bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
164,496 sequences; 82,071,388 total letters
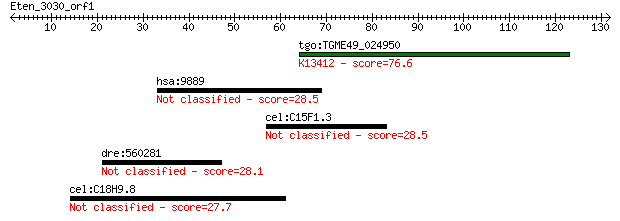
Query= Eten_3030_orf1
Length=131
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_024950 protein kinase 6, putative (EC:1.6.3.1 2.7.1... 76.6 2e-14
hsa:9889 ZBED4; zinc finger, BED-type containing 4 28.5
cel:C15F1.3 tra-2; TRAnsformer : XX animals transformed into m... 28.5 5.8
dre:560281 flii, im:7141769; flightless I homolog (Drosophila) 28.1 6.7
cel:C18H9.8 ift-74; IFT (Chlamydomonas IntraFlagellar Transpor... 27.7 9.6
> tgo:TGME49_024950 protein kinase 6, putative (EC:1.6.3.1 2.7.11.17);
K13412 calcium-dependent protein kinase [EC:2.7.11.1]
Length=681
Score = 76.6 bits (187), Expect = 2e-14, Method: Compositional matrix adjust.
Identities = 38/67 (56%), Positives = 45/67 (67%), Gaps = 8/67 (11%)
Query 64 ADHHCHG-----PEEKPHFFSLCTKCATDGKDKPL---REPAKGFITRSESYVNASAASR 115
AD+ C G KPHFFSLCTKCA +GKDK REP +G I RSES++NAS ASR
Sbjct 92 ADYGCRGGDCCDTSTKPHFFSLCTKCANEGKDKSQHDRREPVRGLIIRSESFMNASEASR 151
Query 116 WEVDELC 122
WE++ C
Sbjct 152 WEIEAQC 158
> hsa:9889 ZBED4; zinc finger, BED-type containing 4
Length=1171
Score = 28.5 bits (62), Expect = 5.3, Method: Compositional matrix adjust.
Identities = 15/36 (41%), Positives = 17/36 (47%), Gaps = 1/36 (2%)
Query 33 GGPPASNMQKDKRLDTAEGRPLESPCRPNCAADHHC 68
G SN ++ TA ESP RP C DHHC
Sbjct 732 SGIWMSNQTREYLTLTAHWVSFESPARPRC-DDHHC 766
> cel:C15F1.3 tra-2; TRAnsformer : XX animals transformed into
males family member (tra-2)
Length=1475
Score = 28.5 bits (62), Expect = 5.8, Method: Composition-based stats.
Identities = 11/27 (40%), Positives = 16/27 (59%), Gaps = 1/27 (3%)
Query 57 PCRPNCAAD-HHCHGPEEKPHFFSLCT 82
P + CAA H C P + H+F++CT
Sbjct 238 PRQKTCAASIHSCDTPLDSEHYFNICT 264
> dre:560281 flii, im:7141769; flightless I homolog (Drosophila)
Length=1259
Score = 28.1 bits (61), Expect = 6.7, Method: Composition-based stats.
Identities = 12/26 (46%), Positives = 14/26 (53%), Gaps = 0/26 (0%)
Query 21 GAEPRPAAAGGGGGPPASNMQKDKRL 46
GA P AA GGG P +M + RL
Sbjct 404 GASPATVAAAGGGNSPRDHMARKMRL 429
> cel:C18H9.8 ift-74; IFT (Chlamydomonas IntraFlagellar Transport)
homolog family member (ift-74)
Length=674
Score = 27.7 bits (60), Expect = 9.6, Method: Composition-based stats.
Identities = 15/47 (31%), Positives = 17/47 (36%), Gaps = 0/47 (0%)
Query 14 PDSPDGEGAEPRPAAAGGGGGPPASNMQKDKRLDTAEGRPLESPCRP 60
P P G P P GGPP + R T RP + RP
Sbjct 114 PVPPSRSGGPPAPMPVSRAGGPPRAPTSMGGRPMTGMARPPTAGLRP 160
Lambda K H
0.310 0.131 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2105161088
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
Posted date: Sep 17, 2011 2:57 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40