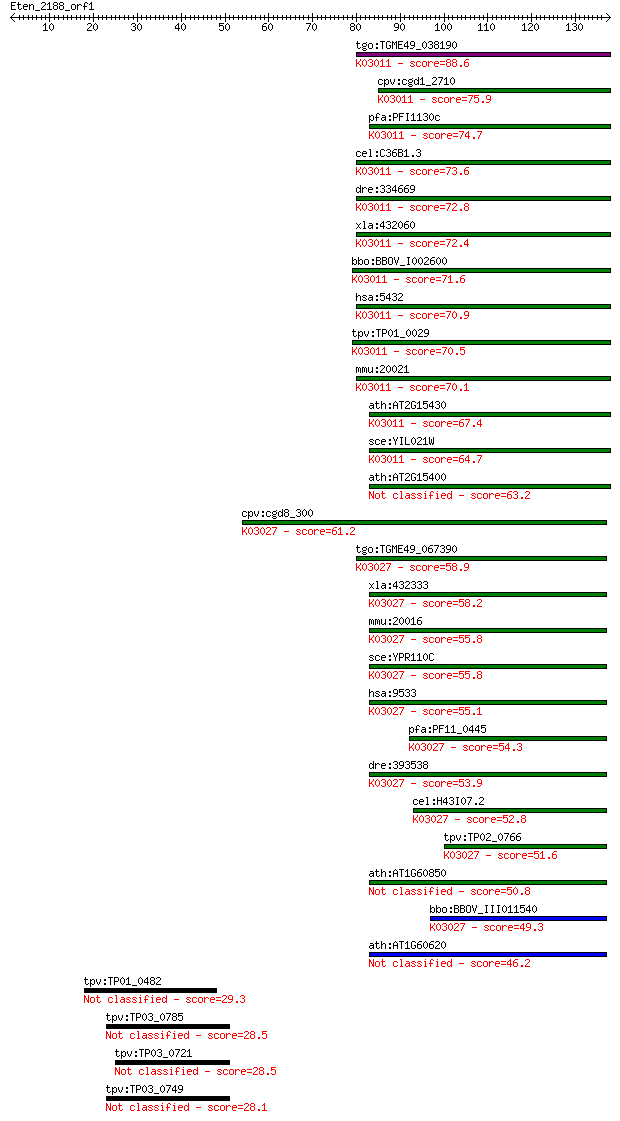
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
164,496 sequences; 82,071,388 total letters
Query= Eten_2188_orf1
Length=137
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_038190 DNA-directed RNA polymerase II, putative (EC... 88.6 6e-18
cpv:cgd1_2710 RNA polymerase II B3 subunit ; K03011 DNA-direct... 75.9 3e-14
pfa:PFI1130c DNA-directed RNA polymerase II, putative (EC:2.7.... 74.7 8e-14
cel:C36B1.3 rpb-3; RNA Polymerase II (B) subunit family member... 73.6 2e-13
dre:334669 polr2c, wu:fk48d01, zgc:56285; polymerase (RNA) II ... 72.8 3e-13
xla:432060 polr2c, MGC81245; polymerase (RNA) II (DNA directed... 72.4 4e-13
bbo:BBOV_I002600 19.m02023; DNA-directed RNA polymerase II (EC... 71.6 7e-13
hsa:5432 POLR2C, RPB3, RPB31, hRPB33, hsRPB3; polymerase (RNA)... 70.9 1e-12
tpv:TP01_0029 DNA-directed RNA polymerase II; K03011 DNA-direc... 70.5 2e-12
mmu:20021 Polr2c, 33kDa, MGC118242, RPB3, Rpo2-3, mRBP31; poly... 70.1 2e-12
ath:AT2G15430 NRPB3; NRPB3; DNA binding / DNA-directed RNA pol... 67.4 1e-11
sce:YIL021W RPB3; B44; K03011 DNA-directed RNA polymerase II s... 64.7 8e-11
ath:AT2G15400 NRPE3B; NRPE3B; DNA binding / DNA-directed RNA p... 63.2 2e-10
cpv:cgd8_300 RNA polymerase III C5 subunit ; K03027 DNA-direct... 61.2 9e-10
tgo:TGME49_067390 DNA-directed RNA polymerases I and III subun... 58.9 4e-09
xla:432333 polr1c, MGC78824, rpa39, rpa40, rpa5, rpac1; polyme... 58.2 7e-09
mmu:20016 Polr1c, 40kDa, AA409007, AA959927, AL024089, MGC1611... 55.8 3e-08
sce:YPR110C RPC40, RPC5; AC40; K03027 DNA-directed RNA polymer... 55.8 4e-08
hsa:9533 POLR1C, RPA39, RPA40, RPA5, RPAC1, TCS3; polymerase (... 55.1 7e-08
pfa:PF11_0445 DNA-directed RNA polymerase I, putative; K03027 ... 54.3 1e-07
dre:393538 polr1c, MGC65973, zgc:65973; polymerase (RNA) I pol... 53.9 2e-07
cel:H43I07.2 hypothetical protein; K03027 DNA-directed RNA pol... 52.8 3e-07
tpv:TP02_0766 DNA-directed RNA polymerase I; K03027 DNA-direct... 51.6 7e-07
ath:AT1G60850 ATRPAC42; DNA binding / DNA-directed RNA polymer... 50.8 1e-06
bbo:BBOV_III011540 17.m07985; DNA-directed RNA polymerase, alp... 49.3 4e-06
ath:AT1G60620 ATRPAC43; DNA binding / DNA-directed RNA polymer... 46.2 3e-05
tpv:TP01_0482 hypothetical protein 29.3 3.8
tpv:TP03_0785 hypothetical protein 28.5 6.3
tpv:TP03_0721 hypothetical protein 28.5 6.7
tpv:TP03_0749 hypothetical protein 28.1 7.7
> tgo:TGME49_038190 DNA-directed RNA polymerase II, putative (EC:2.7.7.6);
K03011 DNA-directed RNA polymerase II subunit RPB3
Length=484
Score = 88.6 bits (218), Expect = 6e-18, Method: Compositional matrix adjust.
Identities = 40/58 (68%), Positives = 47/58 (81%), Gaps = 0/58 (0%)
Query 80 QAAIRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDEY 137
+ +I V E+S V+F L NCD +IAN LRRVMISEVP+LAIDL TV+ENTSVLHDEY
Sbjct 150 EPSIVVKEVSQNKVKFLLENCDVSIANGLRRVMISEVPSLAIDLVTVYENTSVLHDEY 207
> cpv:cgd1_2710 RNA polymerase II B3 subunit ; K03011 DNA-directed
RNA polymerase II subunit RPB3
Length=355
Score = 75.9 bits (185), Expect = 3e-14, Method: Compositional matrix adjust.
Identities = 31/53 (58%), Positives = 45/53 (84%), Gaps = 0/53 (0%)
Query 85 VTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDEY 137
+TEL+ +RFTL+N D ++AN LRR++++E+PTLAIDL T+ +NTSVLHDE+
Sbjct 14 ITELTPNSIRFTLSNTDLSMANTLRRIILAEIPTLAIDLVTMIDNTSVLHDEF 66
> pfa:PFI1130c DNA-directed RNA polymerase II, putative (EC:2.7.7.6);
K03011 DNA-directed RNA polymerase II subunit RPB3
Length=335
Score = 74.7 bits (182), Expect = 8e-14, Method: Compositional matrix adjust.
Identities = 31/55 (56%), Positives = 43/55 (78%), Gaps = 0/55 (0%)
Query 83 IRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDEY 137
I +T+L+ + FTL N ++ IANALRR+M+SE+PTLAID+ V+ENTS HDE+
Sbjct 13 INITKLTKDELYFTLYNSNSGIANALRRIMLSEIPTLAIDVVNVYENTSAFHDEF 67
> cel:C36B1.3 rpb-3; RNA Polymerase II (B) subunit family member
(rpb-3); K03011 DNA-directed RNA polymerase II subunit RPB3
Length=282
Score = 73.6 bits (179), Expect = 2e-13, Method: Compositional matrix adjust.
Identities = 31/58 (53%), Positives = 44/58 (75%), Gaps = 0/58 (0%)
Query 80 QAAIRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDEY 137
Q I VTEL+N +++F L + D ++AN+LRRV ++EVPT+AID + NTSVLHDE+
Sbjct 6 QPNIEVTELTNDIIKFVLWDTDLSVANSLRRVFMAEVPTIAIDWVQIETNTSVLHDEF 63
> dre:334669 polr2c, wu:fk48d01, zgc:56285; polymerase (RNA) II
(DNA directed) polypeptide C (EC:2.7.7.6); K03011 DNA-directed
RNA polymerase II subunit RPB3
Length=275
Score = 72.8 bits (177), Expect = 3e-13, Method: Compositional matrix adjust.
Identities = 30/58 (51%), Positives = 45/58 (77%), Gaps = 0/58 (0%)
Query 80 QAAIRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDEY 137
Q +++TELS++ V+F + N D A+AN++RRV +SEVPT+AID + N+SVLHDE+
Sbjct 6 QPTVKITELSDENVKFVIENTDLAVANSIRRVFMSEVPTIAIDWVQIDANSSVLHDEF 63
> xla:432060 polr2c, MGC81245; polymerase (RNA) II (DNA directed)
polypeptide C, 33kDa; K03011 DNA-directed RNA polymerase
II subunit RPB3
Length=275
Score = 72.4 bits (176), Expect = 4e-13, Method: Compositional matrix adjust.
Identities = 31/58 (53%), Positives = 45/58 (77%), Gaps = 0/58 (0%)
Query 80 QAAIRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDEY 137
Q +R+TEL+++ V+F + N D AIAN++RRV I+EVPT+AID + N+SVLHDE+
Sbjct 6 QPTVRLTELTDENVKFIVENTDLAIANSIRRVFIAEVPTIAIDWVQIDANSSVLHDEF 63
> bbo:BBOV_I002600 19.m02023; DNA-directed RNA polymerase II (EC:2.7.7.6);
K03011 DNA-directed RNA polymerase II subunit RPB3
Length=382
Score = 71.6 bits (174), Expect = 7e-13, Method: Compositional matrix adjust.
Identities = 31/59 (52%), Positives = 45/59 (76%), Gaps = 0/59 (0%)
Query 79 MQAAIRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDEY 137
++ +I + L++ + F L N D A AN+LRR+M+SE+P+LAI++ TV ENTSVLHDEY
Sbjct 8 LRPSIDIVSLTSTKMIFILLNADVATANSLRRIMLSEIPSLAIEVVTVLENTSVLHDEY 66
> hsa:5432 POLR2C, RPB3, RPB31, hRPB33, hsRPB3; polymerase (RNA)
II (DNA directed) polypeptide C, 33kDa (EC:2.7.7.6); K03011
DNA-directed RNA polymerase II subunit RPB3
Length=275
Score = 70.9 bits (172), Expect = 1e-12, Method: Compositional matrix adjust.
Identities = 29/58 (50%), Positives = 44/58 (75%), Gaps = 0/58 (0%)
Query 80 QAAIRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDEY 137
Q +R+TEL+++ V+F + N D A+AN++RRV I+EVP +AID + N+SVLHDE+
Sbjct 6 QPTVRITELTDENVKFIIENTDLAVANSIRRVFIAEVPIIAIDWVQIDANSSVLHDEF 63
> tpv:TP01_0029 DNA-directed RNA polymerase II; K03011 DNA-directed
RNA polymerase II subunit RPB3
Length=354
Score = 70.5 bits (171), Expect = 2e-12, Method: Compositional matrix adjust.
Identities = 30/59 (50%), Positives = 44/59 (74%), Gaps = 0/59 (0%)
Query 79 MQAAIRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDEY 137
++ +I + +L + F L N D + ANA+RRV++SE+P+LAI++ TV ENTSVLHDEY
Sbjct 8 IKPSIEIVDLRKDRMDFILLNSDVSTANAIRRVILSEIPSLAIEIVTVLENTSVLHDEY 66
> mmu:20021 Polr2c, 33kDa, MGC118242, RPB3, Rpo2-3, mRBP31; polymerase
(RNA) II (DNA directed) polypeptide C (EC:2.7.7.6);
K03011 DNA-directed RNA polymerase II subunit RPB3
Length=275
Score = 70.1 bits (170), Expect = 2e-12, Method: Compositional matrix adjust.
Identities = 29/58 (50%), Positives = 43/58 (74%), Gaps = 0/58 (0%)
Query 80 QAAIRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDEY 137
Q +R+TEL+ + V+F + N D A+AN++RRV I+EVP +AID + N+SVLHDE+
Sbjct 6 QPTVRITELTEENVKFIIENTDLAVANSIRRVFIAEVPIIAIDWVQIDANSSVLHDEF 63
> ath:AT2G15430 NRPB3; NRPB3; DNA binding / DNA-directed RNA polymerase/
protein dimerization; K03011 DNA-directed RNA polymerase
II subunit RPB3
Length=319
Score = 67.4 bits (163), Expect = 1e-11, Method: Compositional matrix adjust.
Identities = 30/55 (54%), Positives = 41/55 (74%), Gaps = 0/55 (0%)
Query 83 IRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDEY 137
I++ EL + +F L D ++ANALRRVMISEVPT+AIDL + N+SVL+DE+
Sbjct 12 IKIRELKDDYAKFELRETDVSMANALRRVMISEVPTVAIDLVEIEVNSSVLNDEF 66
> sce:YIL021W RPB3; B44; K03011 DNA-directed RNA polymerase II
subunit RPB3
Length=318
Score = 64.7 bits (156), Expect = 8e-11, Method: Compositional matrix adjust.
Identities = 30/55 (54%), Positives = 40/55 (72%), Gaps = 0/55 (0%)
Query 83 IRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDEY 137
+++ E S V F L+N D A+AN+LRRVMI+E+PTLAID V NT+VL DE+
Sbjct 8 VKIREASKDNVDFILSNVDLAMANSLRRVMIAEIPTLAIDSVEVETNTTVLADEF 62
> ath:AT2G15400 NRPE3B; NRPE3B; DNA binding / DNA-directed RNA
polymerase/ protein dimerization
Length=235
Score = 63.2 bits (152), Expect = 2e-10, Method: Compositional matrix adjust.
Identities = 28/55 (50%), Positives = 40/55 (72%), Gaps = 0/55 (0%)
Query 83 IRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDEY 137
+++ EL + +F L D ++ANALRRVMISEVPT+AI L + N+SVL+DE+
Sbjct 12 VKIRELKDDYAKFELRETDVSMANALRRVMISEVPTMAIHLVKIEVNSSVLNDEF 66
> cpv:cgd8_300 RNA polymerase III C5 subunit ; K03027 DNA-directed
RNA polymerases I and III subunit RPAC1
Length=344
Score = 61.2 bits (147), Expect = 9e-10, Method: Compositional matrix adjust.
Identities = 25/84 (29%), Positives = 51/84 (60%), Gaps = 1/84 (1%)
Query 54 AAAAAAALQRLQQQPQLQQQQQQQLMQAA-IRVTELSNKVVRFTLNNCDAAIANALRRVM 112
+ L + + ++Q +++++ I+V EL + + F ++ DA+IANALRR++
Sbjct 27 GYIGSNELNSIIKGDNYRKQTVEEIIETIDIKVNELKSDYINFDISGIDASIANALRRIL 86
Query 113 ISEVPTLAIDLATVFENTSVLHDE 136
+ E+PT+AI+ +++N V+ DE
Sbjct 87 LVEIPTVAIETVQIYQNNGVIQDE 110
> tgo:TGME49_067390 DNA-directed RNA polymerases I and III subunit
RPAC1, putative (EC:2.7.7.6); K03027 DNA-directed RNA polymerases
I and III subunit RPAC1
Length=346
Score = 58.9 bits (141), Expect = 4e-09, Method: Compositional matrix adjust.
Identities = 26/57 (45%), Positives = 40/57 (70%), Gaps = 0/57 (0%)
Query 80 QAAIRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDE 136
+ +++ +S + V F L DAAI NALRR++ISEVPT+AI+ +++NT V+ DE
Sbjct 58 RTSLKFLSVSPERVVFELKGVDAAIPNALRRILISEVPTVAIETVQIWQNTGVVQDE 114
> xla:432333 polr1c, MGC78824, rpa39, rpa40, rpa5, rpac1; polymerase
(RNA) I polypeptide C, 30kDa; K03027 DNA-directed RNA
polymerases I and III subunit RPAC1
Length=347
Score = 58.2 bits (139), Expect = 7e-09, Method: Compositional matrix adjust.
Identities = 22/54 (40%), Positives = 39/54 (72%), Gaps = 0/54 (0%)
Query 83 IRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDE 136
I + ++ + ++ F + DA+IANA RR+++SEVPT+A++ V+ NTS++ DE
Sbjct 51 IDIVKMEDDILEFDMVGIDASIANAFRRILLSEVPTVAVEKVYVYNNTSIIQDE 104
> mmu:20016 Polr1c, 40kDa, AA409007, AA959927, AL024089, MGC161175,
Polr1e, RPA40, Rpo1-1; polymerase (RNA) I polypeptide
C; K03027 DNA-directed RNA polymerases I and III subunit RPAC1
Length=346
Score = 55.8 bits (133), Expect = 3e-08, Method: Compositional matrix adjust.
Identities = 22/54 (40%), Positives = 37/54 (68%), Gaps = 0/54 (0%)
Query 83 IRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDE 136
+ V ++ + F + DAAIANA RR++++EVPT+A++ V+ NTS++ DE
Sbjct 51 VDVVQMDEDTLEFDMVGIDAAIANAFRRILLAEVPTMAVEKVLVYNNTSIVQDE 104
> sce:YPR110C RPC40, RPC5; AC40; K03027 DNA-directed RNA polymerases
I and III subunit RPAC1
Length=335
Score = 55.8 bits (133), Expect = 4e-08, Method: Compositional matrix adjust.
Identities = 25/54 (46%), Positives = 35/54 (64%), Gaps = 0/54 (0%)
Query 83 IRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDE 136
+ ++ L + F L N D +IANA RR+MISEVP++A + F NTSV+ DE
Sbjct 42 VNISSLDAREANFDLINIDTSIANAFRRIMISEVPSVAAEYVYFFNNTSVIQDE 95
> hsa:9533 POLR1C, RPA39, RPA40, RPA5, RPAC1, TCS3; polymerase
(RNA) I polypeptide C, 30kDa; K03027 DNA-directed RNA polymerases
I and III subunit RPAC1
Length=342
Score = 55.1 bits (131), Expect = 7e-08, Method: Compositional matrix adjust.
Identities = 22/54 (40%), Positives = 36/54 (66%), Gaps = 0/54 (0%)
Query 83 IRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDE 136
+ V + + F + DAAIANA RR++++EVPT+A++ V+ NTS++ DE
Sbjct 51 VDVVHMDENSLEFDMVGIDAAIANAFRRILLAEVPTMAVEKVLVYNNTSIVQDE 104
> pfa:PF11_0445 DNA-directed RNA polymerase I, putative; K03027
DNA-directed RNA polymerases I and III subunit RPAC1
Length=333
Score = 54.3 bits (129), Expect = 1e-07, Method: Compositional matrix adjust.
Identities = 21/45 (46%), Positives = 34/45 (75%), Gaps = 0/45 (0%)
Query 92 VVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDE 136
++ + N D +IANALRR+M+SEVPT+AI+ +++NT ++ DE
Sbjct 59 ILILEIKNMDVSIANALRRIMLSEVPTIAIEKVNMYQNTGIIADE 103
> dre:393538 polr1c, MGC65973, zgc:65973; polymerase (RNA) I polypeptide
C; K03027 DNA-directed RNA polymerases I and III
subunit RPAC1
Length=347
Score = 53.9 bits (128), Expect = 2e-07, Method: Compositional matrix adjust.
Identities = 22/54 (40%), Positives = 35/54 (64%), Gaps = 0/54 (0%)
Query 83 IRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDE 136
I + + F + DAAIANA RR++++EVPT+AI+ ++ NTS++ DE
Sbjct 52 IEIISSDENTLEFDMIGIDAAIANAFRRILLAEVPTMAIEKVFIYNNTSIIQDE 105
> cel:H43I07.2 hypothetical protein; K03027 DNA-directed RNA polymerases
I and III subunit RPAC1
Length=366
Score = 52.8 bits (125), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 22/44 (50%), Positives = 34/44 (77%), Gaps = 0/44 (0%)
Query 93 VRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDE 136
+ F + +A IANALRRV+I+EVPT+A++ +++NTSV+ DE
Sbjct 68 LEFDITRIEAPIANALRRVLIAEVPTMALEKIYLYQNTSVIQDE 111
> tpv:TP02_0766 DNA-directed RNA polymerase I; K03027 DNA-directed
RNA polymerases I and III subunit RPAC1
Length=303
Score = 51.6 bits (122), Expect = 7e-07, Method: Compositional matrix adjust.
Identities = 20/37 (54%), Positives = 30/37 (81%), Gaps = 0/37 (0%)
Query 100 CDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDE 136
D ++ANALRR++ISEVPTLA++ +++NT V+ DE
Sbjct 52 LDVSLANALRRILISEVPTLAVETVQIYQNTGVIQDE 88
> ath:AT1G60850 ATRPAC42; DNA binding / DNA-directed RNA polymerase/
protein dimerization
Length=375
Score = 50.8 bits (120), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 23/54 (42%), Positives = 35/54 (64%), Gaps = 0/54 (0%)
Query 83 IRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDE 136
+ V L+ + F + DAA ANA RR++I+EVP++AI+ + NTSV+ DE
Sbjct 76 VDVVSLTKTDMEFDMIGIDAAFANAFRRILIAEVPSMAIEKVLIAYNTSVIIDE 129
> bbo:BBOV_III011540 17.m07985; DNA-directed RNA polymerase, alpha
subunit, N terminal domain containing protein; K03027 DNA-directed
RNA polymerases I and III subunit RPAC1
Length=319
Score = 49.3 bits (116), Expect = 4e-06, Method: Compositional matrix adjust.
Identities = 19/40 (47%), Positives = 31/40 (77%), Gaps = 0/40 (0%)
Query 97 LNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDE 136
+ + AIANA+RR++++EVPTLAI+ +++NT V+ DE
Sbjct 50 IEGLNVAIANAIRRILLAEVPTLAIESVHIYQNTGVIQDE 89
> ath:AT1G60620 ATRPAC43; DNA binding / DNA-directed RNA polymerase/
protein dimerization
Length=385
Score = 46.2 bits (108), Expect = 3e-05, Method: Compositional matrix adjust.
Identities = 21/54 (38%), Positives = 34/54 (62%), Gaps = 0/54 (0%)
Query 83 IRVTELSNKVVRFTLNNCDAAIANALRRVMISEVPTLAIDLATVFENTSVLHDE 136
+ V L+ + F + A IANA RR++++E+P++AI+ V NTSV+ DE
Sbjct 83 VDVISLTETDMVFDMIGVHAGIANAFRRILLAELPSMAIEKVYVANNTSVIQDE 136
> tpv:TP01_0482 hypothetical protein
Length=918
Score = 29.3 bits (64), Expect = 3.8, Method: Composition-based stats.
Identities = 13/38 (34%), Positives = 18/38 (47%), Gaps = 8/38 (21%)
Query 18 RPSSSSGC--------CSYFHNSCCCCCCSCCCCCRSS 47
+PSS S C C + +SC CC C CC + +
Sbjct 805 QPSSQSQCECCKSQGKCCFKTDSCTCCKDPCDCCSKGN 842
> tpv:TP03_0785 hypothetical protein
Length=1796
Score = 28.5 bits (62), Expect = 6.3, Method: Composition-based stats.
Identities = 13/31 (41%), Positives = 16/31 (51%), Gaps = 3/31 (9%)
Query 23 SGCCSYFHNSCCCCCCS---CCCCCRSSEDK 50
S CCS N+ C CC CC C SS+ +
Sbjct 784 SKCCSSDGNNNCTCCNGDKVKCCLCPSSKSE 814
> tpv:TP03_0721 hypothetical protein
Length=1181
Score = 28.5 bits (62), Expect = 6.7, Method: Composition-based stats.
Identities = 12/29 (41%), Positives = 14/29 (48%), Gaps = 3/29 (10%)
Query 25 CCSYFHNSCCCCCCS---CCCCCRSSEDK 50
CCS CC C S C CCC+ + K
Sbjct 162 CCSPKKVQCCYCAESKADCNCCCKENNKK 190
> tpv:TP03_0749 hypothetical protein
Length=1024
Score = 28.1 bits (61), Expect = 7.7, Method: Composition-based stats.
Identities = 13/31 (41%), Positives = 16/31 (51%), Gaps = 3/31 (9%)
Query 23 SGCCSYFHNSCCCCCCS---CCCCCRSSEDK 50
S CCS N+ C CC CC C SS+ +
Sbjct 12 SKCCSSDGNNNCTCCNGDKVKCCLCPSSKSE 42
Lambda K H
0.322 0.127 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2428006156
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
Posted date: Sep 17, 2011 2:57 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40