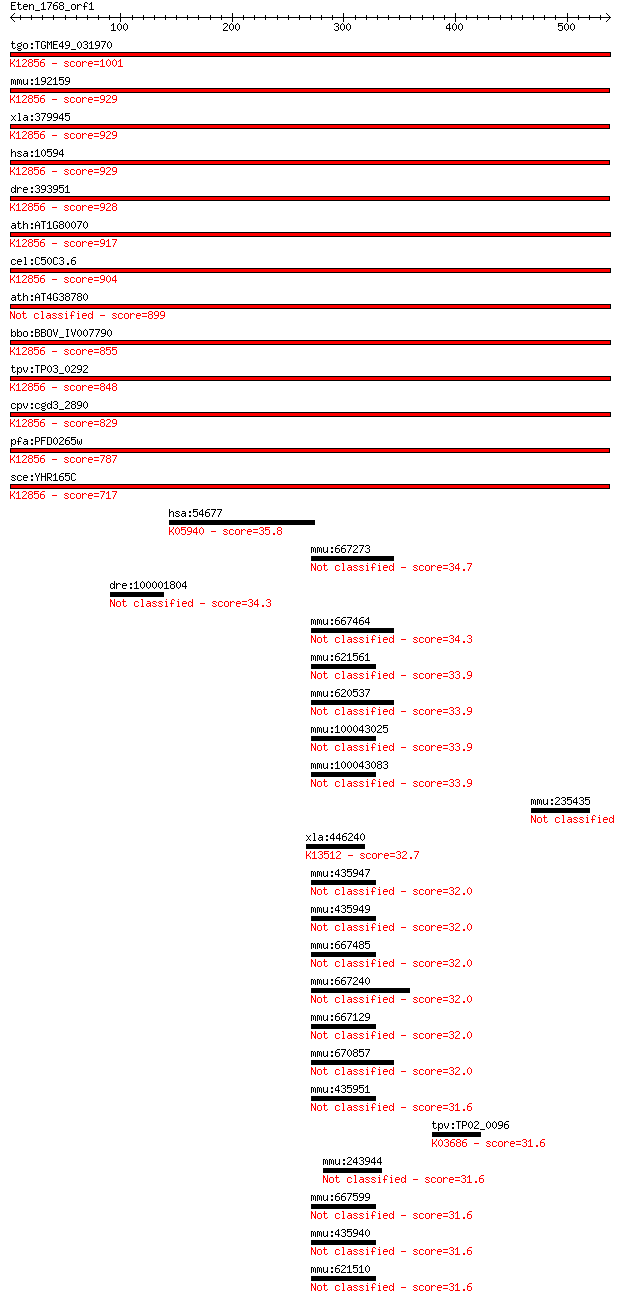
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
164,496 sequences; 82,071,388 total letters
Query= Eten_1768_orf1
Length=538
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_031970 pre-mRNA splicing factor PRP8, putative ; K1... 1001 0.0
mmu:192159 Prpf8, AU019467, D11Bwg0410e, DBF3/PRP8, Prp8, Sfpr... 929 0.0
xla:379945 prpf8, MGC52804, hprp8, prp-8, prp8, prpc8, rp13; P... 929 0.0
hsa:10594 PRPF8, HPRP8, PRP8, PRPC8, RP13; PRP8 pre-mRNA proce... 929 0.0
dre:393951 prpf8, MGC162871, MGC56504, id:ibd1257, ik:tdsubc_2... 928 0.0
ath:AT1G80070 SUS2; SUS2 (ABNORMAL SUSPENSOR 2); K12856 pre-mR... 917 0.0
cel:C50C3.6 prp-8; yeast PRP (splicing factor) related family ... 904 0.0
ath:AT4G38780 splicing factor, putative 899 0.0
bbo:BBOV_IV007790 23.m06497; processing splicing factor 8; K12... 855 0.0
tpv:TP03_0292 splicing factor Prp8; K12856 pre-mRNA-processing... 848 0.0
cpv:cgd3_2890 Prp8. JAB/PAD domain ; K12856 pre-mRNA-processin... 829 0.0
pfa:PFD0265w pre-mRNA splicing factor, putative; K12856 pre-mR... 787 0.0
sce:YHR165C PRP8, DBF3, DNA39, RNA8, SLT21, USA2; Component of... 717 0.0
hsa:54677 CROT, COT; carnitine O-octanoyltransferase (EC:2.3.1... 35.8 0.48
mmu:667273 Vmn1r115, EG667273, Gm8549; vomeronasal 1 receptor 115 34.7
dre:100001804 novel protein containing lectin C-type domains 34.3 1.4
mmu:667464 Vmn1r142, EG667464, Gm8647; vomeronasal 1 receptor 142 34.3
mmu:621561 Vmn1r127, Gm6239; vomeronasal 1 receptor 127 33.9
mmu:620537 Vmn1r94, EG620537, Gm6160; vomeronasal 1 receptor 94 33.9
mmu:100043025 Gm4177, Vmn1r134; predicted gene 4177 33.9
mmu:100043083 Gm4216, Vmn1r162; predicted gene 4216 33.9
mmu:235435 Lctl, E130104I05Rik, KLPH; lactase-like 32.7 3.3
xla:446240 lpcat4, agpat7, aytl3; lysophosphatidylcholine acyl... 32.7 4.1
mmu:435947 Vmn1r151, EG435947, Gm5727; vomeronasal 1 receptor 151 32.0
mmu:435949 Gm5728, EG435949, Vmn1r99; predicted gene 5728 32.0
mmu:667485 Gm8660, EG667485, Vmn1r147; predicted gene 8660 32.0
mmu:667240 Vmn1r121, EG667240, Gm8533; vomeronasal 1 receptor 121 32.0
mmu:667129 Vmn1r103, EG667129, Gm8474; vomeronasal 1 receptor 103 32.0
mmu:670857 Vmn1r159, Gm16507; vomeronasal 1 receptor 159 32.0
mmu:435951 Vmn1r122, EG435951, Gm5729; vomeronasal 1 receptor 122 31.6
tpv:TP02_0096 chaperone protein DnaJ; K03686 molecular chapero... 31.6 8.2
mmu:243944 4930433I11Rik, Gm481; RIKEN cDNA 4930433I11 gene 31.6 8.4
mmu:667599 Gm8720, EG667599, Vmn1r164; predicted gene 8720 31.6
mmu:435940 Gm5725, EG435940, Vmn1r136; predicted gene 5725 31.6
mmu:621510 Vmn1r129, EG621510, Gm6237; vomeronasal 1 receptor 129 31.6
> tgo:TGME49_031970 pre-mRNA splicing factor PRP8, putative ;
K12856 pre-mRNA-processing factor 8
Length=2538
Score = 1001 bits (2588), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 467/538 (86%), Positives = 511/538 (94%), Gaps = 0/538 (0%)
Query 1 VHESIVMDLCQVFDMELDSLEVETVQKETIHPRKSYKMNSSCADILLFATYKWSISKPSL 60
VHESIVMDLCQVFD+ELDSLE+E VQKETIHPRKSYKMNSSCADILLFA YKW ISKPSL
Sbjct 1787 VHESIVMDLCQVFDLELDSLEIEMVQKETIHPRKSYKMNSSCADILLFAAYKWQISKPSL 1846
Query 61 LAEGKDIMEGATATKHWLDVQLRWGDYDSHDIERYCRSKFLDYTTDNMSIYPSPSGVLVG 120
LA+GKD+M+G T +K+WLD+QLRWGD+DSHDIERYCRSKFLDYTTDNMSIYPSP+GVL+G
Sbjct 1847 LADGKDVMDGTTTSKYWLDIQLRWGDFDSHDIERYCRSKFLDYTTDNMSIYPSPTGVLLG 1906
Query 121 VDLAYNLHSAFGNWIPGLKPLMQRAMNKIMKANPALYVLRERIRKGLQLYSSEPTEPYLN 180
VDLAYNLHS FGNW PGLKPLMQRAMNKIMK+NPALYVLRERIRKGLQLYSSEPTEPYL
Sbjct 1907 VDLAYNLHSGFGNWFPGLKPLMQRAMNKIMKSNPALYVLRERIRKGLQLYSSEPTEPYLT 1966
Query 181 SQNYGELFSNQTVWFIDDTNVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKIIHT 240
SQNYGELFSNQT+WF+DDTNVYRVTIHKTFEGNLTTKPVNG IFIFNPRTGQLFLKIIHT
Sbjct 1967 SQNYGELFSNQTIWFVDDTNVYRVTIHKTFEGNLTTKPVNGAIFIFNPRTGQLFLKIIHT 2026
Query 241 SVWAGQKRLIQLARWKTAEEVAALIRSLPVEEQPKQIIATRKGMLDPLEVHLLDFPNIVI 300
SVWAGQKRL QLA+WKTAEEVAALIRSLPVEEQPKQ+IATRKGMLDPLEVHLLDFPNIVI
Sbjct 2027 SVWAGQKRLTQLAKWKTAEEVAALIRSLPVEEQPKQLIATRKGMLDPLEVHLLDFPNIVI 2086
Query 301 KGSELNLPFQALMRIEKFGDMILRATQPEMVLFNMFDDWLKSISAYTAFSRLLLLLRALQ 360
KGSELNLPFQA+M++EKFGDMIL+ATQPEMVLFNM+DDWLKSIS+YTAFSRLLLLLRA+
Sbjct 2087 KGSELNLPFQAIMKVEKFGDMILKATQPEMVLFNMYDDWLKSISSYTAFSRLLLLLRAMH 2146
Query 361 VNTERAKVLLRPHKSVVTQPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVASLTQ 420
VNTER K++LRP+K+ VTQ HHIWPSLTDEEWIHVEV LKDLIL DY KKNNVNVASLTQ
Sbjct 2147 VNTERTKIILRPNKTTVTQSHHIWPSLTDEEWIHVEVALKDLILADYGKKNNVNVASLTQ 2206
Query 421 SEIRDIILGMEIAPPSLQRQQIAEIEAQAREQQQVTSTTTRSVNIHGEEMIVATQSPHEQ 480
SEIRDIILGMEI+PPSLQRQQIAEIEAQ ++ QVT+TTTR+VN HG+E+IV+TQSPHEQ
Sbjct 2207 SEIRDIILGMEISPPSLQRQQIAEIEAQTKDVSQVTATTTRTVNAHGDEIIVSTQSPHEQ 2266
Query 481 QVFSSKTDWRVRAISAASLHLRTRHLYVSSEAVAPAGCTYILPKNLLKKFISIADLRT 538
QVFSSKTDWR+RAISAASLHLRT H+YV+S+ + +G TY+LPKNLLKKFI ++DLRT
Sbjct 2267 QVFSSKTDWRIRAISAASLHLRTHHIYVNSDDIKESGYTYVLPKNLLKKFICVSDLRT 2324
> mmu:192159 Prpf8, AU019467, D11Bwg0410e, DBF3/PRP8, Prp8, Sfprp8l;
pre-mRNA processing factor 8; K12856 pre-mRNA-processing
factor 8
Length=2335
Score = 929 bits (2402), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 433/537 (80%), Positives = 493/537 (91%), Gaps = 0/537 (0%)
Query 1 VHESIVMDLCQVFDMELDSLEVETVQKETIHPRKSYKMNSSCADILLFATYKWSISKPSL 60
+HESIVMDLCQVFD ELD+LE+ETVQKETIHPRKSYKMNSSCADILLFA+YKW++S+PSL
Sbjct 1585 IHESIVMDLCQVFDQELDALEIETVQKETIHPRKSYKMNSSCADILLFASYKWNVSRPSL 1644
Query 61 LAEGKDIMEGATATKHWLDVQLRWGDYDSHDIERYCRSKFLDYTTDNMSIYPSPSGVLVG 120
LA+ KD+M+ T K+W+D+QLRWGDYDSHDIERY R+KFLDYTTDNMSIYPSP+GVL+
Sbjct 1645 LADSKDVMDSTTTQKYWIDIQLRWGDYDSHDIERYARAKFLDYTTDNMSIYPSPTGVLIA 1704
Query 121 VDLAYNLHSAFGNWIPGLKPLMQRAMNKIMKANPALYVLRERIRKGLQLYSSEPTEPYLN 180
+DLAYNLHSA+GNW PG KPL+Q+AM KIMKANPALYVLRERIRKGLQLYSSEPTEPYL+
Sbjct 1705 IDLAYNLHSAYGNWFPGSKPLIQQAMAKIMKANPALYVLRERIRKGLQLYSSEPTEPYLS 1764
Query 181 SQNYGELFSNQTVWFIDDTNVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKIIHT 240
SQNYGELFSNQ +WF+DDTNVYRVTIHKTFEGNLTTKP+NG IFIFNPRTGQLFLKIIHT
Sbjct 1765 SQNYGELFSNQIIWFVDDTNVYRVTIHKTFEGNLTTKPINGAIFIFNPRTGQLFLKIIHT 1824
Query 241 SVWAGQKRLIQLARWKTAEEVAALIRSLPVEEQPKQIIATRKGMLDPLEVHLLDFPNIVI 300
SVWAGQKRL QLA+WKTAEEVAALIRSLPVEEQPKQII TRKGMLDPLEVHLLDFPNIVI
Sbjct 1825 SVWAGQKRLGQLAKWKTAEEVAALIRSLPVEEQPKQIIVTRKGMLDPLEVHLLDFPNIVI 1884
Query 301 KGSELNLPFQALMRIEKFGDMILRATQPEMVLFNMFDDWLKSISAYTAFSRLLLLLRALQ 360
KGSEL LPFQA +++EKFGD+IL+AT+P+MVLFN++DDWLK+IS+YTAFSRL+L+LRAL
Sbjct 1885 KGSELQLPFQACLKVEKFGDLILKATEPQMVLFNLYDDWLKTISSYTAFSRLILILRALH 1944
Query 361 VNTERAKVLLRPHKSVVTQPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVASLTQ 420
VN +RAKV+L+P K+ VT+PHHIWP+LTDEEWI VEV LKDLIL DY KKNNVNVASLTQ
Sbjct 1945 VNNDRAKVILKPDKTTVTEPHHIWPTLTDEEWIKVEVQLKDLILADYGKKNNVNVASLTQ 2004
Query 421 SEIRDIILGMEIAPPSLQRQQIAEIEAQAREQQQVTSTTTRSVNIHGEEMIVATQSPHEQ 480
SEIRDIILGMEI+ PS QRQQIAEIE Q +EQ Q+T+T TR+VN HG+E+I +T S +E
Sbjct 2005 SEIRDIILGMEISAPSQQRQQIAEIEKQTKEQSQLTATQTRTVNKHGDEIITSTTSNYET 2064
Query 481 QVFSSKTDWRVRAISAASLHLRTRHLYVSSEAVAPAGCTYILPKNLLKKFISIADLR 537
Q FSSKT+WRVRAISAA+LHLRT H+YVSS+ + G TYILPKN+LKKFI I+DLR
Sbjct 2065 QTFSSKTEWRVRAISAANLHLRTNHIYVSSDDIKETGYTYILPKNVLKKFICISDLR 2121
> xla:379945 prpf8, MGC52804, hprp8, prp-8, prp8, prpc8, rp13;
PRP8 pre-mRNA processing factor 8 homolog; K12856 pre-mRNA-processing
factor 8
Length=2335
Score = 929 bits (2401), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 432/537 (80%), Positives = 493/537 (91%), Gaps = 0/537 (0%)
Query 1 VHESIVMDLCQVFDMELDSLEVETVQKETIHPRKSYKMNSSCADILLFATYKWSISKPSL 60
+HESIVMDLCQVFD ELD+LE+ETVQKETIHPRKSYKMNSSCADILLFA+YKW++S+PSL
Sbjct 1585 IHESIVMDLCQVFDQELDALEIETVQKETIHPRKSYKMNSSCADILLFASYKWNVSRPSL 1644
Query 61 LAEGKDIMEGATATKHWLDVQLRWGDYDSHDIERYCRSKFLDYTTDNMSIYPSPSGVLVG 120
LA+ KD+M+ T K+W+D+QLRWGDYDSHDIERY R+KFLDYTTDNMSIYPSP+GVL+
Sbjct 1645 LADSKDVMDSTTTQKYWIDIQLRWGDYDSHDIERYARAKFLDYTTDNMSIYPSPTGVLIA 1704
Query 121 VDLAYNLHSAFGNWIPGLKPLMQRAMNKIMKANPALYVLRERIRKGLQLYSSEPTEPYLN 180
+DLAYNLHSA+GNW PG KPL+Q+AM KIMKANPALYVLRERIRKGLQLYSSEPTEPYL+
Sbjct 1705 IDLAYNLHSAYGNWFPGGKPLIQQAMAKIMKANPALYVLRERIRKGLQLYSSEPTEPYLS 1764
Query 181 SQNYGELFSNQTVWFIDDTNVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKIIHT 240
SQNYGELFSNQ +WF+DDTNVYRVTIHKTFEGNLTTKP+NG IFIFNPRTGQLFLKIIHT
Sbjct 1765 SQNYGELFSNQIIWFVDDTNVYRVTIHKTFEGNLTTKPINGAIFIFNPRTGQLFLKIIHT 1824
Query 241 SVWAGQKRLIQLARWKTAEEVAALIRSLPVEEQPKQIIATRKGMLDPLEVHLLDFPNIVI 300
SVWAGQKRL QLA+WKTAEEVAALIRSLPVEEQPKQII TRKGMLDPLEVHLLDFPNIVI
Sbjct 1825 SVWAGQKRLGQLAKWKTAEEVAALIRSLPVEEQPKQIIVTRKGMLDPLEVHLLDFPNIVI 1884
Query 301 KGSELNLPFQALMRIEKFGDMILRATQPEMVLFNMFDDWLKSISAYTAFSRLLLLLRALQ 360
KGSEL LPFQA +++EKFGD+IL+AT+P+MVLFN++DDWLK+IS+YTAFSRL+L+LRAL
Sbjct 1885 KGSELQLPFQACLKVEKFGDLILKATEPQMVLFNLYDDWLKTISSYTAFSRLILILRALH 1944
Query 361 VNTERAKVLLRPHKSVVTQPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVASLTQ 420
VN +RAKV+L+P K+ +T+PHHIWP+LTDEEWI VEV LKDLIL DY KKNNVNVASLTQ
Sbjct 1945 VNNDRAKVILKPDKTTITEPHHIWPTLTDEEWIKVEVQLKDLILADYGKKNNVNVASLTQ 2004
Query 421 SEIRDIILGMEIAPPSLQRQQIAEIEAQAREQQQVTSTTTRSVNIHGEEMIVATQSPHEQ 480
SEIRDIILGMEI+ PS QRQQIAEIE Q +EQ Q+T+T TR+VN HG+E+I +T S +E
Sbjct 2005 SEIRDIILGMEISAPSQQRQQIAEIEKQTKEQSQLTATQTRTVNKHGDEIITSTTSNYET 2064
Query 481 QVFSSKTDWRVRAISAASLHLRTRHLYVSSEAVAPAGCTYILPKNLLKKFISIADLR 537
Q FSSKT+WRVRAISAA+LHLRT H+YVSS+ + G TYILPKN+LKKFI I+DLR
Sbjct 2065 QTFSSKTEWRVRAISAANLHLRTNHIYVSSDDIKETGYTYILPKNVLKKFICISDLR 2121
> hsa:10594 PRPF8, HPRP8, PRP8, PRPC8, RP13; PRP8 pre-mRNA processing
factor 8 homolog (S. cerevisiae); K12856 pre-mRNA-processing
factor 8
Length=2335
Score = 929 bits (2401), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 432/537 (80%), Positives = 493/537 (91%), Gaps = 0/537 (0%)
Query 1 VHESIVMDLCQVFDMELDSLEVETVQKETIHPRKSYKMNSSCADILLFATYKWSISKPSL 60
+HESIVMDLCQVFD ELD+LE+ETVQKETIHPRKSYKMNSSCADILLFA+YKW++S+PSL
Sbjct 1585 IHESIVMDLCQVFDQELDALEIETVQKETIHPRKSYKMNSSCADILLFASYKWNVSRPSL 1644
Query 61 LAEGKDIMEGATATKHWLDVQLRWGDYDSHDIERYCRSKFLDYTTDNMSIYPSPSGVLVG 120
LA+ KD+M+ T K+W+D+QLRWGDYDSHDIERY R+KFLDYTTDNMSIYPSP+GVL+
Sbjct 1645 LADSKDVMDSTTTQKYWIDIQLRWGDYDSHDIERYARAKFLDYTTDNMSIYPSPTGVLIA 1704
Query 121 VDLAYNLHSAFGNWIPGLKPLMQRAMNKIMKANPALYVLRERIRKGLQLYSSEPTEPYLN 180
+DLAYNLHSA+GNW PG KPL+Q+AM KIMKANPALYVLRERIRKGLQLYSSEPTEPYL+
Sbjct 1705 IDLAYNLHSAYGNWFPGSKPLIQQAMAKIMKANPALYVLRERIRKGLQLYSSEPTEPYLS 1764
Query 181 SQNYGELFSNQTVWFIDDTNVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKIIHT 240
SQNYGELFSNQ +WF+DDTNVYRVTIHKTFEGNLTTKP+NG IFIFNPRTGQLFLKIIHT
Sbjct 1765 SQNYGELFSNQIIWFVDDTNVYRVTIHKTFEGNLTTKPINGAIFIFNPRTGQLFLKIIHT 1824
Query 241 SVWAGQKRLIQLARWKTAEEVAALIRSLPVEEQPKQIIATRKGMLDPLEVHLLDFPNIVI 300
SVWAGQKRL QLA+WKTAEEVAALIRSLPVEEQPKQII TRKGMLDPLEVHLLDFPNIVI
Sbjct 1825 SVWAGQKRLGQLAKWKTAEEVAALIRSLPVEEQPKQIIVTRKGMLDPLEVHLLDFPNIVI 1884
Query 301 KGSELNLPFQALMRIEKFGDMILRATQPEMVLFNMFDDWLKSISAYTAFSRLLLLLRALQ 360
KGSEL LPFQA +++EKFGD+IL+AT+P+MVLFN++DDWLK+IS+YTAFSRL+L+LRAL
Sbjct 1885 KGSELQLPFQACLKVEKFGDLILKATEPQMVLFNLYDDWLKTISSYTAFSRLILILRALH 1944
Query 361 VNTERAKVLLRPHKSVVTQPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVASLTQ 420
VN +RAKV+L+P K+ +T+PHHIWP+LTDEEWI VEV LKDLIL DY KKNNVNVASLTQ
Sbjct 1945 VNNDRAKVILKPDKTTITEPHHIWPTLTDEEWIKVEVQLKDLILADYGKKNNVNVASLTQ 2004
Query 421 SEIRDIILGMEIAPPSLQRQQIAEIEAQAREQQQVTSTTTRSVNIHGEEMIVATQSPHEQ 480
SEIRDIILGMEI+ PS QRQQIAEIE Q +EQ Q+T+T TR+VN HG+E+I +T S +E
Sbjct 2005 SEIRDIILGMEISAPSQQRQQIAEIEKQTKEQSQLTATQTRTVNKHGDEIITSTTSNYET 2064
Query 481 QVFSSKTDWRVRAISAASLHLRTRHLYVSSEAVAPAGCTYILPKNLLKKFISIADLR 537
Q FSSKT+WRVRAISAA+LHLRT H+YVSS+ + G TYILPKN+LKKFI I+DLR
Sbjct 2065 QTFSSKTEWRVRAISAANLHLRTNHIYVSSDDIKETGYTYILPKNVLKKFICISDLR 2121
> dre:393951 prpf8, MGC162871, MGC56504, id:ibd1257, ik:tdsubc_2a9,
im:7141966, tdsubc_2a9, wu:fb37c02, wu:fb73e06, xx:tdsubc_2a9,
zgc:56504; pre-mRNA processing factor 8; K12856 pre-mRNA-processing
factor 8
Length=2342
Score = 928 bits (2398), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 432/537 (80%), Positives = 492/537 (91%), Gaps = 0/537 (0%)
Query 1 VHESIVMDLCQVFDMELDSLEVETVQKETIHPRKSYKMNSSCADILLFATYKWSISKPSL 60
+HESIVMDLCQVFD ELD+LE+ETVQKETIHPRKSYKMNSSCADILLFA+YKW++S+PSL
Sbjct 1592 IHESIVMDLCQVFDQELDALEIETVQKETIHPRKSYKMNSSCADILLFASYKWNVSRPSL 1651
Query 61 LAEGKDIMEGATATKHWLDVQLRWGDYDSHDIERYCRSKFLDYTTDNMSIYPSPSGVLVG 120
LA+ KD+M+ T K W+D+QLRWGDYDSHDIERY R+KFLDYTTDNMSIYPSP+GVL+
Sbjct 1652 LADSKDVMDSTTTQKFWIDIQLRWGDYDSHDIERYARAKFLDYTTDNMSIYPSPTGVLIA 1711
Query 121 VDLAYNLHSAFGNWIPGLKPLMQRAMNKIMKANPALYVLRERIRKGLQLYSSEPTEPYLN 180
+DLAYNLHSA+GNW PG KPL+Q+AM KIMKANPALYVLRERIRKGLQLYSSEPTEPYL+
Sbjct 1712 IDLAYNLHSAYGNWFPGAKPLIQQAMAKIMKANPALYVLRERIRKGLQLYSSEPTEPYLS 1771
Query 181 SQNYGELFSNQTVWFIDDTNVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKIIHT 240
SQNYGELFSNQ +WF+DDTNVYRVTIHKTFEGNLTTKP+NG IFIFNPRTGQLFLKIIHT
Sbjct 1772 SQNYGELFSNQIIWFVDDTNVYRVTIHKTFEGNLTTKPINGAIFIFNPRTGQLFLKIIHT 1831
Query 241 SVWAGQKRLIQLARWKTAEEVAALIRSLPVEEQPKQIIATRKGMLDPLEVHLLDFPNIVI 300
SVWAGQKRL QLA+WKTAEEVAALIRSLPVEEQPKQII TRKGMLDPLEVHLLDFPNIVI
Sbjct 1832 SVWAGQKRLGQLAKWKTAEEVAALIRSLPVEEQPKQIIVTRKGMLDPLEVHLLDFPNIVI 1891
Query 301 KGSELNLPFQALMRIEKFGDMILRATQPEMVLFNMFDDWLKSISAYTAFSRLLLLLRALQ 360
KGSEL LPFQA +++EKFGD+IL+AT+P+MVLFN++DDWLK+IS+YTAFSRL+L+LRAL
Sbjct 1892 KGSELQLPFQACLKVEKFGDLILKATEPQMVLFNLYDDWLKTISSYTAFSRLILILRALH 1951
Query 361 VNTERAKVLLRPHKSVVTQPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVASLTQ 420
VN +RAKV+L+P K+ +T+PHHIWP+LTDEEWI VEV LKDLIL DY KKNNVNVASLTQ
Sbjct 1952 VNNDRAKVILKPDKTTITEPHHIWPTLTDEEWIKVEVQLKDLILADYGKKNNVNVASLTQ 2011
Query 421 SEIRDIILGMEIAPPSLQRQQIAEIEAQAREQQQVTSTTTRSVNIHGEEMIVATQSPHEQ 480
SEIRDIILGMEI+ PS QRQQIAEIE Q +EQ Q+T+T TR+VN HG+E+I +T S +E
Sbjct 2012 SEIRDIILGMEISAPSQQRQQIAEIEKQTKEQSQLTATQTRTVNKHGDEIITSTTSNYET 2071
Query 481 QVFSSKTDWRVRAISAASLHLRTRHLYVSSEAVAPAGCTYILPKNLLKKFISIADLR 537
Q FSSKT+WRVRAISAA+LHLRT H+YVSS+ + G TYILPKN+LKKFI I+DLR
Sbjct 2072 QTFSSKTEWRVRAISAANLHLRTNHIYVSSDDIKETGYTYILPKNVLKKFICISDLR 2128
> ath:AT1G80070 SUS2; SUS2 (ABNORMAL SUSPENSOR 2); K12856 pre-mRNA-processing
factor 8
Length=2382
Score = 917 bits (2371), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 421/538 (78%), Positives = 489/538 (90%), Gaps = 0/538 (0%)
Query 1 VHESIVMDLCQVFDMELDSLEVETVQKETIHPRKSYKMNSSCADILLFATYKWSISKPSL 60
+HES+VMDLCQV D ELD+LE+ETVQKETIHPRKSYKMNSSCAD+LLFA +KW +SKPSL
Sbjct 1632 IHESVVMDLCQVLDQELDALEIETVQKETIHPRKSYKMNSSCADVLLFAAHKWPMSKPSL 1691
Query 61 LAEGKDIMEGATATKHWLDVQLRWGDYDSHDIERYCRSKFLDYTTDNMSIYPSPSGVLVG 120
+AE KD+ + + K+W+DVQLRWGDYDSHDIERY R+KF+DYTTDNMSIYPSP+GV++G
Sbjct 1692 VAESKDMFDQKASNKYWIDVQLRWGDYDSHDIERYTRAKFMDYTTDNMSIYPSPTGVMIG 1751
Query 121 VDLAYNLHSAFGNWIPGLKPLMQRAMNKIMKANPALYVLRERIRKGLQLYSSEPTEPYLN 180
+DLAYNLHSAFGNW PG KPL+ +AMNKIMK+NPALYVLRERIRKGLQLYSSEPTEPYL+
Sbjct 1752 LDLAYNLHSAFGNWFPGSKPLLAQAMNKIMKSNPALYVLRERIRKGLQLYSSEPTEPYLS 1811
Query 181 SQNYGELFSNQTVWFIDDTNVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKIIHT 240
SQNYGE+FSNQ +WF+DDTNVYRVTIHKTFEGNLTTKP+NG IFIFNPRTGQLFLK+IHT
Sbjct 1812 SQNYGEIFSNQIIWFVDDTNVYRVTIHKTFEGNLTTKPINGAIFIFNPRTGQLFLKVIHT 1871
Query 241 SVWAGQKRLIQLARWKTAEEVAALIRSLPVEEQPKQIIATRKGMLDPLEVHLLDFPNIVI 300
SVWAGQKRL QLA+WKTAEEVAAL+RSLPVEEQPKQII TRKGMLDPLEVHLLDFPNIVI
Sbjct 1872 SVWAGQKRLGQLAKWKTAEEVAALVRSLPVEEQPKQIIVTRKGMLDPLEVHLLDFPNIVI 1931
Query 301 KGSELNLPFQALMRIEKFGDMILRATQPEMVLFNMFDDWLKSISAYTAFSRLLLLLRALQ 360
KGSEL LPFQA ++IEKFGD+IL+AT+P+MVLFN++DDWLKSIS+YTAFSRL+L+LRAL
Sbjct 1932 KGSELQLPFQACLKIEKFGDLILKATEPQMVLFNIYDDWLKSISSYTAFSRLILILRALH 1991
Query 361 VNTERAKVLLRPHKSVVTQPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVASLTQ 420
VN E+AK+LL+P KSVVT+PHHIWPSLTD++W+ VEV L+DLIL DYAKKNNVN ++LTQ
Sbjct 1992 VNNEKAKMLLKPDKSVVTEPHHIWPSLTDDQWMKVEVALRDLILSDYAKKNNVNTSALTQ 2051
Query 421 SEIRDIILGMEIAPPSLQRQQIAEIEAQAREQQQVTSTTTRSVNIHGEEMIVATQSPHEQ 480
SEIRDIILG EI PPS QRQQIAEIE QA+E Q+T+ TTR+ N+HG+E+IV T SP+EQ
Sbjct 2052 SEIRDIILGAEITPPSQQRQQIAEIEKQAKEASQLTAVTTRTTNVHGDELIVTTTSPYEQ 2111
Query 481 QVFSSKTDWRVRAISAASLHLRTRHLYVSSEAVAPAGCTYILPKNLLKKFISIADLRT 538
F SKTDWRVRAISA +L+LR H+YV+S+ + G TYI+PKN+LKKFI +ADLRT
Sbjct 2112 SAFGSKTDWRVRAISATNLYLRVNHIYVNSDDIKETGYTYIMPKNILKKFICVADLRT 2169
> cel:C50C3.6 prp-8; yeast PRP (splicing factor) related family
member (prp-8); K12856 pre-mRNA-processing factor 8
Length=2329
Score = 904 bits (2335), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 409/538 (76%), Positives = 491/538 (91%), Gaps = 0/538 (0%)
Query 1 VHESIVMDLCQVFDMELDSLEVETVQKETIHPRKSYKMNSSCADILLFATYKWSISKPSL 60
+HES+VMDLCQVFD ELD+LE++TVQKETIHPRKSYKMNSSCAD+LLFA YKW++S+PSL
Sbjct 1578 IHESVVMDLCQVFDQELDALEIQTVQKETIHPRKSYKMNSSCADVLLFAQYKWNVSRPSL 1637
Query 61 LAEGKDIMEGATATKHWLDVQLRWGDYDSHDIERYCRSKFLDYTTDNMSIYPSPSGVLVG 120
+A+ KD+M+ T K+WLDVQLRWGDYDSHD+ERY R+KFLDYTTDNMSIYPSP+GVL+
Sbjct 1638 MADSKDVMDNTTTQKYWLDVQLRWGDYDSHDVERYARAKFLDYTTDNMSIYPSPTGVLIA 1697
Query 121 VDLAYNLHSAFGNWIPGLKPLMQRAMNKIMKANPALYVLRERIRKGLQLYSSEPTEPYLN 180
+DLAYNL+SA+GNW PG+KPL+++AM KI+KANPA YVLRERIRKGLQLYSSEPTEPYL
Sbjct 1698 IDLAYNLYSAYGNWFPGMKPLIRQAMAKIIKANPAFYVLRERIRKGLQLYSSEPTEPYLT 1757
Query 181 SQNYGELFSNQTVWFIDDTNVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKIIHT 240
SQNYGELFSNQ +WF+DDTNVYRVTIHKTFEGNLTTKP+NG IFIFNPRTGQLFLKIIHT
Sbjct 1758 SQNYGELFSNQIIWFVDDTNVYRVTIHKTFEGNLTTKPINGAIFIFNPRTGQLFLKIIHT 1817
Query 241 SVWAGQKRLIQLARWKTAEEVAALIRSLPVEEQPKQIIATRKGMLDPLEVHLLDFPNIVI 300
SVWAGQKRL QLA+WKTAEEVAALIRSLPVEEQP+QII TRK MLDPLEVHLLDFPNIVI
Sbjct 1818 SVWAGQKRLSQLAKWKTAEEVAALIRSLPVEEQPRQIIVTRKAMLDPLEVHLLDFPNIVI 1877
Query 301 KGSELNLPFQALMRIEKFGDMILRATQPEMVLFNMFDDWLKSISAYTAFSRLLLLLRALQ 360
KGSEL LPFQA+M++EKFGD+IL+AT+P+MVLFN++DDWLK+IS+YTAFSR++L++R +
Sbjct 1878 KGSELMLPFQAIMKVEKFGDLILKATEPQMVLFNLYDDWLKTISSYTAFSRVVLIMRGMH 1937
Query 361 VNTERAKVLLRPHKSVVTQPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVASLTQ 420
+N ++ KV+L+P K+ +T+PHHIWP+L+D++WI VE+ LKD+IL DY KKNNVNVASLTQ
Sbjct 1938 INPDKTKVILKPDKTTITEPHHIWPTLSDDDWIKVELALKDMILADYGKKNNVNVASLTQ 1997
Query 421 SEIRDIILGMEIAPPSLQRQQIAEIEAQAREQQQVTSTTTRSVNIHGEEMIVATQSPHEQ 480
SE+RDIILGMEI+ PS QRQQIA+IE Q +EQ QVT+TTTR+VN HG+E+I AT S +E
Sbjct 1998 SEVRDIILGMEISAPSQQRQQIADIEKQTKEQSQVTATTTRTVNKHGDEIITATTSNYET 2057
Query 481 QVFSSKTDWRVRAISAASLHLRTRHLYVSSEAVAPAGCTYILPKNLLKKFISIADLRT 538
F+S+T+WRVRAIS+ +LHLRT+H+YV+S+ V G TYILPKN+LKKFI+I+DLRT
Sbjct 2058 ASFASRTEWRVRAISSTNLHLRTQHIYVNSDDVKDTGYTYILPKNILKKFITISDLRT 2115
> ath:AT4G38780 splicing factor, putative
Length=2332
Score = 899 bits (2322), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 412/538 (76%), Positives = 483/538 (89%), Gaps = 0/538 (0%)
Query 1 VHESIVMDLCQVFDMELDSLEVETVQKETIHPRKSYKMNSSCADILLFATYKWSISKPSL 60
+HES+VMDLCQV D EL+ LE+ETVQKETIHPRKSYKMNSSCAD+LLFA +KW +SKPSL
Sbjct 1584 IHESVVMDLCQVLDQELEPLEIETVQKETIHPRKSYKMNSSCADVLLFAAHKWPMSKPSL 1643
Query 61 LAEGKDIMEGATATKHWLDVQLRWGDYDSHDIERYCRSKFLDYTTDNMSIYPSPSGVLVG 120
+AE KD+ + + K+W+DVQLRWGDYDSHDIERY ++KF+DYTTDNMSIYPSP+GV++G
Sbjct 1644 IAESKDVFDQKASNKYWIDVQLRWGDYDSHDIERYTKAKFMDYTTDNMSIYPSPTGVIIG 1703
Query 121 VDLAYNLHSAFGNWIPGLKPLMQRAMNKIMKANPALYVLRERIRKGLQLYSSEPTEPYLN 180
+DLAYNLHSAFGNW PG KPL+ +AMNKIMK+NPALYVLRERIRKGLQLYSSEPTEPYL+
Sbjct 1704 LDLAYNLHSAFGNWFPGSKPLLAQAMNKIMKSNPALYVLRERIRKGLQLYSSEPTEPYLS 1763
Query 181 SQNYGELFSNQTVWFIDDTNVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKIIHT 240
SQNYGE+FSNQ +WF+DDTNVYRVTIHKTFEGNLTTKP+NGVIFIFNPRTGQLFLKIIHT
Sbjct 1764 SQNYGEIFSNQIIWFVDDTNVYRVTIHKTFEGNLTTKPINGVIFIFNPRTGQLFLKIIHT 1823
Query 241 SVWAGQKRLIQLARWKTAEEVAALIRSLPVEEQPKQIIATRKGMLDPLEVHLLDFPNIVI 300
SVWAGQKRL QLA+WKTAEEVAAL+RSLPVEEQPKQ+I TRKGMLDPLEVHLLDFPNIVI
Sbjct 1824 SVWAGQKRLGQLAKWKTAEEVAALVRSLPVEEQPKQVIVTRKGMLDPLEVHLLDFPNIVI 1883
Query 301 KGSELNLPFQALMRIEKFGDMILRATQPEMVLFNMFDDWLKSISAYTAFSRLLLLLRALQ 360
KGSEL LPFQA ++IEKFGD+IL+AT+P+M LFN++DDWL ++S+YTAF RL+L+LRAL
Sbjct 1884 KGSELQLPFQACLKIEKFGDLILKATEPQMALFNIYDDWLMTVSSYTAFQRLILILRALH 1943
Query 361 VNTERAKVLLRPHKSVVTQPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVASLTQ 420
VN E+AK+LL+P SVVT+P+HIWPSLTD++W+ VEV L+DLIL DYAKKN VN ++LTQ
Sbjct 1944 VNNEKAKMLLKPDMSVVTEPNHIWPSLTDDQWMKVEVALRDLILSDYAKKNKVNTSALTQ 2003
Query 421 SEIRDIILGMEIAPPSLQRQQIAEIEAQAREQQQVTSTTTRSVNIHGEEMIVATQSPHEQ 480
SEIRDIILG EI PPS QRQQIAEIE QA+E Q+T+ TTR+ N+HG+E+I T SP+EQ
Sbjct 2004 SEIRDIILGAEITPPSQQRQQIAEIEKQAKEASQLTAVTTRTTNVHGDELISTTISPYEQ 2063
Query 481 QVFSSKTDWRVRAISAASLHLRTRHLYVSSEAVAPAGCTYILPKNLLKKFISIADLRT 538
F SKTDWRVRAISA +L+LR H+YV+S+ + G TYI+PKN+LKKFI IADLRT
Sbjct 2064 SAFGSKTDWRVRAISATNLYLRVNHIYVNSDDIKETGYTYIMPKNILKKFICIADLRT 2121
> bbo:BBOV_IV007790 23.m06497; processing splicing factor 8; K12856
pre-mRNA-processing factor 8
Length=2343
Score = 855 bits (2210), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 399/538 (74%), Positives = 461/538 (85%), Gaps = 7/538 (1%)
Query 1 VHESIVMDLCQVFDMELDSLEVETVQKETIHPRKSYKMNSSCADILLFATYKWSISKPSL 60
+HES+VMDLCQV D+ELD+LE+ETVQKETIHPRKSYKMNSSCADILL A YKW + KPSL
Sbjct 1607 IHESLVMDLCQVLDLELDALEIETVQKETIHPRKSYKMNSSCADILLCAAYKWHVGKPSL 1666
Query 61 LAEGKDIMEGATATKHWLDVQLRWGDYDSHDIERYCRSKFLDYTTDNMSIYPSPSGVLVG 120
L +G + K W+DVQLRWGDYDSHDIERYCR+KFLDYTTD+MSIYP P+G L+G
Sbjct 1667 LTDGDSGDATTSTNKFWIDVQLRWGDYDSHDIERYCRAKFLDYTTDSMSIYPCPTGCLIG 1726
Query 121 VDLAYNLHSAFGNWIPGLKPLMQRAMNKIMKANPALYVLRERIRKGLQLYSSEPTEPYLN 180
VDLAYN+HS FG W PG+K L RAMNKIMKANPA++VLRERIRKGLQLYSSEPTEPYL+
Sbjct 1727 VDLAYNMHSGFGYWFPGMKTLCGRAMNKIMKANPAMFVLRERIRKGLQLYSSEPTEPYLS 1786
Query 181 SQNYGELFSNQTVWFIDDTNVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKIIHT 240
SQN+GELF +QT+WF+DDTNVYRVTIHKTFEGNLTTKPVNG IF+FNP+TGQLFLKIIHT
Sbjct 1787 SQNFGELFGSQTIWFVDDTNVYRVTIHKTFEGNLTTKPVNGAIFVFNPKTGQLFLKIIHT 1846
Query 241 SVWAGQKRLIQLARWKTAEEVAALIRSLPVEEQPKQIIATRKGMLDPLEVHLLDFPNIVI 300
SVWAGQKRL QLA+WKTAEEVAALIRSLPVEEQPKQII TRKGMLDPLEVHLLDFPNIVI
Sbjct 1847 SVWAGQKRLSQLAKWKTAEEVAALIRSLPVEEQPKQIIVTRKGMLDPLEVHLLDFPNIVI 1906
Query 301 KGSELNLPFQALMRIEKFGDMILRATQPEMVLFNMFDDWLKSISAYTAFSRLLLLLRALQ 360
KGSEL LP+Q++M++EKFGDMILRATQPEMVLFN++DDWLKSIS+YTAFSRL+L+LRA+
Sbjct 1907 KGSELQLPYQSIMKLEKFGDMILRATQPEMVLFNLYDDWLKSISSYTAFSRLILILRAMH 1966
Query 361 VNTERAKVLLRPHKSVVTQPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVASLTQ 420
VN +RAKVLL+P K+ VT PHH+WPSLTD EWI+ EV LKDLIL DYAK+N +N SLTQ
Sbjct 1967 VNCDRAKVLLKPSKTTVTLPHHVWPSLTDTEWINCEVALKDLILADYAKRNGINATSLTQ 2026
Query 421 SEIRDIILGMEIAPPSLQRQQIAEIEAQAREQQQVTSTTTRSVNIHGEEMIVATQSPHEQ 480
SEIRDIILGMEIAPP LQRQ++ E A + TTRSVN+HG+E+IV TQ+PHEQ
Sbjct 2027 SEIRDIILGMEIAPPDLQRQELEENRADVAAKM----VTTRSVNVHGDEIIVTTQTPHEQ 2082
Query 481 QVFSSKTDWRVRAISAASLHLRTRHLYVSSEAVAPAGCTYILPKNLLKKFISIADLRT 538
+VF+SKTDWR R ++A LH R L V S + T ++P NL++K + ++DLRT
Sbjct 2083 KVFASKTDWRNRCLAAGLLHRRLEGLSVES---VDSDDTLVIPLNLIRKLVMVSDLRT 2137
> tpv:TP03_0292 splicing factor Prp8; K12856 pre-mRNA-processing
factor 8
Length=2736
Score = 848 bits (2192), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 400/542 (73%), Positives = 467/542 (86%), Gaps = 16/542 (2%)
Query 1 VHESIVMDLCQVFDMELDSLEVETVQKETIHPRKSYKMNSSCADILLFATYKWSISKPSL 60
+HESIVMDLCQV DMELDSLE+ETVQKETIHPRKSYKMNSSCADIL+ ++YKW KPSL
Sbjct 2013 IHESIVMDLCQVLDMELDSLEIETVQKETIHPRKSYKMNSSCADILVTSSYKWRFGKPSL 2072
Query 61 LAEGKDIMEGATATKHWLDVQLRWGDYDSHDIERYCRSKFLDYTTDNMSIYPSPSGVLVG 120
L + I + TK+W+DVQLRWGDYDSHDIERY RSKFLDYT D+MSIYP P+G L+
Sbjct 2073 LTDTTPIEKLENGTKYWIDVQLRWGDYDSHDIERYSRSKFLDYTGDSMSIYPCPTGCLIA 2132
Query 121 VDLAYNLHSAFGNWIPGLKPLMQRAMNKIMKANPALYVLRERIRKGLQLYSSEPTEPYLN 180
VDLAYNLHS +G W G++ LM RAMNKIMKANPAL+VLRERIRK LQLYSSEPTEPYL+
Sbjct 2133 VDLAYNLHSGYGYWFEGMRELMVRAMNKIMKANPALFVLRERIRKSLQLYSSEPTEPYLS 2192
Query 181 SQNYGELFSNQTVWFIDDTNVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKIIHT 240
SQN GELF +QT+WF+DDTNVYRVTIHKTFEGNLTTKPVNG IFIFNP+TGQLFLK+IHT
Sbjct 2193 SQNMGELFGSQTIWFVDDTNVYRVTIHKTFEGNLTTKPVNGAIFIFNPKTGQLFLKVIHT 2252
Query 241 SVWAGQKRLIQLARWKTAEEVAALIRSLPVEEQPKQIIATRKGMLDPLEVHLLDFPNIVI 300
SVWAGQKRL QL++WKTAEEV ALIRSLPVEEQPKQII TR+GMLDPLEVHL+DFPNIVI
Sbjct 2253 SVWAGQKRLSQLSKWKTAEEVVALIRSLPVEEQPKQIIVTRRGMLDPLEVHLVDFPNIVI 2312
Query 301 KGSELNLPFQALMRIEKFGDMILRATQPEMVLFNMFDDWLKSISAYTAFSRLLLLLRALQ 360
KGSEL LP+Q++M++EKFGD+ILRATQPEMVLFN++DDWLKSIS+YTAFSRL+L+LRA+
Sbjct 2313 KGSELQLPYQSIMKLEKFGDLILRATQPEMVLFNLYDDWLKSISSYTAFSRLILILRAIH 2372
Query 361 VNTERAKVLLRPHKSVVTQPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVASLTQ 420
VN ERAK +L+P+K+ +T PHH+WP+LTD EWI+VE+ LKDLIL DYAK+N+V V SLTQ
Sbjct 2373 VNPERAKCILKPNKTTITLPHHVWPNLTDNEWINVEIALKDLILADYAKRNSVTVTSLTQ 2432
Query 421 SEIRDIILGMEIAPPSLQRQQIAE-IEAQAREQQQVTSTTTRSVNIHGEEMIVATQSPHE 479
+EIRDIILGMEIAPP +QRQQI E IE + S TT++VN+HG+E++V TQSPHE
Sbjct 2433 TEIRDIILGMEIAPPDVQRQQIEENIEVGPK------SVTTKTVNVHGDEIVVTTQSPHE 2486
Query 480 QQVFSSKTDWRVRAISAASLHLRTRHLY---VSSEAVAPAGCTYILPKNLLKKFISIADL 536
QQVF+SKTDWR R +++ +LHLR +H+Y V SE V +LP NL+KK ISIADL
Sbjct 2487 QQVFASKTDWRNRCLASGTLHLRAKHIYVVPVESEQVI------VLPNNLIKKLISIADL 2540
Query 537 RT 538
RT
Sbjct 2541 RT 2542
> cpv:cgd3_2890 Prp8. JAB/PAD domain ; K12856 pre-mRNA-processing
factor 8
Length=2379
Score = 829 bits (2142), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 385/557 (69%), Positives = 473/557 (84%), Gaps = 23/557 (4%)
Query 1 VHESIVMDLCQVFDMELDSLEVETVQKETIHPRKSYKMNSSCADILLFATYKWSISKPSL 60
+HESIVMD+CQV D E+D+L +E VQKE IHPRKSYKMNSSCADILL ++YKW + PSL
Sbjct 1592 IHESIVMDICQVLDNEVDALGIEMVQKEAIHPRKSYKMNSSCADILLLSSYKWVATNPSL 1651
Query 61 LAEGKDIMEGAT---ATKHWLDVQLRWGDYDSHDIERYCRSKFLDYTTDNMSIYPSPSGV 117
L + KD + + K W+D+QLRWGDYDSHDIERYCR+KFLDYT+D MSIYPSP+GV
Sbjct 1652 LLDKKDDISSNSLINTNKFWIDIQLRWGDYDSHDIERYCRAKFLDYTSDAMSIYPSPTGV 1711
Query 118 LVGVDLAYNLHSAFGNWIPGLKPLMQRAMNKIMKANPALYVLRERIRKGLQLYSSEPTEP 177
L+ VDLAYNL+SA+GNWIPGLK L+Q+AM KIMK+NPALYVLRERIRKGLQLY SEPTEP
Sbjct 1712 LIAVDLAYNLYSAYGNWIPGLKELIQKAMAKIMKSNPALYVLRERIRKGLQLYCSEPTEP 1771
Query 178 YLNSQNYGELFSNQTVWFIDDTNVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKI 237
YLNSQNY ELFSNQT WF+DDTNVYRV+IHKTFEGNLTTKPVNG I I NP G+LF+K+
Sbjct 1772 YLNSQNYNELFSNQTTWFVDDTNVYRVSIHKTFEGNLTTKPVNGCILILNPCNGKLFMKV 1831
Query 238 IHTSVWAGQKRLIQLARWKTAEEVAALIRSLPVEEQPKQIIATRKGMLDPLEVHLLDFPN 297
IHTSVWAGQKRL QLA+WKTAEEV ALIRSLP+EEQPKQ+I TRKGMLDPLEVHLLDFPN
Sbjct 1832 IHTSVWAGQKRLSQLAKWKTAEEVVALIRSLPIEEQPKQVIVTRKGMLDPLEVHLLDFPN 1891
Query 298 IVIKGSELNLPFQALMRIEKFGDMILRATQPEMVLFNMFDDWLKSISAYTAFSRLLLLLR 357
IVIKGS+L+LPFQAL++IEKFGD++L++TQP MVLF+++DDWLK+IS +TAFSRL+L+LR
Sbjct 1892 IVIKGSDLSLPFQALLKIEKFGDLVLKSTQPSMVLFSLYDDWLKTISPFTAFSRLVLILR 1951
Query 358 ALQVNTERAKVLLRPHKSVVTQPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVAS 417
++ +N ER KV+L+P+K+++T +HIWPSLTDEEW +VEV +KD+IL DY+KKNNV++++
Sbjct 1952 SMHINPERTKVILKPNKNIITMHNHIWPSLTDEEWANVEVAMKDIILDDYSKKNNVHISA 2011
Query 418 LTQSEIRDIILGMEIAPPSLQRQQIAEIEAQAR-------EQQQVTSTTTRSVNIHGEEM 470
LTQSEIRDIILGMEI PPS+QRQQIAEIE Q + E +TS TT++VN+HG+E+
Sbjct 2012 LTQSEIRDIILGMEITPPSIQRQQIAEIERQVKDLANKTTECSDITSLTTKTVNVHGQEI 2071
Query 471 IVATQSPHEQQVFSSKTDWRVRAISAASLHLRTRHLY---------VSSEAVAPAGCTYI 521
+V TQ+ +EQ+ FSSKTDWR RA+++ +L LR+ ++Y +S+ ++ P YI
Sbjct 2072 VVTTQTQYEQKTFSSKTDWRARALASTTLSLRSDNIYILSDDSILNISNNSIIP----YI 2127
Query 522 LPKNLLKKFISIADLRT 538
+PKNLLK FI I+DLRT
Sbjct 2128 IPKNLLKTFIEISDLRT 2144
> pfa:PFD0265w pre-mRNA splicing factor, putative; K12856 pre-mRNA-processing
factor 8
Length=3136
Score = 787 bits (2032), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 380/619 (61%), Positives = 471/619 (76%), Gaps = 82/619 (13%)
Query 1 VHESIVMDLCQVFDMELDSLEVETVQKETIHPRKSYKMNSSCADILLFATYKWSISKPSL 60
+HES+VMD+CQVFD+ D L++ETVQKETIHPRKSYKMNSSCADILLFA YKW ISKPSL
Sbjct 2241 IHESLVMDICQVFDLNCDLLDIETVQKETIHPRKSYKMNSSCADILLFANYKWGISKPSL 2300
Query 61 LAEGKDIMEGAT-----------------------------------------ATKHWLD 79
L + I T + + W+D
Sbjct 2301 LTDEDHIFTNNTLGSTSGTNNNIMLNSNMINSGSNNSSSNNMNSVSFGSFPYTSNQFWID 2360
Query 80 VQLRWGDYDSHDIERYCRSKFLDYTTDNMSIYPSPSGVLVGVDLAYNLHSAFGNWIPGLK 139
+QLRWGD+DSHDIERY R+KFLDYTTDN+SIYP +GVL+GVDLAYNL+SA+GNW LK
Sbjct 2361 IQLRWGDFDSHDIERYSRAKFLDYTTDNLSIYPCLTGVLIGVDLAYNLYSAYGNWFNNLK 2420
Query 140 PLMQRAMNKIMKANPALYVLRERIRKGLQLYSSEPTEPYLNSQNYGELFSNQTVWFIDDT 199
PLMQ+A+ KI+++NP+LYVLRERIRKGLQLYSSEPTEPYLN+QNY ELFS+QT+WF+DDT
Sbjct 2421 PLMQKALQKIVQSNPSLYVLRERIRKGLQLYSSEPTEPYLNTQNYNELFSSQTIWFVDDT 2480
Query 200 NVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKIIHTSVWAGQKRLIQLARWKTAE 259
NVYRVTIHKTFEGNLTTKP+NG IFI NP+TGQLFLKIIHTSVW GQKRL QLA+WKTAE
Sbjct 2481 NVYRVTIHKTFEGNLTTKPINGAIFILNPKTGQLFLKIIHTSVWIGQKRLSQLAKWKTAE 2540
Query 260 EVAALIRSLPVEEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLPFQALMRIEKFG 319
EVA+LIRSLP+EEQPKQII TRKGMLDPLEVHLLDFPNI+IKG+ELNLPFQAL+++ K G
Sbjct 2541 EVASLIRSLPIEEQPKQIIVTRKGMLDPLEVHLLDFPNIIIKGTELNLPFQALLKLNKIG 2600
Query 320 DMILRATQPEMVLFNMFDDWLKSISAYTAFSRLLLLLRALQVNTERAKVLLRPHKSVV-T 378
D+IL+ATQP+M+LFN++DDWL SIS++TAFSRL+L+LR+L +N ++ K+LL+P+K++V T
Sbjct 2601 DLILKATQPQMLLFNLYDDWLNSISSFTAFSRLILILRSLHINPQQTKILLQPNKNIVTT 2660
Query 379 QPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVASLTQSEIRDIILGMEIAPPSLQ 438
QPHHIWPS + +WIH+EV LKDLIL DY+K+NNV++ASLTQ+EIRDI+LGMEI PPS+Q
Sbjct 2661 QPHHIWPSFNNNQWIHLEVQLKDLILNDYSKRNNVHIASLTQNEIRDILLGMEITPPSIQ 2720
Query 439 RQQIAEIEAQARE--QQQVTSTTTRSVNIHGEEMIVATQSPHEQQVFSSKTDWRVRAISA 496
RQQIAE+E + +QQ+ TT+++ HG E+IV+T SPHEQQ F++KTDW++R ++
Sbjct 2721 RQQIAELEKNNLDLMEQQMKVTTSKTTTKHGNEIIVSTLSPHEQQTFTTKTDWKIRYLAN 2780
Query 497 ASLHLRTRHLYV-------------------SSEAVAPAGC------------------- 518
SL RT+++YV S + G
Sbjct 2781 NSLLFRTKNIYVNNNNMSNMSNINTISASASSHNILNKNGTNSDNQNSHYHTSINSINDY 2840
Query 519 TYILPKNLLKKFISIADLR 537
TY++ KNLL+KFI I+DL+
Sbjct 2841 TYVIAKNLLEKFICISDLK 2859
> sce:YHR165C PRP8, DBF3, DNA39, RNA8, SLT21, USA2; Component
of the U4/U6-U5 snRNP complex, involved in the second catalytic
step of splicing; mutations of human Prp8 cause retinitis
pigmentosa; K12856 pre-mRNA-processing factor 8
Length=2413
Score = 717 bits (1852), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 330/544 (60%), Positives = 424/544 (77%), Gaps = 7/544 (1%)
Query 1 VHESIVMDLCQVFDMELDSLEVETVQKETIHPRKSYKMNSSCADILLFATYKWSISKPSL 60
+HESIV D+CQ+ D ELD L++E+V KET+HPRKSYKMNSS ADI + + ++W +SKPSL
Sbjct 1657 IHESIVFDICQILDGELDVLQIESVTKETVHPRKSYKMNSSAADITMESVHEWEVSKPSL 1716
Query 61 LAEGKDIMEGATATKHWLDVQLRWGDYDSHDIERYCRSKFLDYTTDNMSIYPSPSGVLVG 120
L E D +G K W DVQLR+GDYDSHDI RY R+KFLDYTTDN+S+YPSP+GV++G
Sbjct 1717 LHETNDSFKGLITNKMWFDVQLRYGDYDSHDISRYVRAKFLDYTTDNVSMYPSPTGVMIG 1776
Query 121 VDLAYNLHSAFGNWIPGLKPLMQRAMNKIMKANPALYVLRERIRKGLQLYSSEPTEPYLN 180
+DLAYN++ A+GNW GLKPL+Q +M IMKANPALYVLRERIRKGLQ+Y S EP+LN
Sbjct 1777 IDLAYNMYDAYGNWFNGLKPLIQNSMRTIMKANPALYVLRERIRKGLQIYQSSVQEPFLN 1836
Query 181 SQNYGELFSNQTVWFIDDTNVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKIIHT 240
S NY ELF+N F+DDTNVYRVT+HKTFEGN+ TK +NG IF NP+TG LFLKIIHT
Sbjct 1837 SSNYAELFNNDIKLFVDDTNVYRVTVHKTFEGNVATKAINGCIFTLNPKTGHLFLKIIHT 1896
Query 241 SVWAGQKRLIQLARWKTAEEVAALIRSLPVEEQPKQIIATRKGMLDPLEVHLLDFPNIVI 300
SVWAGQKRL QLA+WKTAEEV+AL+RSLP EEQPKQII TRK MLDPLEVH+LDFPNI I
Sbjct 1897 SVWAGQKRLSQLAKWKTAEEVSALVRSLPKEEQPKQIIVTRKAMLDPLEVHMLDFPNIAI 1956
Query 301 KGSELNLPFQALMRIEKFGDMILRATQPEMVLFNMFDDWLKSISAYTAFSRLLLLLRALQ 360
+ +EL LPF A M I+K D++++AT+P+MVLFN++DDWL IS+YTAFSRL LLLRAL+
Sbjct 1957 RPTELRLPFSAAMSIDKLSDVVMKATEPQMVLFNIYDDWLDRISSYTAFSRLTLLLRALK 2016
Query 361 VNTERAKVLLRPHKSVVTQPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVASLTQ 420
N E AK++L ++ + +H+WPS TDE+WI +E ++DLIL +Y +K NVN+++LTQ
Sbjct 2017 TNEESAKMILLSDPTITIKSYHLWPSFTDEQWITIESQMRDLILTEYGRKYNVNISALTQ 2076
Query 421 SEIRDIILGMEIAPPSLQRQQIAEIEAQAREQQQ-------VTSTTTRSVNIHGEEMIVA 473
+EI+DIILG I PS++RQ++AE+EA E+Q T T+++N GEE++V
Sbjct 2077 TEIKDIILGQNIKAPSVKRQKMAELEAARSEKQNDEEAAGASTVMKTKTINAQGEEIVVV 2136
Query 474 TQSPHEQQVFSSKTDWRVRAISAASLHLRTRHLYVSSEAVAPAGCTYILPKNLLKKFISI 533
+ +E Q FSSK +WR AI+ L+LR +++YVS++ Y+LPKNLLKKFI I
Sbjct 2137 ASADYESQTFSSKNEWRKSAIANTLLYLRLKNIYVSADDFVEEQNVYVLPKNLLKKFIEI 2196
Query 534 ADLR 537
+D++
Sbjct 2197 SDVK 2200
> hsa:54677 CROT, COT; carnitine O-octanoyltransferase (EC:2.3.1.137);
K05940 carnitine O-octanoyltransferase [EC:2.3.1.137]
Length=640
Score = 35.8 bits (81), Expect = 0.48, Method: Compositional matrix adjust.
Identities = 30/135 (22%), Positives = 58/135 (42%), Gaps = 22/135 (16%)
Query 144 RAMNKIMKANPALYVLRERIRKGLQLYSSEPTEPYLNSQNYGE-----LFSNQTVWFIDD 198
+A ++ +P L E+I+ L +YS E + P++ ++Y E L + TV + D
Sbjct 278 KAREYLIGLDPENLALLEKIQSSLLVYSMEDSSPHVTPEDYSEIIAAILIGDPTVRWGDK 337
Query 199 TNVYRVTIHKTFEGNLTTKPVNGVIFIFNPRTGQLFLKIIHTSVWAGQKRLIQLARWKTA 258
+ + F N P + +I +++ S + +K RWK +
Sbjct 338 SYNLISFSNGVFGCNCDHAPFDAMI-------------MVNISYYVDEKIFQNEGRWKGS 384
Query 259 EEVAALIRSLPVEEQ 273
E+V R +P+ E+
Sbjct 385 EKV----RDIPLPEE 395
> mmu:667273 Vmn1r115, EG667273, Gm8549; vomeronasal 1 receptor
115
Length=295
Score = 34.7 bits (78), Expect = 0.87, Method: Compositional matrix adjust.
Identities = 23/102 (22%), Positives = 42/102 (41%), Gaps = 29/102 (28%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIK---------------------GSELNLPF 309
+++P+Q+I + M + L + L FPN +I NL
Sbjct 47 KQRPRQVILSHMAMANALTLFLTIFPNNMIAFAPKTPPTELKCKLEFFSHMVARSTNLCS 106
Query 310 QALMRIEKF--------GDMILRATQPEMVLFNMFDDWLKSI 343
++ I +F G +ILRA+ P+ V ++ + W S+
Sbjct 107 TCVLSIHQFVTLVLVNRGKLILRASVPKFVSYSCYSCWFFSV 148
> dre:100001804 novel protein containing lectin C-type domains
Length=328
Score = 34.3 bits (77), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 23/64 (35%), Positives = 30/64 (46%), Gaps = 17/64 (26%)
Query 91 DIERYCRSKFLDYTT-DNM--------SIYPSPSGVLVGVD--LAYNLHSAFG------N 133
D +RYCR K+ D T DNM S+ + GV +G+ AYN H + G N
Sbjct 36 DAQRYCREKYTDLATVDNMNDMIQLNKSVKVNDKGVWIGLQGTRAYNWHWSSGDPVLFLN 95
Query 134 WIPG 137
W G
Sbjct 96 WASG 99
> mmu:667464 Vmn1r142, EG667464, Gm8647; vomeronasal 1 receptor
142
Length=305
Score = 34.3 bits (77), Expect = 1.5, Method: Compositional matrix adjust.
Identities = 23/102 (22%), Positives = 42/102 (41%), Gaps = 29/102 (28%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIK---------------------GSELNLPF 309
+++P+Q+I + M + L + L FPN +I NL
Sbjct 47 KQRPRQVILSHMAMANALTLFLTIFPNNMIAFAPKTPPTELKCKLEFFSHMVARSTNLCS 106
Query 310 QALMRIEKF--------GDMILRATQPEMVLFNMFDDWLKSI 343
++ I +F G +ILRA+ P+ V ++ + W S+
Sbjct 107 TCVLSIHQFVTLVLVNRGKLILRASVPKFVSYSCYSCWFFSV 148
> mmu:621561 Vmn1r127, Gm6239; vomeronasal 1 receptor 127
Length=305
Score = 33.9 bits (76), Expect = 1.7, Method: Compositional matrix adjust.
Identities = 16/57 (28%), Positives = 31/57 (54%), Gaps = 0/57 (0%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLPFQALMRIEKFGDMILRATQ 327
+++P+Q+I + M + L + L FPN +I + P + ++E F M+ R+T
Sbjct 47 KQRPRQVILSHMAMANALTLFLTIFPNNMIAFAPKTPPTELKCKLEFFSHMVARSTN 103
> mmu:620537 Vmn1r94, EG620537, Gm6160; vomeronasal 1 receptor
94
Length=305
Score = 33.9 bits (76), Expect = 1.7, Method: Compositional matrix adjust.
Identities = 23/102 (22%), Positives = 42/102 (41%), Gaps = 29/102 (28%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIK---------------------GSELNLPF 309
+++P+Q+I + M + L + L FPN +I NL
Sbjct 47 KQRPRQVILSHMAMANALTLFLTIFPNNMIAFAPKTPPTELKCKLEFFSHMVARSTNLCS 106
Query 310 QALMRIEKF--------GDMILRATQPEMVLFNMFDDWLKSI 343
++ I +F G +ILRA+ P+ V ++ + W S+
Sbjct 107 TCVLSIHQFVTLVLVNRGKLILRASVPKFVSYSCYSCWFFSV 148
> mmu:100043025 Gm4177, Vmn1r134; predicted gene 4177
Length=305
Score = 33.9 bits (76), Expect = 1.7, Method: Compositional matrix adjust.
Identities = 16/57 (28%), Positives = 31/57 (54%), Gaps = 0/57 (0%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLPFQALMRIEKFGDMILRATQ 327
+++P+Q+I + M + L + L FPN +I + P + ++E F M+ R+T
Sbjct 47 KQRPRQVILSHMAMANALTLFLTIFPNNMIAFAPKTPPTELKCKLEFFSHMVARSTN 103
> mmu:100043083 Gm4216, Vmn1r162; predicted gene 4216
Length=305
Score = 33.9 bits (76), Expect = 1.7, Method: Compositional matrix adjust.
Identities = 16/57 (28%), Positives = 31/57 (54%), Gaps = 0/57 (0%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLPFQALMRIEKFGDMILRATQ 327
+++P+Q+I + M + L + L FPN +I + P + ++E F M+ R+T
Sbjct 47 KQRPRQVILSHMAMANALTLFLTIFPNNMIAFAPKTPPTELKCKLEFFSHMVARSTN 103
> mmu:235435 Lctl, E130104I05Rik, KLPH; lactase-like
Length=566
Score = 32.7 bits (73), Expect = 3.3, Method: Compositional matrix adjust.
Identities = 19/59 (32%), Positives = 33/59 (55%), Gaps = 11/59 (18%)
Query 468 EEMIVATQSPHEQQVFSSKTDWRVRAISAASLHLRT-------RHLYVSSEAVAPAGCT 519
+E+I A+ P+ Q+V S WR++A+ S++ + RH++V+SE V P C
Sbjct 494 KEIITASGFPNPQEVES----WRLKALETCSINNQMLATEPLLRHMHVASEIVVPTVCA 548
> xla:446240 lpcat4, agpat7, aytl3; lysophosphatidylcholine acyltransferase
4 (EC:2.3.1.51); K13512 lysophospholipid acyltransferase
[EC:2.3.1.23 2.3.1.-]
Length=522
Score = 32.7 bits (73), Expect = 4.1, Method: Compositional matrix adjust.
Identities = 16/53 (30%), Positives = 26/53 (49%), Gaps = 1/53 (1%)
Query 266 RSLPVEEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLP-FQALMRIEK 317
R P E P ++A DP+ + D P++V + LN+P AL+R +
Sbjct 115 RRAPASEAPILVVAPHSTFFDPIVTVVCDLPSVVSRVENLNIPVIGALLRFNQ 167
> mmu:435947 Vmn1r151, EG435947, Gm5727; vomeronasal 1 receptor
151
Length=305
Score = 32.0 bits (71), Expect = 5.8, Method: Compositional matrix adjust.
Identities = 15/57 (26%), Positives = 31/57 (54%), Gaps = 0/57 (0%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLPFQALMRIEKFGDMILRATQ 327
+++P+Q+I + + + L + L FPN +I + P + ++E F M+ R+T
Sbjct 47 KQRPRQVILSHMAVANALTLFLTIFPNNMIAFAPKTPPTELKCKLEFFSHMVARSTN 103
> mmu:435949 Gm5728, EG435949, Vmn1r99; predicted gene 5728
Length=305
Score = 32.0 bits (71), Expect = 5.9, Method: Compositional matrix adjust.
Identities = 15/57 (26%), Positives = 31/57 (54%), Gaps = 0/57 (0%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLPFQALMRIEKFGDMILRATQ 327
+++P+Q+I + + + L + L FPN +I + P + ++E F M+ R+T
Sbjct 47 KQRPRQVILSHMAVANALTLFLTIFPNNMIAFAPKTPPTELKCKLEFFSHMVARSTN 103
> mmu:667485 Gm8660, EG667485, Vmn1r147; predicted gene 8660
Length=305
Score = 32.0 bits (71), Expect = 5.9, Method: Compositional matrix adjust.
Identities = 15/57 (26%), Positives = 31/57 (54%), Gaps = 0/57 (0%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLPFQALMRIEKFGDMILRATQ 327
+++P+Q+I + + + L + L FPN +I + P + ++E F M+ R+T
Sbjct 47 KQRPRQVILSHMAVANALTLFLTIFPNNMIAFAPKTPPTELKCKLEFFSHMVARSTN 103
> mmu:667240 Vmn1r121, EG667240, Gm8533; vomeronasal 1 receptor
121
Length=314
Score = 32.0 bits (71), Expect = 6.0, Method: Compositional matrix adjust.
Identities = 23/91 (25%), Positives = 45/91 (49%), Gaps = 10/91 (10%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLPFQALMRIEKFGDMILRATQPEM 330
+++P+Q+I + + + L + L FPN ++ + N P + +++ F ++ R+T
Sbjct 47 KQRPRQVILSHMAVANALTLFLTVFPNNMMAFAPRNPPTELKCKLKFFSHLVTRST---- 102
Query 331 VLFNMFDDWLKSISAYTAF---SRLLLLLRA 358
N+ + SI + SR LLLRA
Sbjct 103 ---NLCSTCVLSIHQFVTLVPISRGKLLLRA 130
> mmu:667129 Vmn1r103, EG667129, Gm8474; vomeronasal 1 receptor
103
Length=305
Score = 32.0 bits (71), Expect = 6.0, Method: Compositional matrix adjust.
Identities = 15/57 (26%), Positives = 31/57 (54%), Gaps = 0/57 (0%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLPFQALMRIEKFGDMILRATQ 327
+++P+Q+I + + + L + L FPN +I + P + ++E F M+ R+T
Sbjct 47 KQRPRQVILSHMAVANALTLFLTIFPNNMIAFAPKTPPTELKCKLEFFSHMVARSTN 103
> mmu:670857 Vmn1r159, Gm16507; vomeronasal 1 receptor 159
Length=305
Score = 32.0 bits (71), Expect = 7.0, Method: Compositional matrix adjust.
Identities = 22/102 (21%), Positives = 42/102 (41%), Gaps = 29/102 (28%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIK---------------------GSELNLPF 309
+++P+Q+I + + + L + L FPN +I NL
Sbjct 47 KQRPRQVILSHMSVANALTLFLTIFPNNMIAFAPKTPPTELKCKLEFFSHMVARSTNLCS 106
Query 310 QALMRIEKF--------GDMILRATQPEMVLFNMFDDWLKSI 343
++ I +F G +ILRA+ P +V ++ + W S+
Sbjct 107 TCVLSIHQFVTLVLVNRGKLILRASVPNLVSYSCYGCWFFSV 148
> mmu:435951 Vmn1r122, EG435951, Gm5729; vomeronasal 1 receptor
122
Length=305
Score = 31.6 bits (70), Expect = 7.5, Method: Compositional matrix adjust.
Identities = 15/57 (26%), Positives = 31/57 (54%), Gaps = 0/57 (0%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLPFQALMRIEKFGDMILRATQ 327
+++P+Q+I + + + L + L FPN +I + P + ++E F M+ R+T
Sbjct 47 KQRPRQVILSHMSVANALTLFLTIFPNNMIAFAPKTPPTELKCKLEFFSHMVARSTN 103
> tpv:TP02_0096 chaperone protein DnaJ; K03686 molecular chaperone
DnaJ
Length=458
Score = 31.6 bits (70), Expect = 8.2, Method: Compositional matrix adjust.
Identities = 15/43 (34%), Positives = 25/43 (58%), Gaps = 5/43 (11%)
Query 379 QPHHIWPSLTDEEWIHVEVGLKDLILGDYAKKNNVNVASLTQS 421
QPHHI+ + D+ +HV + LK +LG N+++ +L S
Sbjct 333 QPHHIFKWIDDDIHVHVPISLKQCLLG-----GNISIPTLDGS 370
> mmu:243944 4930433I11Rik, Gm481; RIKEN cDNA 4930433I11 gene
Length=632
Score = 31.6 bits (70), Expect = 8.4, Method: Compositional matrix adjust.
Identities = 16/54 (29%), Positives = 33/54 (61%), Gaps = 2/54 (3%)
Query 282 KGMLDPLEVHLLDFPNIVIKGSELNLP--FQALMRIEKFGDMILRATQPEMVLF 333
KG LD L+V +DFP+I ++++LP F+ L ++++ D + ++ V++
Sbjct 141 KGTLDSLKVSTIDFPDITTLVADIHLPQLFKFLTGLDQYQDSTVTESKDSTVVW 194
> mmu:667599 Gm8720, EG667599, Vmn1r164; predicted gene 8720
Length=307
Score = 31.6 bits (70), Expect = 9.0, Method: Compositional matrix adjust.
Identities = 13/57 (22%), Positives = 32/57 (56%), Gaps = 0/57 (0%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLPFQALMRIEKFGDMILRATQ 327
+++P+Q+I + + + L + L FPN ++ + P + ++E F +++R+T
Sbjct 47 KQRPRQVILSHMAVANALTLFLTIFPNNMMAFAPKTPPTELKCKLESFSHLVVRSTN 103
> mmu:435940 Gm5725, EG435940, Vmn1r136; predicted gene 5725
Length=307
Score = 31.6 bits (70), Expect = 9.0, Method: Compositional matrix adjust.
Identities = 13/57 (22%), Positives = 32/57 (56%), Gaps = 0/57 (0%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLPFQALMRIEKFGDMILRATQ 327
+++P+Q+I + + + L + L FPN ++ + P + ++E F +++R+T
Sbjct 47 KQRPRQVILSHMAVANALTLFLTIFPNNMMAFAPKTPPTELKCKLESFSHLVVRSTN 103
> mmu:621510 Vmn1r129, EG621510, Gm6237; vomeronasal 1 receptor
129
Length=307
Score = 31.6 bits (70), Expect = 9.2, Method: Compositional matrix adjust.
Identities = 13/57 (22%), Positives = 32/57 (56%), Gaps = 0/57 (0%)
Query 271 EEQPKQIIATRKGMLDPLEVHLLDFPNIVIKGSELNLPFQALMRIEKFGDMILRATQ 327
+++P+Q+I + + + L + L FPN ++ + P + ++E F +++R+T
Sbjct 47 KQRPRQVILSHMAVANALTLFLTIFPNNMMAFAPKTPPTELKCKLESFSHLVVRSTN 103
Lambda K H
0.320 0.134 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 26317561200
Database: egene_temp_file_orthology_annotation_similarity_blast_database_866
Posted date: Sep 17, 2011 2:57 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40