42
tgo:TGME49_075690 chaperone clpB 1 protein, putative (EC:3.4.2... 168 2e-41
bbo:BBOV_III008980 17.m07783; Clp amino terminal domain contai... 157 4e-38
eco:b0882 clpA, ECK0873, JW0866, lopD; ATPase and specificity ... 154 4e-37
tpv:TP04_0800 ATP-dependent Clp protease ATP-binding subunit (... 151 3e-36
ath:AT5G51070 ERD1; ERD1 (EARLY RESPONSIVE TO DEHYDRATION 1); ... 144 4e-34
pfa:PF14_0063 ATP-dependent CLP protease, putative; K03695 ATP... 142 1e-33
ath:AT3G45450 Clp amino terminal domain-containing protein 107 4e-23
bbo:BBOV_I001700 19.m02115; chaperone clpB 103 8e-22
sce:YNL218W MGS1; Mgs1p; K07478 putative ATPase 52.0 2e-06
hsa:23560 GTPBP4, CRFG, FLJ10686, FLJ10690, FLJ39774, NGB, NOG... 46.2 1e-04
dre:334050 gtpbp4, fi28d07, wu:fi28d07, zgc:55757; GTP binding... 45.8 1e-04
mmu:69237 Gtpbp4, 2610028C09Rik, Crfg, Gtpbp3, NGB, Nog1; GTP ... 45.4 2e-04
cpv:cgd7_2620 ClpB ATpase (bacterial), signal peptide 42.7 0.001
mmu:78903 Wrnip1, 4833444L21Rik, WHIP, Wrnip; Werner helicase ... 42.4 0.002
pfa:PF14_0548 ATPase, putative; K12196 vacuolar protein-sortin... 42.0 0.002
cel:Y34D9A.10 vps-4; related to yeast Vacuolar Protein Sorting... 40.8 0.005
dre:100331225 valosin-containing protein-like 40.4 0.006
ath:AT1G05910 cell division cycle protein 48-related / CDC48-r... 40.0 0.009
hsa:56897 WRNIP1, FLJ22526, RP11-420G6.2, WHIP, bA420G6.2; Wer... 39.3 0.015
ath:AT2G27600 SKD1; SKD1 (SUPPRESSOR OF K+ TRANSPORT GROWTH DE... 38.5 0.023
sce:YOL094C RFC4; Rfc4p; K10755 replication factor C subunit 2/4 38.5
dre:405856 MGC85976; zgc:85976; K07478 putative ATPase 38.5 0.027
hsa:54454 ATAD2B, KIAA1240, MGC88424; ATPase family, AAA domai... 38.5 0.027
cpv:cgd2_1550 origin recognition complex 4 orc4p like AAA+ ATpase 38.5 0.028
mmu:320817 Atad2b, 1110014E10Rik, BC032887, C79189, D530031C13... 38.1 0.029
mmu:70472 Atad2, 2610509G12Rik, MGC38189; ATPase family, AAA d... 38.1 0.030
sce:YPR173C VPS4, CSC1, DID6, END13, GRD13, VPL4, VPT10; AAA-A... 38.1 0.032
mmu:20479 Vps4b, 8030489C12Rik, Skd1; vacuolar protein sorting... 38.1 0.035
hsa:29028 ATAD2, ANCCA, DKFZp667N1320, MGC131938, MGC142216, M... 37.7 0.041
xla:100158428 atad2b; ATPase family, AAA domain containing 2B 37.7 0.042
xla:379801 vps4b, MGC53483; vacuolar protein sorting 4 homolog... 37.7 0.043
hsa:9525 VPS4B, SKD1, SKD1B, VPS4-2; vacuolar protein sorting ... 37.7 0.044
xla:444796 MGC82073 protein; K12196 vacuolar protein-sorting-a... 37.7 0.046
> tgo:TGME49_082200 clpB protein, putative
Length=970
Score = 210 bits (535), Expect = 4e-54, Method: Compositional matrix adjust.
Identities = 117/268 (43%), Positives = 167/268 (62%), Gaps = 18/268 (6%)
Query 10 TAAVVEVFLRARRMAEAES-CLVSMQHIFLALLHSPTFYRTLVNSGCDIDLFKEELALAY 68
T + V LRA +A+ + + M+H+ +L ++L +GC++ +F E Y
Sbjct 141 TPQLKRVALRALEIAQKDKKQTIGMEHLITSLFEENNLRKSLEQAGCNVKVFLEHSTTMY 200
Query 69 KNT-------------PM---PSTSGSTEHVAKDFDLSQYGEDLTEKARLGKLQPVIGRQ 112
+N P P+TS + + + + +G D+T+ A GKL+PV+GR
Sbjct 201 ENKFKQDVASRSMKKQPTEGSPATSRAKPDASDEEFVQSFGVDMTKLAAEGKLEPVVGRN 260
Query 113 EEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLN 172
+EI + +LSR P L+GEPGVGKTAV+EGLAQR+VE VPK+L+++ +F++DL
Sbjct 261 KEIKEVLTVLSRKGKGNPCLVGEPGVGKTAVVEGLAQRLVEGMVPKSLENKILFAVDLGA 320
Query 173 LSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPLA 232
L AG + RGEFE+RMK +I Y + VILF+DE+H L+GAGK+ G+ DA +LK P+A
Sbjct 321 LIAGATYRGEFEKRMKALIRYAVNQEGRVILFIDELHMLMGAGKSDGTMDAANLLKPPMA 380
Query 233 RGEIVLVGATTLEEYKLHIEKDAAFCRR 260
RGEI LVGATT EEYK+ IEKDAA RR
Sbjct 381 RGEIRLVGATTQEEYKI-IEKDAAMERR 407
> tgo:TGME49_002580 heat shock protein, putative (EC:3.4.21.53)
Length=983
Score = 204 bits (519), Expect = 3e-52, Method: Compositional matrix adjust.
Identities = 112/238 (47%), Positives = 151/238 (63%), Gaps = 13/238 (5%)
Query 30 LVSMQHIFLALLHSPTFYRTLVNSGCDIDLFKEELALAY-------KNTPMPSTSGSTEH 82
LV + + AL F L N G D+ +F + + A+ + P+ +GS E
Sbjct 202 LVGVHDLVTALARCDRFRARLANVGTDVSVFLKLVTSAFIVHTEAEERKATPARTGSQEW 261
Query 83 VAKDFDLSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTA 142
+ +G D+TE+A+ GK+ V GR+ EI++I ++SRM LLIGEPGVGKTA
Sbjct 262 I------KTFGVDMTEQAKEGKIGTVTGREAEIEQITSVMSRMSKANCLLIGEPGVGKTA 315
Query 143 VIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVI 202
V+EGLA+RIV+ VP AL ++FSLD+ +L +G+SMRGEFERRMK I+ YL R I
Sbjct 316 VVEGLAKRIVDGDVPDALLGVQVFSLDVGSLLSGSSMRGEFERRMKGILDYLFASDRSTI 375
Query 203 LFVDEIHTLVGAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
LF+DEIHTL+GAGK G DA +LK LARG + ++GATT EY+ HIE+D AF RR
Sbjct 376 LFIDEIHTLMGAGKADGPMDAANLLKPALARGALRVIGATTRAEYRKHIERDMAFARR 433
> ath:AT5G15450 CLPB3; CLPB3 (CASEIN LYTIC PROTEINASE B3); ATP
binding / ATPase/ nucleoside-triphosphatase/ nucleotide binding
/ protein binding; K03695 ATP-dependent Clp protease ATP-binding
subunit ClpB
Length=968
Score = 199 bits (505), Expect = 1e-50, Method: Compositional matrix adjust.
Identities = 113/246 (45%), Positives = 154/246 (62%), Gaps = 4/246 (1%)
Query 16 VFLRARRMA-EAESCLVSMQHIFLALLHSPTFYRTLVNSGCDIDLFKEELALAYKNTPMP 74
+F RAR+ + + VS++H+ LA F + L D + + L A ++
Sbjct 166 LFQRARQFKKDLKDSYVSVEHLVLAFADDKRFGKQLFK---DFQISERSLKSAIESIRGK 222
Query 75 STSGSTEHVAKDFDLSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIG 134
+ + K L +YG+DLT AR GKL PVIGR +EI R QILSR P+LIG
Sbjct 223 QSVIDQDPEGKYEALEKYGKDLTAMAREGKLDPVIGRDDEIRRCIQILSRRTKNNPVLIG 282
Query 135 EPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYL 194
EPGVGKTA+ EGLAQRIV+ VP+AL +R++ SLD+ L AG RGEFE R+K ++ +
Sbjct 283 EPGVGKTAISEGLAQRIVQGDVPQALMNRKLISLDMGALIAGAKYRGEFEDRLKAVLKEV 342
Query 195 QRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKD 254
+ ++ILF+DEIHT+VGAG T G+ DA +LK L RGE+ +GATTL+EY+ +IEKD
Sbjct 343 TDSEGQIILFIDEIHTVVGAGATNGAMDAGNLLKPMLGRGELRCIGATTLDEYRKYIEKD 402
Query 255 AAFCRR 260
A RR
Sbjct 403 PALERR 408
> ath:AT1G74310 ATHSP101 (ARABIDOPSIS THALIANA HEAT SHOCK PROTEIN
101); ATP binding / ATPase/ nucleoside-triphosphatase/ nucleotide
binding / protein binding
Length=911
Score = 196 bits (499), Expect = 5e-50, Method: Compositional matrix adjust.
Identities = 105/186 (56%), Positives = 131/186 (70%), Gaps = 6/186 (3%)
Query 75 STSGSTEHVAKDFDLSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIG 134
S SG T A L YG DL E+A GKL PVIGR EEI R+ +ILSR P+LIG
Sbjct 154 SASGDTNFQA----LKTYGRDLVEQA--GKLDPVIGRDEEIRRVVRILSRRTKNNPVLIG 207
Query 135 EPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYL 194
EPGVGKTAV+EGLAQRIV+ VP +L D R+ SLD+ L AG RGEFE R+K ++ +
Sbjct 208 EPGVGKTAVVEGLAQRIVKGDVPNSLTDVRLISLDMGALVAGAKYRGEFEERLKSVLKEV 267
Query 195 QRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKD 254
+ + +VILF+DEIH ++GAGKT GS DA + K LARG++ +GATTLEEY+ ++EKD
Sbjct 268 EDAEGKVILFIDEIHLVLGAGKTEGSMDAANLFKPMLARGQLRCIGATTLEEYRKYVEKD 327
Query 255 AAFCRR 260
AAF RR
Sbjct 328 AAFERR 333
> eco:b2592 clpB, ECK2590, htpM, JW2573; protein disaggregation
chaperone; K03695 ATP-dependent Clp protease ATP-binding subunit
ClpB
Length=857
Score = 196 bits (497), Expect = 1e-49, Method: Compositional matrix adjust.
Identities = 98/172 (56%), Positives = 124/172 (72%), Gaps = 0/172 (0%)
Query 89 LSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLA 148
L +Y DLTE+A GKL PVIGR EEI R Q+L R P+LIGEPGVGKTA++EGLA
Sbjct 161 LKKYTIDLTERAEQGKLDPVIGRDEEIRRTIQVLQRRTKNNPVLIGEPGVGKTAIVEGLA 220
Query 149 QRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEI 208
QRI+ VP+ L+ RR+ +LD+ L AG RGEFE R+K ++ L + + VILF+DE+
Sbjct 221 QRIINGEVPEGLKGRRVLALDMGALVAGAKYRGEFEERLKGVLNDLAKQEGNVILFIDEL 280
Query 209 HTLVGAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
HT+VGAGK G+ DA +LK LARGE+ VGATTL+EY+ +IEKDAA RR
Sbjct 281 HTMVGAGKADGAMDAGNMLKPALARGELHCVGATTLDEYRQYIEKDAALERR 332
> tpv:TP04_0174 hypothetical protein; K03695 ATP-dependent Clp
protease ATP-binding subunit ClpB
Length=985
Score = 194 bits (493), Expect = 3e-49, Method: Composition-based stats.
Identities = 113/260 (43%), Positives = 164/260 (63%), Gaps = 6/260 (2%)
Query 4 LGGAKVTAAVVEVFLRARRMAEAE--SCLVSMQHIFLALLHSPT-FYRTLVNSGCDIDLF 60
G K+ ++E L + ++E +S+ H+ L L T F+R + + L
Sbjct 178 FGDRKILGRILENVLNISKRYKSEFGDKYISVDHLLLGLAAEDTKFFRPYLTRN-KVTLE 236
Query 61 KEELALAYKNTPMPSTSGSTEHVAKDFDLSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQ 120
K + ++ TS +TE+ K L+++ +DLT+ AR GKL PVIGR EI R +
Sbjct 237 KLKDSVLSIRGKRKITSRNTENSYKL--LNKFSKDLTDMARNGKLDPVIGRDNEIRRTIE 294
Query 121 ILSRMMTKAPLLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMR 180
ILSR P+L+G+PGVGKTA+ EGLA RIV VP +L++R++ SLD+ + AGT R
Sbjct 295 ILSRRTKNNPVLLGDPGVGKTAIAEGLANRIVSGDVPDSLKNRKVLSLDIAAIVAGTMYR 354
Query 181 GEFERRMKDIIAYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPLARGEIVLVG 240
GEFE R+K+I++ ++ + E+++F+DEIHTLVGAG++ GS DA ILK LARGE+ +G
Sbjct 355 GEFEERLKEILSEIENSQGEIVMFIDEIHTLVGAGESQGSLDAGNILKPMLARGELRCIG 414
Query 241 ATTLEEYKLHIEKDAAFCRR 260
ATTL+EY+ IEKD A RR
Sbjct 415 ATTLQEYRQKIEKDKALERR 434
> bbo:BBOV_II004100 18.m06340; ClpB; K03695 ATP-dependent Clp
protease ATP-binding subunit ClpB
Length=931
Score = 192 bits (488), Expect = 1e-48, Method: Compositional matrix adjust.
Identities = 112/264 (42%), Positives = 156/264 (59%), Gaps = 14/264 (5%)
Query 4 LGGAKVTAAVVEVFLRARRMAEAE--SCLVSMQHIFLALLHSPTFYRTLVNSGCDIDLFK 61
G KV ++ L R ++E +S++H+ LAL T + L K
Sbjct 118 FGDQKVLGRTLQNVLTVTRRIKSEYNDHFISVEHLLLALACEDTKFTKPW-------LTK 170
Query 62 EELALAYKNTPMPSTSGSTEHVAKDFD-----LSQYGEDLTEKARLGKLQPVIGRQEEID 116
++ + S G + +K+ + L +Y +DLT AR GKL PVIGR EI
Sbjct 171 HKVGYDKLKRAVESVRGKRKVTSKNPEMLFGVLEKYSKDLTMMARSGKLDPVIGRDNEIR 230
Query 117 RIAQILSRMMTKAPLLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAG 176
R +ILSR P+L+G+PGVGKTA+ EGLA RIV VP +L++ R+ SLDL ++ AG
Sbjct 231 RTVEILSRRTKNNPILLGDPGVGKTAIAEGLANRIVSGDVPDSLKNTRVISLDLASMLAG 290
Query 177 TSMRGEFERRMKDIIAYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPLARGEI 236
+ RGEFE R+K+I+ +Q + E+I+F+DEIHT+VGAG G+ DA ILK LARGE+
Sbjct 291 SQYRGEFEERLKNILKEVQDSQGEIIMFIDEIHTVVGAGDAQGAMDAGNILKPMLARGEL 350
Query 237 VLVGATTLEEYKLHIEKDAAFCRR 260
+GATTL+EY+ IEKD A RR
Sbjct 351 RCIGATTLQEYRQRIEKDKALERR 374
> ath:AT5G50920 CLPC1; CLPC1; ATP binding / ATP-dependent peptidase/
ATPase; K03696 ATP-dependent Clp protease ATP-binding
subunit ClpC
Length=929
Score = 191 bits (484), Expect = 3e-48, Method: Compositional matrix adjust.
Identities = 101/214 (47%), Positives = 137/214 (64%), Gaps = 2/214 (0%)
Query 48 RTLVNSGCDIDLFKEE-LALAYKNTPMPSTSGSTEHVAKDFDLSQYGEDLTEKARLGKLQ 106
R L N G D + + + + +N + + G K L +YG +LT+ A GKL
Sbjct 215 RVLENLGADPSNIRTQVIRMVGENNEVTANVGGGSSSNKMPTLEEYGTNLTKLAEEGKLD 274
Query 107 PVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIF 166
PV+GRQ +I+R+ QIL R P LIGEPGVGKTA+ EGLAQRI VP+ ++ +++
Sbjct 275 PVVGRQPQIERVVQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIASGDVPETIEGKKVI 334
Query 167 SLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQI 226
+LD+ L AGT RGEFE R+K ++ + R E+ILF+DE+HTL+GAG G+ DA I
Sbjct 335 TLDMGLLVAGTKYRGEFEERLKKLMEEI-RQSDEIILFIDEVHTLIGAGAAEGAIDAANI 393
Query 227 LKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
LK LARGE+ +GATTL+EY+ HIEKD A RR
Sbjct 394 LKPALARGELQCIGATTLDEYRKHIEKDPALERR 427
> ath:AT2G25140 CLPB4; CLPB4 (CASEIN LYTIC PROTEINASE B4); ATP
binding / ATPase/ nucleoside-triphosphatase/ nucleotide binding
/ protein binding
Length=964
Score = 190 bits (482), Expect = 5e-48, Method: Compositional matrix adjust.
Identities = 104/231 (45%), Positives = 143/231 (61%), Gaps = 5/231 (2%)
Query 31 VSMQHIFLALLHSPTFYRTLV-NSGCDIDLFKEELALAYKNTPMPSTSGSTEHVAKDFDL 89
VS++H LA F + + DI + K+ + + + + +++ A L
Sbjct 187 VSVEHFLLAYYSDTRFGQEFFRDMKLDIQVLKDAIKDVRGDQRVTDRNPESKYQA----L 242
Query 90 SQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLAQ 149
+YG DLTE AR GKL PVIGR +EI R QIL R P++IGEPGVGKTA+ EGLAQ
Sbjct 243 EKYGNDLTEMARRGKLDPVIGRDDEIRRCIQILCRRTKNNPVIIGEPGVGKTAIAEGLAQ 302
Query 150 RIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEIH 209
RIV VP+ L +R++ SLD+ +L AG RG+FE R+K ++ + + ILF+DEIH
Sbjct 303 RIVRGDVPEPLMNRKLISLDMGSLLAGAKFRGDFEERLKAVMKEVSASNGQTILFIDEIH 362
Query 210 TLVGAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
T+VGAG G+ DA+ +LK L RGE+ +GATTL EY+ +IEKD A RR
Sbjct 363 TVVGAGAMDGAMDASNLLKPMLGRGELRCIGATTLTEYRKYIEKDPALERR 413
> sce:YLL026W HSP104; Heat shock protein that cooperates with
Ydj1p (Hsp40) and Ssa1p (Hsp70) to refold and reactivate previously
denatured, aggregated proteins; responsive to stresses
including: heat, ethanol, and sodium arsenite; involved in
[PSI+] propagation
Length=908
Score = 189 bits (480), Expect = 8e-48, Method: Compositional matrix adjust.
Identities = 99/241 (41%), Positives = 150/241 (62%), Gaps = 6/241 (2%)
Query 20 ARRMAEAESCLVSMQHIFLALLHSPTFYRTLVNSGCDIDLFKEELALAYKNTPMPSTSGS 79
A+ + + ++ HI AL + + + + DI+ K++ AL + + G+
Sbjct 100 AKIQKQQKDSFIAQDHILFALFNDSSIQQIFKEAQVDIEAIKQQ-ALELRGNTRIDSRGA 158
Query 80 TEHVAKDFDLSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVG 139
+ ++ LS+Y D+TE+AR GKL PVIGR+EEI ++L+R + P LIGEPG+G
Sbjct 159 DTNTPLEY-LSKYAIDMTEQARQGKLDPVIGREEEIRSTIRVLARRIKSNPCLIGEPGIG 217
Query 140 KTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKR 199
KTA+IEG+AQRI++ VP LQ ++FSLDL L+AG +G+FE R K ++ ++ K
Sbjct 218 KTAIIEGVAQRIIDDDVPTILQGAKLFSLDLAALTAGAKYKGDFEERFKGVLKEIEESKT 277
Query 200 EVILFVDEIHTLVGAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCR 259
++LF+DEIH L+G GK DA ILK L+RG++ ++GATT EY+ +EKD AF R
Sbjct 278 LIVLFIDEIHMLMGNGK----DDAANILKPALSRGQLKVIGATTNNEYRSIVEKDGAFER 333
Query 260 R 260
R
Sbjct 334 R 334
> ath:AT3G48870 HSP93-III; ATP binding / ATPase/ DNA binding /
nuclease/ nucleoside-triphosphatase/ nucleotide binding / protein
binding; K03696 ATP-dependent Clp protease ATP-binding
subunit ClpC
Length=952
Score = 185 bits (469), Expect = 2e-46, Method: Compositional matrix adjust.
Identities = 92/172 (53%), Positives = 122/172 (70%), Gaps = 1/172 (0%)
Query 89 LSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLA 148
L +YG +LT+ A GKL PV+GRQ +I+R+ QIL+R P LIGEPGVGKTA+ EGLA
Sbjct 278 LEEYGTNLTKLAEEGKLDPVVGRQPQIERVVQILARRTKNNPCLIGEPGVGKTAIAEGLA 337
Query 149 QRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEI 208
QRI VP+ ++ + + +LD+ L AGT RGEFE R+K ++ + R E+ILF+DE+
Sbjct 338 QRIASGDVPETIEGKTVITLDMGLLVAGTKYRGEFEERLKKLMEEI-RQSDEIILFIDEV 396
Query 209 HTLVGAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
HTL+GAG G+ DA ILK LARGE+ +GATT++EY+ HIEKD A RR
Sbjct 397 HTLIGAGAAEGAIDAANILKPALARGELQCIGATTIDEYRKHIEKDPALERR 448
> ath:AT4G14670 CLPB2; CLPB2; ATP binding / nucleoside-triphosphatase/
nucleotide binding / protein binding
Length=623
Score = 185 bits (469), Expect = 2e-46, Method: Compositional matrix adjust.
Identities = 92/172 (53%), Positives = 126/172 (73%), Gaps = 2/172 (1%)
Query 89 LSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLA 148
L YG DL E+A GKL PVIGR EI R+ ++LSR P+LIGEPGVGKTAV+EGLA
Sbjct 129 LKTYGTDLVEQA--GKLDPVIGRHREIRRVIEVLSRRTKNNPVLIGEPGVGKTAVVEGLA 186
Query 149 QRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEI 208
QRI++ VP L ++ SL+ + AGT++RG+FE R+K ++ ++ + +V+LF+DEI
Sbjct 187 QRILKGDVPINLTGVKLISLEFGAMVAGTTLRGQFEERLKSVLKAVEEAQGKVVLFIDEI 246
Query 209 HTLVGAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
H +GA K +GS+DA ++LK LARG++ +GATTLEEY+ H+EKDAAF RR
Sbjct 247 HMALGACKASGSTDAAKLLKPMLARGQLRFIGATTLEEYRTHVEKDAAFERR 298
> tgo:TGME49_057990 heat shock protein, putative (EC:3.4.21.53)
Length=921
Score = 183 bits (465), Expect = 4e-46, Method: Compositional matrix adjust.
Identities = 106/238 (44%), Positives = 147/238 (61%), Gaps = 15/238 (6%)
Query 28 SCLVSMQHIFLALLHSPTFYRTLVNSGCDIDLFKEELALAYKNTPMPSTSGSTEHVAKDF 87
L+S +F AL+ L +G + +E+ S GS + + D
Sbjct 103 DSLMSADSLFSALVQEKGIRSHLTAAGFMMKQIEEK---------AKSVRGSKKIASSDD 153
Query 88 D-----LSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTA 142
D L +YG D T+ A GKL PVIGR++EI R+ +IL R P+LIGEPGVGK+A
Sbjct 154 DANFEALKKYGTDFTDLAEKGKLDPVIGREDEIRRVIRILCRRTKNNPVLIGEPGVGKSA 213
Query 143 VIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVI 202
V+EGLA+RIVE VP L+ R+ SLD+ +L AG RGEFE R+ ++ ++ ++I
Sbjct 214 VVEGLARRIVEHDVPSNLR-CRLVSLDVGSLIAGAKFRGEFEERLTAVLQEVKDAAGKII 272
Query 203 LFVDEIHTLVGAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
LF+DEIH ++GAGKT G+ DA +LK LARGE+ +GATTL+EY+ ++EKDAAF RR
Sbjct 273 LFIDEIHVILGAGKTEGALDAANLLKPMLARGELRCIGATTLDEYRKYVEKDAAFERR 330
> tgo:TGME49_068650 clp ATP-binding chain B1, putative (EC:3.4.21.53)
Length=929
Score = 182 bits (461), Expect = 2e-45, Method: Compositional matrix adjust.
Identities = 100/193 (51%), Positives = 127/193 (65%), Gaps = 11/193 (5%)
Query 70 NTPMPSTSGSTEHVAKDFDLSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKA 129
NT P S + L +YG DLTE A +L PVIGR +E+ R+ QILSR
Sbjct 167 NTKTPEVSYQS--------LKKYGRDLTEAAMANELDPVIGRDKEVRRVIQILSRRTKNN 218
Query 130 PLLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKD 189
P+++G+PGVGKTA+ EGLAQRIV VP L R++ SLDL L AG +RGEFE R+K
Sbjct 219 PIILGDPGVGKTAIAEGLAQRIVSGDVPDTLAGRQLISLDLGALLAGAKLRGEFEERLKS 278
Query 190 IIAYLQRHKREVILFVDEIHTLVGAGKTAGSS--DATQILKVPLARGEIVLVGATTLEEY 247
+I +Q ++ILF+DEIH +VGAG +AG S DA ILK LARGE+ +GATTL+EY
Sbjct 279 VIREVQESSGQIILFIDEIHMVVGAG-SAGESGMDAGNILKPMLARGELRCIGATTLDEY 337
Query 248 KLHIEKDAAFCRR 260
+ +IEKD A RR
Sbjct 338 RKYIEKDKALERR 350
> pfa:PF08_0063 ClpB protein, putative
Length=1070
Score = 179 bits (455), Expect = 7e-45, Method: Composition-based stats.
Identities = 93/173 (53%), Positives = 122/173 (70%), Gaps = 1/173 (0%)
Query 89 LSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLA 148
L +Y DLT AR GKL PVIGR EI R QILSR P+L+G+PGVGKTA++EGLA
Sbjct 314 LEKYSRDLTALARAGKLDPVIGRDNEIRRAIQILSRRTKNNPILLGDPGVGKTAIVEGLA 373
Query 149 QRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEI 208
+IV+ VP +L+ R++ SLD+ +L AG RG+FE R+K I+ +Q + +V++F+DEI
Sbjct 374 IKIVQGDVPDSLKGRKLVSLDMSSLIAGAKYRGDFEERLKSILKEVQDAEGQVVMFIDEI 433
Query 209 HTLVGAGKTA-GSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
HT+VGAG A G+ DA ILK LARGE+ +GATT+ EY+ IEKD A RR
Sbjct 434 HTVVGAGAVAEGALDAGNILKPMLARGELRCIGATTVSEYRQFIEKDKALERR 486
> sce:YDR258C HSP78; Hsp78p
Length=811
Score = 174 bits (442), Expect = 2e-43, Method: Compositional matrix adjust.
Identities = 94/172 (54%), Positives = 121/172 (70%), Gaps = 2/172 (1%)
Query 89 LSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLA 148
L Q+G +LT+ AR GKL PVIGR EEI R QILSR P LIG GVGKTA+I+GLA
Sbjct 98 LEQFGTNLTKLARDGKLDPVIGRDEEIARAIQILSRRTKNNPCLIGRAGVGKTALIDGLA 157
Query 149 QRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEI 208
QRIV VP +L+D+ + +LDL +L AG RGEFE R+K ++ + + +VI+F+DE+
Sbjct 158 QRIVAGEVPDSLKDKDLVALDLGSLIAGAKYRGEFEERLKKVLEEIDKANGKVIVFIDEV 217
Query 209 HTLVGAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
H L+G GKT GS DA+ ILK LARG + + ATTL+E+K+ IEKD A RR
Sbjct 218 HMLLGLGKTDGSMDASNILKPKLARG-LRCISATTLDEFKI-IEKDPALSRR 267
> pfa:PF11_0175 heat shock protein 101, putative
Length=906
Score = 172 bits (436), Expect = 1e-42, Method: Composition-based stats.
Identities = 87/172 (50%), Positives = 116/172 (67%), Gaps = 0/172 (0%)
Query 89 LSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLA 148
+ Q+G ++ EK R GKLQ + GR EEI I + L R +P+L+G PG GKT ++EGL
Sbjct 190 IEQFGSNMNEKVRNGKLQGIYGRDEEIRAIIESLLRYNKNSPVLVGNPGTGKTTIVEGLV 249
Query 149 QRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEI 208
RI + VPK LQ + SL+ ++GTS RGEFE RMK+II L+ K ++ILFVDEI
Sbjct 250 YRIEKGDVPKELQGYTVISLNFRKFTSGTSYRGEFETRMKNIIKELKNKKNKIILFVDEI 309
Query 209 HTLVGAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
H L+GAGK G +DA +LK L++GEI L+GATT+ EY+ IE +AF RR
Sbjct 310 HLLLGAGKAEGGTDAANLLKPVLSKGEIKLIGATTIAEYRKFIESCSAFERR 361
> tgo:TGME49_075690 chaperone clpB 1 protein, putative (EC:3.4.21.53);
K03695 ATP-dependent Clp protease ATP-binding subunit
ClpB
Length=898
Score = 168 bits (425), Expect = 2e-41, Method: Compositional matrix adjust.
Identities = 108/239 (45%), Positives = 146/239 (61%), Gaps = 6/239 (2%)
Query 25 EAESCLVSMQHIFLALLHSPT-FYRTLVNSGCDIDLFKEELALAYKNTPMPSTSGSTEHV 83
E + +S++H+ LAL T F R + G ++ K A+ TS + E
Sbjct 246 EFKDQYLSVEHLVLALAAEDTKFTRPFLTRG-NVSFNKLRSAVEDIRGKKKVTSKNPELA 304
Query 84 AKDFDLSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTAV 143
+ L +Y DLT AR GKL PVIGR +EI R QILSR P+L+G+PGVGKTA+
Sbjct 305 YQA--LERYSRDLTAAARAGKLDPVIGRDDEIRRTIQILSRRTKNNPVLLGDPGVGKTAI 362
Query 144 IEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVIL 203
+EGLAQRI+ VP +L+ RR+ SLD+ L AG RGEFE R+K ++ +Q + +V++
Sbjct 363 VEGLAQRIISGDVPDSLKGRRVISLDMAALIAGAKYRGEFEERLKAVLKEVQDAEGDVVM 422
Query 204 FVDEIHTLV--GAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
F+DEIHT+V GAG G+ DA +LK LARGE +GATT EY+ +IEKD A RR
Sbjct 423 FIDEIHTVVGAGAGGEGGAMDAGNMLKPMLARGEFRCIGATTTNEYRQYIEKDKALERR 481
> bbo:BBOV_III008980 17.m07783; Clp amino terminal domain containing
protein; K01358 ATP-dependent Clp protease, protease
subunit [EC:3.4.21.92]
Length=1005
Score = 157 bits (397), Expect = 4e-38, Method: Compositional matrix adjust.
Identities = 105/266 (39%), Positives = 146/266 (54%), Gaps = 32/266 (12%)
Query 25 EAESC---LVSMQHIFLALL--HSPTFYRTLVNSGCDIDLFKEELALAY-------KNTP 72
EAES + +HI L +L HS TF + L +ID + L L Y
Sbjct 172 EAESMGNKTLETEHILLGMLGTHSYTFRQILDALTVNIDAMRN-LTLKYIAEEKEKPAEE 230
Query 73 MPS--------------TSGSTEHVAKDF----DLSQYGEDLTEKARLGKLQPVIGRQEE 114
MP+ T G +E + +D L+ + D+T KA GKLQ V+ R E
Sbjct 231 MPTDTWKQALSQYTYIPTPGRSEDILRDTYQVSPLNAFTIDITRKAEEGKLQKVLCRDSE 290
Query 115 IDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLS 174
IDR + L R + P+LIGEPGVGKTAV+EG+A ++ E V + + ++R+ LD+ L
Sbjct 291 IDRSIRTLCRKYKRNPILIGEPGVGKTAVVEGIAMQLREGHVLEKMLNKRLRQLDVGLLV 350
Query 175 AGTSMRGEFERRMKDIIAYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPLARG 234
AG RG+FE R+ +I + ++ + +IL +DE H LVGAG G+ DA +LK LARG
Sbjct 351 AGARFRGQFEERLTRLIEEI-KNAKNIILVIDEAHMLVGAGAGEGALDAANLLKPTLARG 409
Query 235 EIVLVGATTLEEYKLHIEKDAAFCRR 260
EI + TT +EY+ H EKDAA CRR
Sbjct 410 EIQCIAITTPKEYQKHFEKDAALCRR 435
> eco:b0882 clpA, ECK0873, JW0866, lopD; ATPase and specificity
subunit of ClpA-ClpP ATP-dependent serine protease, chaperone
activity; K03694 ATP-dependent Clp protease ATP-binding
subunit ClpA
Length=758
Score = 154 bits (388), Expect = 4e-37, Method: Compositional matrix adjust.
Identities = 80/187 (42%), Positives = 120/187 (64%), Gaps = 2/187 (1%)
Query 75 STSGSTEHVAKDFDLSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIG 134
S S E + + + +L + AR+G + P+IGR++E++R Q+L R PLL+G
Sbjct 155 SQPNSEEQAGGEERMENFTTNLNQLARVGGIDPLIGREKELERAIQVLCRRRKNNPLLVG 214
Query 135 EPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYL 194
E GVGKTA+ EGLA RIV+ VP+ + D I+SLD+ +L AGT RG+FE+R K ++ L
Sbjct 215 ESGVGKTAIAEGLAWRIVQGDVPEVMADCTIYSLDIGSLLAGTKYRGDFEKRFKALLKQL 274
Query 195 QRHKREVILFVDEIHTLVGAGKTAGSS-DATQILKVPLARGEIVLVGATTLEEYKLHIEK 253
++ ILF+DEIHT++GAG +G DA ++K L+ G+I ++G+TT +E+ EK
Sbjct 275 EQDTNS-ILFIDEIHTIIGAGAASGGQVDAANLIKPLLSSGKIRVIGSTTYQEFSNIFEK 333
Query 254 DAAFCRR 260
D A RR
Sbjct 334 DRALARR 340
> tpv:TP04_0800 ATP-dependent Clp protease ATP-binding subunit
(EC:3.4.21.92); K01358 ATP-dependent Clp protease, protease
subunit [EC:3.4.21.92]
Length=900
Score = 151 bits (381), Expect = 3e-36, Method: Compositional matrix adjust.
Identities = 81/172 (47%), Positives = 106/172 (61%), Gaps = 1/172 (0%)
Query 89 LSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLA 148
+S + DLTEKAR G+L VI R EI+R LSRM PLL+GEPGVGKTA++EG+A
Sbjct 250 ISMFTVDLTEKARNGQLPKVIHRDNEIERAIITLSRMTKSNPLLVGEPGVGKTAIVEGIA 309
Query 149 QRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEI 208
RI + + +RI L L AGT RG+FE R+ +I + + ++IL +DE
Sbjct 310 NRISQGISQPQISKKRILQLQFGLLIAGTKFRGQFEERLTKLIDEI-KSAGDIILVIDEA 368
Query 209 HTLVGAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
H L+G G GS DA +LK PL+RGEI + TT +EYK + EKD A RR
Sbjct 369 HMLIGGGAGDGSIDAANLLKPPLSRGEIQCIAITTPKEYKKYFEKDMALSRR 420
> ath:AT5G51070 ERD1; ERD1 (EARLY RESPONSIVE TO DEHYDRATION 1);
ATP binding / ATPase/ nucleoside-triphosphatase/ nucleotide
binding / protein binding
Length=945
Score = 144 bits (362), Expect = 4e-34, Method: Compositional matrix adjust.
Identities = 84/189 (44%), Positives = 116/189 (61%), Gaps = 7/189 (3%)
Query 77 SGSTEHVAKDFDLSQYGEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEP 136
SG AK+ L Q+ DLT +A G + PVIGR++E+ R+ QIL R P+L+GE
Sbjct 260 SGPGGKKAKNV-LEQFCVDLTARASEGLIDPVIGREKEVQRVIQILCRRTKNNPILLGEA 318
Query 137 GVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQR 196
GVGKTA+ EGLA I E + P L +RI SLD+ L AG RGE E R+ +I+ +++
Sbjct 319 GVGKTAIAEGLAISIAEASAPGFLLTKRIMSLDIGLLMAGAKERGELEARVTALISEVKK 378
Query 197 HKREVILFVDEIHTLVGAGKTA----GSS-DATQILKVPLARGEIVLVGATTLEEYKLHI 251
+ VILF+DE+HTL+G+G GS D +LK L RGE+ + +TTL+E++
Sbjct 379 SGK-VILFIDEVHTLIGSGTVGRGNKGSGLDIANLLKPSLGRGELQCIASTTLDEFRSQF 437
Query 252 EKDAAFCRR 260
EKD A RR
Sbjct 438 EKDKALARR 446
> pfa:PF14_0063 ATP-dependent CLP protease, putative; K03695 ATP-dependent
Clp protease ATP-binding subunit ClpB
Length=1341
Score = 142 bits (358), Expect = 1e-33, Method: Composition-based stats.
Identities = 76/166 (45%), Positives = 105/166 (63%), Gaps = 1/166 (0%)
Query 95 DLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLAQRIVER 154
D+ +A+ GR++EI RI +IL R PLLIGE GVGKTA+IE L+ I++
Sbjct 512 DMVHEAQEKGDDHFFGRKKEIKRIIEILGRKKKSNPLLIGESGVGKTAIIEYLSYLILKD 571
Query 155 AVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEIHTLVGA 214
VP L++ RIF L+L N+ AGT RGEFE +MK +++ + + K+ ILF+DEIH +VGA
Sbjct 572 NVPYHLKNCRIFQLNLGNIVAGTKYRGEFEEKMKHLLSNMNKKKKN-ILFIDEIHVIVGA 630
Query 215 GKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
G GS DA+ +LK L+ + +G TT +EY IE D A RR
Sbjct 631 GSGEGSLDASNLLKPFLSSDNLQCIGTTTFQEYSKFIENDKALRRR 676
> ath:AT3G45450 Clp amino terminal domain-containing protein
Length=341
Score = 107 bits (267), Expect = 4e-23, Method: Compositional matrix adjust.
Identities = 65/161 (40%), Positives = 90/161 (55%), Gaps = 18/161 (11%)
Query 101 RLGKLQPVIGRQEEIDRIAQILSRMMTKA-PLLIGEPGVGKTAVIEGLAQRIVERAVPKA 159
R GKL PV+GRQ +I R+ QIL+R + LIG+PGVGK A+ EG+AQRI VP+
Sbjct 149 RRGKLDPVVGRQPQIKRVVQILARRTCRNNACLIGKPGVGKRAIAEGIAQRIASGDVPET 208
Query 160 LQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEIHTLVGAGKTAG 219
++ + +N+ AG E R + I + ++ILF+DE+H L+GAG G
Sbjct 209 IKGK-------MNV-AGNCGWNEIRWRSRGKIEEVYGQSDDIILFIDEMHLLIGAGAVEG 260
Query 220 SSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
+ DA ILK L R E+ +Y+ HIE D A RR
Sbjct 261 AIDAANILKPALERCEL---------QYRKHIENDPALERR 292
> bbo:BBOV_I001700 19.m02115; chaperone clpB
Length=833
Score = 103 bits (256), Expect = 8e-22, Method: Compositional matrix adjust.
Identities = 65/168 (38%), Positives = 95/168 (56%), Gaps = 13/168 (7%)
Query 95 DLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLAQRIVER 154
+LTE A+ G +GR+ E++R+ L+RM LLIGEPGVGKTA++E L
Sbjct 168 NLTEAAKNGSGNVFVGRENELERLKGSLNRMRKNNVLLIGEPGVGKTALVERL------- 220
Query 155 AVPKALQDRRI--FSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEIHTLV 212
AV L+D I +SLDL L +G RGE E ++K I ++ K ILF+DEIH L+
Sbjct 221 AVDMLLEDPNITVYSLDLCRLYSGQGTRGELEAKLKSIFDTVKNGKS--ILFIDEIHHLI 278
Query 213 GAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYKLHIEKDAAFCRR 260
+ + T +LK + + ++G+TT +EY + +D AF RR
Sbjct 279 QNQENG--VNVTNLLKPIMTSTLVKIIGSTTAKEYHQYFRRDRAFERR 324
> sce:YNL218W MGS1; Mgs1p; K07478 putative ATPase
Length=587
Score = 52.0 bits (123), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 47/157 (29%), Positives = 77/157 (49%), Gaps = 26/157 (16%)
Query 96 LTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPL----LIGEPGVGKTAVIEGLAQRI 151
L+EK R +L+ +G+Q + + L + + + + L G PGVGKT+ LA+ +
Sbjct 135 LSEKLRPKELRDYVGQQHILSQDNGTLFKYIKQGTIPSMILWGPPGVGKTS----LARLL 190
Query 152 VERAVPKALQDR---RIFSLDLLNLSAGTS-MRGEFERRMKDIIAYLQRHKREVILFVDE 207
+ A + + R F ++ A T +RG FE+ K+ Q KR +LF+DE
Sbjct 191 TKTATTSSNESNVGSRYFMIETSATKANTQELRGIFEKSKKE----YQLTKRRTVLFIDE 246
Query 208 IHTLVGAGKTAGSSDATQILKVP-LARGEIVLVGATT 243
IH + Q L +P + G+I+L+GATT
Sbjct 247 IHRF---------NKVQQDLLLPHVENGDIILIGATT 274
> hsa:23560 GTPBP4, CRFG, FLJ10686, FLJ10690, FLJ39774, NGB, NOG1;
GTP binding protein 4; K06943 nucleolar GTP-binding protein
Length=634
Score = 46.2 bits (108), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 45/136 (33%), Positives = 68/136 (50%), Gaps = 22/136 (16%)
Query 46 FYRTLVNSGCDIDLFKEELALAYKNTPMPSTSGSTEHVAKDF-DLSQYGEDLT-----EK 99
FY L+N D D +K LAL N ++VAKD+ L +YG+ L ++
Sbjct 78 FYADLMNILYDKDHYK--LALGQINI----AKNLVDNVAKDYVRLMKYGDSLYRCKQLKR 131
Query 100 ARLGKLQPVIGRQ----EEIDRIAQILSRMMTKAP-----LLIGEPGVGKTAVIEGLAQR 150
A LG++ VI RQ E ++++ Q LSR+ T P LL G P VGK++ I + +
Sbjct 132 AALGRMCTVIKRQKQSLEYLEQVRQHLSRLPTIDPNTRTLLLCGYPNVGKSSFINKVTRA 191
Query 151 IVERAVPKALQDRRIF 166
V+ P A + +F
Sbjct 192 DVD-VQPYAFTTKSLF 206
> dre:334050 gtpbp4, fi28d07, wu:fi28d07, zgc:55757; GTP binding
protein 4; K06943 nucleolar GTP-binding protein
Length=631
Score = 45.8 bits (107), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 44/136 (32%), Positives = 68/136 (50%), Gaps = 22/136 (16%)
Query 46 FYRTLVNSGCDIDLFKEELALAYKNTPMPSTSGSTEHVAKDF-DLSQYGEDLT-----EK 99
FY L+N D D +K LAL N ++VAKD+ L +YG+ L ++
Sbjct 78 FYADLMNVLYDKDHYK--LALGQINI----AKNLIDNVAKDYVRLMKYGDSLYRCKQLKR 131
Query 100 ARLGKLQPVIGRQ----EEIDRIAQILSRMMTKAP-----LLIGEPGVGKTAVIEGLAQR 150
A LG++ ++ RQ E ++++ Q LSR+ T P LL G P VGK++ I + +
Sbjct 132 AALGRMCTILKRQKQSLEYLEQVRQHLSRLPTIDPNTRTLLLCGYPNVGKSSFINKVTRA 191
Query 151 IVERAVPKALQDRRIF 166
VE P A + +F
Sbjct 192 DVE-VQPYAFTTKSLF 206
> mmu:69237 Gtpbp4, 2610028C09Rik, Crfg, Gtpbp3, NGB, Nog1; GTP
binding protein 4; K06943 nucleolar GTP-binding protein
Length=634
Score = 45.4 bits (106), Expect = 2e-04, Method: Compositional matrix adjust.
Identities = 44/136 (32%), Positives = 68/136 (50%), Gaps = 22/136 (16%)
Query 46 FYRTLVNSGCDIDLFKEELALAYKNTPMPSTSGSTEHVAKDF-DLSQYGEDLT-----EK 99
FY L+N D D +K LAL N ++VAKD+ L +YG+ L ++
Sbjct 78 FYADLMNILYDKDHYK--LALGQINI----AKNLVDNVAKDYVRLMKYGDSLYRCKQLKR 131
Query 100 ARLGKLQPVIGRQ----EEIDRIAQILSRMMTKAP-----LLIGEPGVGKTAVIEGLAQR 150
A LG++ +I RQ E ++++ Q LSR+ T P LL G P VGK++ I + +
Sbjct 132 AALGRMCTIIKRQRQSLEYLEQVRQHLSRLPTIDPNTRTLLLCGYPNVGKSSFINKVTRA 191
Query 151 IVERAVPKALQDRRIF 166
V+ P A + +F
Sbjct 192 DVD-VQPYAFTTKSLF 206
> cpv:cgd7_2620 ClpB ATpase (bacterial), signal peptide
Length=1263
Score = 42.7 bits (99), Expect = 0.001, Method: Composition-based stats.
Identities = 49/213 (23%), Positives = 93/213 (43%), Gaps = 14/213 (6%)
Query 54 GCDIDLFKEELALAYKNTPMPSTSGSTEHVA--KDFDLSQYGEDLTEKARLGKLQPVI-- 109
G D F E YK +PST G ++ ++ ++ Y L +K + + VI
Sbjct 417 GIDYFKFMNEALKKYKTFFVPSTDGRINSISWFNNYSVNGYTNKLNKKWFVDINELVIEN 476
Query 110 GRQEEIDRIAQI------LSRMMTKAPLLIGEPGVGKTAVIEGLAQRIVERAVPKALQDR 163
G + +D + + LSR + +++ E + K ++E LA RI+ L+
Sbjct 477 GVTKAVDFMDEFHLLEISLSRSGLSSAIIVSESKLLKRTLVEYLAYRIISGNSSIDLRGY 536
Query 164 RIFSLDLLNL--SAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEIHTLVGAGKTAGSS 221
RI S+ L +L S + + E+ + + ++I+F D + + + GS
Sbjct 537 RIISIHLESLLESCKNTKKSLTEQIKIKFDELMGAYDGKIIVFTDNLFS--SFETSTGSK 594
Query 222 DATQILKVPLARGEIVLVGATTLEEYKLHIEKD 254
I+K + RG + ++ + E YK+ EK+
Sbjct 595 RLYDIMKHYIVRGTLKVIATLSNENYKILAEKE 627
> mmu:78903 Wrnip1, 4833444L21Rik, WHIP, Wrnip; Werner helicase
interacting protein 1
Length=660
Score = 42.4 bits (98), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 42/159 (26%), Positives = 69/159 (43%), Gaps = 33/159 (20%)
Query 93 GEDLTEKARLGKLQPVIGRQEEIDR---IAQILSRMMTKAPLLIGEPGVGKTAVIEGLAQ 149
G+ L +K R LQ IG+ + + + +L + +L G PG GKT + +A
Sbjct 219 GKPLADKMRPDTLQDYIGQSRAVGQETLLRSLLEANEIPSLILWGPPGCGKTTLAHIIAN 278
Query 150 RIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHK----REVILFV 205
+ + S+ + LSA + + ++D+I Q K R+ ILF+
Sbjct 279 ------------NSKKHSIRFVTLSATNAKTND----VRDVIKQAQNEKSFFKRKTILFI 322
Query 206 DEIHTLVGAGKTAGSSDATQILKVP-LARGEIVLVGATT 243
DEIH + + Q +P + G I L+GATT
Sbjct 323 DEIHRF---------NKSQQDTFLPHVECGTITLIGATT 352
> pfa:PF14_0548 ATPase, putative; K12196 vacuolar protein-sorting-associated
protein 4
Length=419
Score = 42.0 bits (97), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 34/133 (25%), Positives = 61/133 (45%), Gaps = 29/133 (21%)
Query 123 SRMMTKAPLLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAG---TSM 179
S + K LL G PG GKT + AL +++ N+S+ +
Sbjct 143 STLPYKGILLYGPPGTGKTFL---------------ALACSNECNMNFFNVSSSDLVSKY 187
Query 180 RGEFERRMKDIIAYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPL-------- 231
+GE E+ +K + + H I+F+DEI +L G+ +T G +++T+ +K
Sbjct 188 QGESEKYIKCLFETAKEH-SPAIIFIDEIDSLCGS-RTDGENESTRRIKTEFLINMSGLT 245
Query 232 -ARGEIVLVGATT 243
+ I+++GAT
Sbjct 246 NYKNNIIVMGATN 258
> cel:Y34D9A.10 vps-4; related to yeast Vacuolar Protein Sorting
factor family member (vps-4); K12196 vacuolar protein-sorting-associated
protein 4
Length=430
Score = 40.8 bits (94), Expect = 0.005, Method: Compositional matrix adjust.
Identities = 36/122 (29%), Positives = 55/122 (45%), Gaps = 20/122 (16%)
Query 131 LLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDI 190
LL G PG GK+ + + +A E I S DL+ + GE E+ +K++
Sbjct 155 LLFGPPGTGKSYIAKAVATEAGESTF------FSISSSDLM-----SKWLGESEKLVKNL 203
Query 191 IAYLQRHKREVILFVDEIHTLVGAGKTAGSSDA--------TQILKVPLARGEIVLVGAT 242
A + HK +I F+DEI +L A S A Q+ V L I+++GAT
Sbjct 204 FALAREHKPSII-FIDEIDSLCSARSDNESESARRIKTEFMVQMQGVGLNNDGILVLGAT 262
Query 243 TL 244
+
Sbjct 263 NI 264
> dre:100331225 valosin-containing protein-like
Length=739
Score = 40.4 bits (93), Expect = 0.006, Method: Compositional matrix adjust.
Identities = 39/136 (28%), Positives = 56/136 (41%), Gaps = 22/136 (16%)
Query 120 QILSRMMTKAP---LLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAG 176
Q+ + + P L G PG GKT V LA + DR++
Sbjct 404 QVFEKFKIQPPRGCLFYGPPGTGKTLVARALANECSQ-------GDRKVSFFMRKGADCL 456
Query 177 TSMRGEFERRMKDII--AYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPLA-- 232
+ GE ER+++ + AYL R I+F DEI L + + I+ LA
Sbjct 457 SKWVGESERQLRLLFDQAYLM---RPSIIFFDEIDGLAPVRSSRQDQIHSSIVSTLLALM 513
Query 233 -----RGEIVLVGATT 243
RGEIV++GAT
Sbjct 514 DGLDSRGEIVVIGATN 529
> ath:AT1G05910 cell division cycle protein 48-related / CDC48-related
Length=1210
Score = 40.0 bits (92), Expect = 0.009, Method: Compositional matrix adjust.
Identities = 39/123 (31%), Positives = 54/123 (43%), Gaps = 21/123 (17%)
Query 131 LLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIF---SLDLLNLSAGTSMRGEFERRM 187
LL G PG GKT + LA A KA Q + D+L + GE ER++
Sbjct 419 LLCGPPGTGKTLIARALAC-----AASKAGQKVSFYMRKGADVL-----SKWVGEAERQL 468
Query 188 KDIIAYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPLA-------RGEIVLVG 240
K + QR++ +I F DEI L + I+ LA RG++VL+G
Sbjct 469 KLLFEEAQRNQPSIIFF-DEIDGLAPVRSSKQEQIHNSIVSTLLALMDGLDSRGQVVLIG 527
Query 241 ATT 243
AT
Sbjct 528 ATN 530
> hsa:56897 WRNIP1, FLJ22526, RP11-420G6.2, WHIP, bA420G6.2; Werner
helicase interacting protein 1
Length=665
Score = 39.3 bits (90), Expect = 0.015, Method: Compositional matrix adjust.
Identities = 41/159 (25%), Positives = 69/159 (43%), Gaps = 33/159 (20%)
Query 93 GEDLTEKARLGKLQPVIGRQEEIDRIAQILSRMMTK---APLLIGEPGVGKTAVIEGLAQ 149
G+ L + R LQ G+ + + + + S + T + +L G PG GKT + +A
Sbjct 224 GKPLADTMRPDTLQDYFGQSKAVGQDTLLRSLLETNEIPSLILWGPPGCGKTTLAHIIAS 283
Query 150 RIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHK----REVILFV 205
+ + S+ + LSA + + ++D+I Q K R+ ILF+
Sbjct 284 ------------NSKKHSIRFVTLSATNAKTND----VRDVIKQAQNEKSFFKRKTILFI 327
Query 206 DEIHTLVGAGKTAGSSDATQILKVP-LARGEIVLVGATT 243
DEIH + + Q +P + G I L+GATT
Sbjct 328 DEIHRF---------NKSQQDTFLPHVECGTITLIGATT 357
> ath:AT2G27600 SKD1; SKD1 (SUPPRESSOR OF K+ TRANSPORT GROWTH
DEFECT1); ATP binding / nucleoside-triphosphatase/ nucleotide
binding; K12196 vacuolar protein-sorting-associated protein
4
Length=435
Score = 38.5 bits (88), Expect = 0.023, Method: Compositional matrix adjust.
Identities = 42/145 (28%), Positives = 61/145 (42%), Gaps = 25/145 (17%)
Query 124 RMMTKAPLLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEF 183
R +A LL G PG GK+ + + +A D FS+ +L + GE
Sbjct 162 RRPWRAFLLYGPPGTGKSYLAKAVATEA----------DSTFFSVSSSDLV--SKWMGES 209
Query 184 ERRMKDIIAYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPL--------ARGE 235
E+ + ++ + R I+FVDEI +L G S+A++ +K L E
Sbjct 210 EKLVSNLFE-MARESAPSIIFVDEIDSLCGTRGEGNESEASRRIKTELLVQMQGVGHNDE 268
Query 236 IVLVGATTLEEYKLHIEKDAAFCRR 260
VLV A T Y L D A RR
Sbjct 269 KVLVLAATNTPYAL----DQAIRRR 289
> sce:YOL094C RFC4; Rfc4p; K10755 replication factor C subunit
2/4
Length=323
Score = 38.5 bits (88), Expect = 0.027, Method: Compositional matrix adjust.
Identities = 20/58 (34%), Positives = 31/58 (53%), Gaps = 0/58 (0%)
Query 98 EKARLGKLQPVIGRQEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIEGLAQRIVERA 155
EK R L ++G +E IDR+ QI ++ G PG+GKT + LA ++ R+
Sbjct 13 EKYRPQVLSDIVGNKETIDRLQQIAKDGNMPHMIISGMPGIGKTTSVHCLAHELLGRS 70
> dre:405856 MGC85976; zgc:85976; K07478 putative ATPase
Length=546
Score = 38.5 bits (88), Expect = 0.027, Method: Compositional matrix adjust.
Identities = 43/154 (27%), Positives = 67/154 (43%), Gaps = 28/154 (18%)
Query 96 LTEKARLGKLQPVIGRQEEIDRIAQILSRMMTKA---PLLI--GEPGVGKTAVIEGLAQR 150
L E R L+ G+ + I Q L R + K+ P LI G PG GKT + +A
Sbjct 117 LAELLRPSTLEEYFGQNKLIGE--QTLLRSLLKSQEIPSLILWGPPGCGKTTLAHIIASS 174
Query 151 IVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEIHT 210
I ++ + + LSA ++ + +K L+ KR+ +LF+DEIH
Sbjct 175 IKQKGTGR-----------FVTLSATSASVSDVREVIKQAQNELRLCKRKTVLFIDEIHR 223
Query 211 LVGAGKTAGSSDATQILKVP-LARGEIVLVGATT 243
+ + Q +P + G I L+GATT
Sbjct 224 F---------NKSQQDTFLPHVECGTITLIGATT 248
> hsa:54454 ATAD2B, KIAA1240, MGC88424; ATPase family, AAA domain
containing 2B
Length=1458
Score = 38.5 bits (88), Expect = 0.027, Method: Compositional matrix adjust.
Identities = 38/136 (27%), Positives = 56/136 (41%), Gaps = 22/136 (16%)
Query 120 QILSRMMTKAP---LLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAG 176
+I + + P L G PG GKT V LA + D+++
Sbjct 424 EIFEKFKIQPPRGCLFYGPPGTGKTLVARALANECSQ-------GDKKVAFFMRKGADCL 476
Query 177 TSMRGEFERRMKDII--AYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPLA-- 232
+ GE ER+++ + AYL R I+F DEI L + + I+ LA
Sbjct 477 SKWVGESERQLRLLFDQAYLM---RPSIIFFDEIDGLAPVRSSRQDQIHSSIVSTLLALM 533
Query 233 -----RGEIVLVGATT 243
RGEIV++GAT
Sbjct 534 DGLDNRGEIVVIGATN 549
> cpv:cgd2_1550 origin recognition complex 4 orc4p like AAA+ ATpase
Length=495
Score = 38.5 bits (88), Expect = 0.028, Method: Compositional matrix adjust.
Identities = 36/156 (23%), Positives = 66/156 (42%), Gaps = 11/156 (7%)
Query 94 EDLTEKARLGKLQPVIGRQEEIDRIA-QILSRMMTKAPLLIGEPGVGKTAVIEGLAQRIV 152
E+LTE+ ++ + + +++ I QI+ +++ +LIG+P GKT ++ +
Sbjct 35 ENLTEEHQVSRCDNSLVIPNQLENIFDQIMINSTSRSCILIGKPCAGKTKLLSSYFNKKK 94
Query 153 ERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDIIAYLQRHKREVILFVDEIHTLV 212
V L I L+ LN T + ER I Y H+R LF +
Sbjct 95 SENVSNEL---IIIHLNCLNYDDNTLLSALLER----INEYFPMHRR---LFSGHQKISI 144
Query 213 GAGKTAGSSDATQILKVPLARGEIVLVGATTLEEYK 248
K +G S + L E +++GA+ + +
Sbjct 145 LKEKLSGLSKCGYTIVFALDNCEPIIIGASNISYFN 180
> mmu:320817 Atad2b, 1110014E10Rik, BC032887, C79189, D530031C13Rik,
KIAA1240; ATPase family, AAA domain containing 2B
Length=1460
Score = 38.1 bits (87), Expect = 0.029, Method: Compositional matrix adjust.
Identities = 38/136 (27%), Positives = 56/136 (41%), Gaps = 22/136 (16%)
Query 120 QILSRMMTKAP---LLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAG 176
+I + + P L G PG GKT V LA + D+++
Sbjct 423 EIFEKFKIQPPRGCLFYGPPGTGKTLVARALANECSQ-------GDKKVAFFMRKGADCL 475
Query 177 TSMRGEFERRMKDII--AYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPLA-- 232
+ GE ER+++ + AYL R I+F DEI L + + I+ LA
Sbjct 476 SKWVGESERQLRLLFDQAYLM---RPSIIFFDEIDGLAPVRSSRQDQIHSSIVSTLLALM 532
Query 233 -----RGEIVLVGATT 243
RGEIV++GAT
Sbjct 533 DGLDNRGEIVVIGATN 548
> mmu:70472 Atad2, 2610509G12Rik, MGC38189; ATPase family, AAA
domain containing 2 (EC:3.6.1.3)
Length=1364
Score = 38.1 bits (87), Expect = 0.030, Method: Compositional matrix adjust.
Identities = 35/121 (28%), Positives = 52/121 (42%), Gaps = 17/121 (14%)
Query 131 LLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDI 190
L G PG GKT V LA + D+R+ + GE ER+++
Sbjct 443 LFYGPPGTGKTLVARALANEC-------SRGDKRVAFFMRKGADCLSKWVGESERQLR-- 493
Query 191 IAYLQRHK-REVILFVDEIHTLVGAGKTAGSSDATQILKVPLA-------RGEIVLVGAT 242
+ + Q ++ R I+F DEI L + + I+ LA RGEIV++GAT
Sbjct 494 LLFDQAYQMRPAIIFFDEIDGLAPVRSSRQDQIHSSIVSTLLALMDGLDSRGEIVVIGAT 553
Query 243 T 243
Sbjct 554 N 554
> sce:YPR173C VPS4, CSC1, DID6, END13, GRD13, VPL4, VPT10; AAA-ATPase
involved in multivesicular body (MVB) protein sorting,
ATP-bound Vps4p localizes to endosomes and catalyzes ESCRT-III
disassembly and membrane release; ATPase activity is activated
by Vta1p; regulates cellular sterol metabolism; K12196
vacuolar protein-sorting-associated protein 4
Length=437
Score = 38.1 bits (87), Expect = 0.032, Method: Compositional matrix adjust.
Identities = 41/148 (27%), Positives = 70/148 (47%), Gaps = 30/148 (20%)
Query 123 SRMMTKAPLLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGE 182
+R T LL G PG GK+ + + +A + FS+ +L + GE
Sbjct 162 NRKPTSGILLYGPPGTGKSYLAKAVATEA----------NSTFFSVSSSDLV--SKWMGE 209
Query 183 FERRMKDIIAYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPL----------A 232
E+ +K + A + R + I+F+DE+ L G + G S+A++ +K L +
Sbjct 210 SEKLVKQLFA-MARENKPSIIFIDEVDALTGT-RGEGESEASRRIKTELLVQMNGVGNDS 267
Query 233 RGEIVLVGATTLEEYKLHIEKDAAFCRR 260
+G +VL GAT + ++L D+A RR
Sbjct 268 QGVLVL-GATNI-PWQL----DSAIRRR 289
> mmu:20479 Vps4b, 8030489C12Rik, Skd1; vacuolar protein sorting
4b (yeast); K12196 vacuolar protein-sorting-associated protein
4
Length=444
Score = 38.1 bits (87), Expect = 0.035, Method: Compositional matrix adjust.
Identities = 34/130 (26%), Positives = 61/130 (46%), Gaps = 22/130 (16%)
Query 124 RMMTKAPLLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEF 183
R + LL G PG GK+ + +AV + FS+ +L + GE
Sbjct 164 RTPWRGILLFGPPGTGKS---------YLAKAVATEANNSTFFSISSSDLV--SKWLGES 212
Query 184 ERRMKDIIAYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILK---------VPLARG 234
E+ +K++ L R + I+F+DEI +L G+ ++ S+A + +K V +
Sbjct 213 EKLVKNLFQ-LARENKPSIIFIDEIDSLCGS-RSENESEAARRIKTEFLVQMQGVGVDND 270
Query 235 EIVLVGATTL 244
I+++GAT +
Sbjct 271 GILVLGATNI 280
> hsa:29028 ATAD2, ANCCA, DKFZp667N1320, MGC131938, MGC142216,
MGC29843, MGC5254, PRO2000; ATPase family, AAA domain containing
2 (EC:3.6.1.3)
Length=1390
Score = 37.7 bits (86), Expect = 0.041, Method: Compositional matrix adjust.
Identities = 35/121 (28%), Positives = 52/121 (42%), Gaps = 17/121 (14%)
Query 131 LLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDI 190
L G PG GKT V LA + D+R+ + GE ER+++
Sbjct 464 LFYGPPGTGKTLVARALANECSQ-------GDKRVAFFMRKGADCLSKWVGESERQLR-- 514
Query 191 IAYLQRHK-REVILFVDEIHTLVGAGKTAGSSDATQILKVPLA-------RGEIVLVGAT 242
+ + Q ++ R I+F DEI L + + I+ LA RGEIV++GAT
Sbjct 515 LLFDQAYQMRPSIIFFDEIDGLAPVRSSRQDQIHSSIVSTLLALMDGLDSRGEIVVIGAT 574
Query 243 T 243
Sbjct 575 N 575
> xla:100158428 atad2b; ATPase family, AAA domain containing 2B
Length=872
Score = 37.7 bits (86), Expect = 0.042, Method: Compositional matrix adjust.
Identities = 37/136 (27%), Positives = 56/136 (41%), Gaps = 22/136 (16%)
Query 120 QILSRMMTKAP---LLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAG 176
+I + + P L G PG GKT V LA + D+++
Sbjct 393 EIFEKFRIQPPRGCLFYGPPGTGKTLVARALANECSQ-------GDKKVSFFMRKGADCL 445
Query 177 TSMRGEFERRMKDII--AYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILKVPLA-- 232
+ GE ER+++ + AY+ R I+F DEI L + + I+ LA
Sbjct 446 SKWVGESERQLRLLFDQAYVM---RPSIIFFDEIDGLAPVRSSRQDQIHSSIVSTLLALM 502
Query 233 -----RGEIVLVGATT 243
RGEIV++GAT
Sbjct 503 DGLDNRGEIVVIGATN 518
> xla:379801 vps4b, MGC53483; vacuolar protein sorting 4 homolog
B; K12196 vacuolar protein-sorting-associated protein 4
Length=442
Score = 37.7 bits (86), Expect = 0.043, Method: Compositional matrix adjust.
Identities = 33/123 (26%), Positives = 59/123 (47%), Gaps = 22/123 (17%)
Query 131 LLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDI 190
LL G PG GK+ + +AV + FS+ +L + GE E+ +K++
Sbjct 169 LLFGPPGTGKS---------YLAKAVATEANNSTFFSISSSDLV--SKWLGESEKLVKNL 217
Query 191 IAYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILK---------VPLARGEIVLVGA 241
+ HK +I F+DEI +L G+ ++ S+A + +K V + I+++GA
Sbjct 218 FQLAREHKPSII-FIDEIDSLCGS-RSENESEAARRIKTEFLVQMQGVGVDNEGILVLGA 275
Query 242 TTL 244
T +
Sbjct 276 TNI 278
> hsa:9525 VPS4B, SKD1, SKD1B, VPS4-2; vacuolar protein sorting
4 homolog B (S. cerevisiae); K12196 vacuolar protein-sorting-associated
protein 4
Length=444
Score = 37.7 bits (86), Expect = 0.044, Method: Compositional matrix adjust.
Identities = 33/123 (26%), Positives = 59/123 (47%), Gaps = 22/123 (17%)
Query 131 LLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDI 190
LL G PG GK+ + +AV + FS+ +L + GE E+ +K++
Sbjct 171 LLFGPPGTGKS---------YLAKAVATEANNSTFFSISSSDLV--SKWLGESEKLVKNL 219
Query 191 IAYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILK---------VPLARGEIVLVGA 241
L R + I+F+DEI +L G+ ++ S+A + +K V + I+++GA
Sbjct 220 FQ-LARENKPSIIFIDEIDSLCGS-RSENESEAARRIKTEFLVQMQGVGVDNDGILVLGA 277
Query 242 TTL 244
T +
Sbjct 278 TNI 280
> xla:444796 MGC82073 protein; K12196 vacuolar protein-sorting-associated
protein 4
Length=443
Score = 37.7 bits (86), Expect = 0.046, Method: Compositional matrix adjust.
Identities = 28/99 (28%), Positives = 49/99 (49%), Gaps = 13/99 (13%)
Query 131 LLIGEPGVGKTAVIEGLAQRIVERAVPKALQDRRIFSLDLLNLSAGTSMRGEFERRMKDI 190
LL G PG GK+ + +AV + FS+ +L + GE E+ +K++
Sbjct 170 LLFGPPGTGKS---------YLAKAVATEANNSTFFSISSSDLV--SKWLGESEKLVKNL 218
Query 191 IAYLQRHKREVILFVDEIHTLVGAGKTAGSSDATQILKV 229
+ HK +I F+DEI +L G+ ++ S+A + +K
Sbjct 219 FQLAREHKPSII-FIDEIDSLCGS-RSENESEAARRIKT 255
Lambda K H
0.321 0.136 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 9596524200
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
Posted date: Sep 17, 2011 2:57 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40