bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_6193_orf1
Length=133
Score E
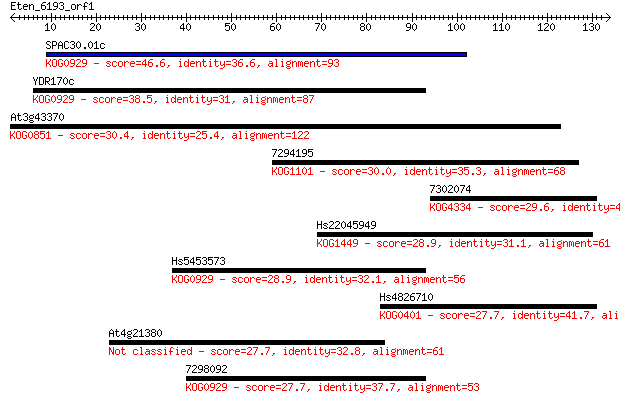
Sequences producing significant alignments: (Bits) Value
SPAC30.01c 46.6 1e-05
YDR170c 38.5 0.003
At3g43370 30.4 1.0
7294195 30.0 1.3
7302074 29.6 1.8
Hs22045949 28.9 2.7
Hs5453573 28.9 2.8
Hs4826710 27.7 5.6
At4g21380 27.7 6.5
7298092 27.7 6.8
> SPAC30.01c
Length=1822
Score = 46.6 bits (109), Expect = 1e-05, Method: Composition-based stats.
Identities = 34/97 (35%), Positives = 51/97 (52%), Gaps = 8/97 (8%)
Query 9 RCVAQLLLI----DLLRKEFIGNLLPVGALRLLVCCLEASFIFAYLYNQQVSLRLWLQRC 64
+C+ QLLLI +LL E + N +P + + + S+ FA +N+ SLR+ L
Sbjct 1624 KCILQLLLISIVAELLDNEEVFNHIPHEHVLKITVAIYDSWQFARKFNEDKSLRITLLNV 1683
Query 65 GFMAGLKQLPGLLKQHREAAAAAAALMLSALSETSNP 101
GFM KQLP LL+Q +A L+ L +T +P
Sbjct 1684 GFM---KQLPNLLRQETASALLYITLLFRLL-KTRDP 1716
> YDR170c
Length=2009
Score = 38.5 bits (88), Expect = 0.003, Method: Composition-based stats.
Identities = 27/91 (29%), Positives = 43/91 (47%), Gaps = 7/91 (7%)
Query 6 VAARCVAQLLLIDLLRKEF----IGNLLPVGALRLLVCCLEASFIFAYLYNQQVSLRLWL 61
+ +CV QLL+I+LL + F + +P + LE S+ F+ +N+ LR L
Sbjct 1818 IVVKCVLQLLMIELLNELFENEDFAHCIPYKEAIRITRLLEKSYEFSRDFNEDYGLRTRL 1877
Query 62 QRCGFMAGLKQLPGLLKQHREAAAAAAALML 92
+ ++P LLKQ AAA +M
Sbjct 1878 VEARV---VDKIPNLLKQETSAAAVLLDIMF 1905
> At3g43370
Length=235
Score = 30.4 bits (67), Expect = 1.0, Method: Compositional matrix adjust.
Identities = 31/123 (25%), Positives = 50/123 (40%), Gaps = 13/123 (10%)
Query 1 FDPQD-VAARCVAQLLLIDLLRKEFIGNLLPVGALRLLVCCLEASFIFAYLYNQQVSLRL 59
FDP D + + + +L+D+ +G L+ VGAL L+ SF YN+ +
Sbjct 118 FDPFDYILEKSAYKNVLVDV-----VGALVDVGALTTDYYGLKLSFEIKDRYNKVLECEA 172
Query 60 WLQRCGFMAGLKQLPGLLKQHREAAAAAAALMLSALSETSNPEKQQQEGAAAVAVSELAP 119
Q+ ++ G Q G AL L+E SNP+ + + V + P
Sbjct 173 RNQQAEYLDGYFQSLG-------KGNFVVALSFWRLTELSNPKLESHGAISKVVANPDRP 225
Query 120 AAA 122
A
Sbjct 226 EVA 228
> 7294195
Length=438
Score = 30.0 bits (66), Expect = 1.3, Method: Composition-based stats.
Identities = 24/78 (30%), Positives = 42/78 (53%), Gaps = 13/78 (16%)
Query 59 LWLQRCGFMAGLK-QL--------PGLLKQHREAAAAAAALMLSALSETSNPEKQQQEGA 109
LWL +C F+ +K QL P L ++ E+++ ++ S ++ E+QQ +
Sbjct 285 LWLSQCRFVKLMKGQLYIDTVAAKPVLAEEKEESSSIGG---VAVASTQASEEEQQTSLS 341
Query 110 AAVAVS-ELAPAAAPAAA 126
+ AVS ++AP+ AP AA
Sbjct 342 SEEAVSGDVAPSVAPTAA 359
> 7302074
Length=642
Score = 29.6 bits (65), Expect = 1.8, Method: Composition-based stats.
Identities = 17/38 (44%), Positives = 23/38 (60%), Gaps = 1/38 (2%)
Query 94 ALSETSNPEKQQQEGAAAVAVSELAPAAA-PAAAASAQ 130
AL E S E+Q+ +GA VAV +A A+A PA A +
Sbjct 210 ALEEESEVEQQKTDGAPPVAVCPMAGASALPAEAMDVE 247
> Hs22045949
Length=1444
Score = 28.9 bits (63), Expect = 2.7, Method: Composition-based stats.
Identities = 19/62 (30%), Positives = 31/62 (50%), Gaps = 1/62 (1%)
Query 69 GLKQLPGL-LKQHREAAAAAAALMLSALSETSNPEKQQQEGAAAVAVSELAPAAAPAAAA 127
GL Q PG L++ + ++ ++ L+ L + NPEK +EG A + +P A A
Sbjct 575 GLSQEPGAHLEEKKTPESSLSSQHLNELEKRPNPEKVVEEGREAGEMESSTLQESPRARA 634
Query 128 SA 129
A
Sbjct 635 EA 636
> Hs5453573
Length=1785
Score = 28.9 bits (63), Expect = 2.8, Method: Composition-based stats.
Identities = 18/56 (32%), Positives = 25/56 (44%), Gaps = 2/56 (3%)
Query 37 LVCCLEASFIFAYLYNQQVSLRLWLQRCGFMAGLKQLPGLLKQHREAAAAAAALML 92
L+ CL+ S F+ +N R L R GF K P LLKQ + A ++
Sbjct 1614 LLDCLQESHSFSKAFNSNYEQRTVLWRAGFKG--KSKPNLLKQETSSLACCLRILF 1667
> Hs4826710
Length=1396
Score = 27.7 bits (60), Expect = 5.6, Method: Compositional matrix adjust.
Identities = 20/51 (39%), Positives = 29/51 (56%), Gaps = 6/51 (11%)
Query 83 AAAAAAALMLSALSETSNPEKQQQEGAAAVA---VSELAPAAAPAAAASAQ 130
A + A+ LS +SE PE+Q +E A+VA + PA AP+A + AQ
Sbjct 210 GAPSPPAVDLSPVSE---PEEQAKEVTASVAPPTIPSATPATAPSATSPAQ 257
> At4g21380
Length=850
Score = 27.7 bits (60), Expect = 6.5, Method: Composition-based stats.
Identities = 20/74 (27%), Positives = 33/74 (44%), Gaps = 15/74 (20%)
Query 23 EFIGNLLPVGALRLLVCCLEAS---FIFAYLYNQQVSLRL----------WLQRCGFMAG 69
+ I L + +RLL CC++A I+ YL N + L W R + G
Sbjct 572 KLIARLQHINLVRLLACCVDAGEKMLIYEYLENLSLDSHLFDKSRNSKLNWQMRFDIING 631
Query 70 LKQLPGLLKQHREA 83
+ + GLL H+++
Sbjct 632 IAR--GLLYLHQDS 643
> 7298092
Length=1643
Score = 27.7 bits (60), Expect = 6.8, Method: Compositional matrix adjust.
Identities = 20/53 (37%), Positives = 25/53 (47%), Gaps = 4/53 (7%)
Query 40 CLEASFIFAYLYNQQVSLRLWLQRCGFMAGLKQLPGLLKQHREAAAAAAALML 92
CL S FA +N R L R GF +K P LLKQ E ++ A L +
Sbjct 1488 CLMQSHRFAKRFNADHDQRSLLWRAGFKGSVK--PNLLKQ--ETSSLACVLRI 1536
Lambda K H
0.323 0.132 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1356426142
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40