bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_5498_orf1
Length=93
Score E
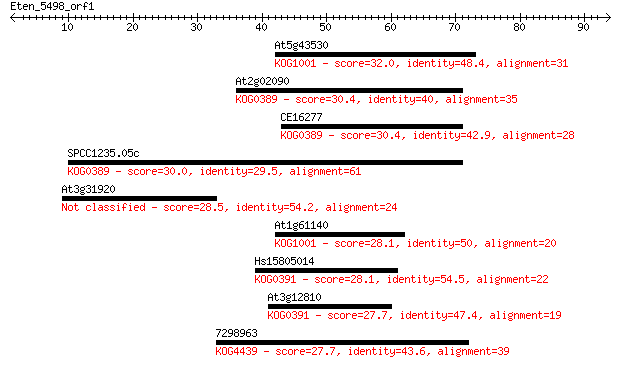
Sequences producing significant alignments: (Bits) Value
At5g43530 32.0 0.28
At2g02090 30.4 0.81
CE16277 30.4 0.85
SPCC1235.05c 30.0 0.97
At3g31920 28.5 3.0
At1g61140 28.1 3.7
Hs15805014 28.1 4.5
At3g12810 27.7 4.8
7298963 27.7 5.0
> At5g43530
Length=1277
Score = 32.0 bits (71), Expect = 0.28, Method: Composition-based stats.
Identities = 15/34 (44%), Positives = 20/34 (58%), Gaps = 3/34 (8%)
Query 42 LLFRLRRICNHSLLMQGRYTSEQK---DRMARHY 72
LL RLR+ CNH L+ R S+Q D +AR +
Sbjct 974 LLLRLRQCCNHPFLVMSRADSQQYADLDSLARRF 1007
> At2g02090
Length=763
Score = 30.4 bits (67), Expect = 0.81, Method: Composition-based stats.
Identities = 14/35 (40%), Positives = 22/35 (62%), Gaps = 0/35 (0%)
Query 36 KRFVSSLLFRLRRICNHSLLMQGRYTSEQKDRMAR 70
KR +S+ + R+I NH LL++ Y+ E R+AR
Sbjct 500 KRQISNYFTQFRKIANHPLLIRRIYSDEDVIRIAR 534
> CE16277
Length=1038
Score = 30.4 bits (67), Expect = 0.85, Method: Compositional matrix adjust.
Identities = 12/28 (42%), Positives = 19/28 (67%), Gaps = 0/28 (0%)
Query 43 LFRLRRICNHSLLMQGRYTSEQKDRMAR 70
L RLR+ NH LL + YT ++ D++A+
Sbjct 673 LMRLRQAANHPLLRRSEYTDQKLDKIAK 700
> SPCC1235.05c
Length=1284
Score = 30.0 bits (66), Expect = 0.97, Method: Composition-based stats.
Identities = 18/62 (29%), Positives = 31/62 (50%), Gaps = 1/62 (1%)
Query 10 AAQKGDTGSSGQQANEGDSREPEAGGKRFVSS-LLFRLRRICNHSLLMQGRYTSEQKDRM 68
A QK + N+ + E+ GK + +L +LR+ NH+LL + Y E+ +M
Sbjct 811 ALQKNQQLRRDDKRNKRSKNDEESDGKSLSAGHVLMQLRKAANHALLFRKFYDDEKLKQM 870
Query 69 AR 70
A+
Sbjct 871 AK 872
> At3g31920
Length=725
Score = 28.5 bits (62), Expect = 3.0, Method: Composition-based stats.
Identities = 13/24 (54%), Positives = 14/24 (58%), Gaps = 0/24 (0%)
Query 9 TAAQKGDTGSSGQQANEGDSREPE 32
TA GD S Q NE DS+EPE
Sbjct 103 TAYFDGDVHSDSQNQNEDDSKEPE 126
> At1g61140
Length=1287
Score = 28.1 bits (61), Expect = 3.7, Method: Composition-based stats.
Identities = 10/20 (50%), Positives = 15/20 (75%), Gaps = 0/20 (0%)
Query 42 LLFRLRRICNHSLLMQGRYT 61
+L RLR+ C+H LL+ G Y+
Sbjct 951 MLLRLRQACDHPLLVNGEYS 970
> Hs15805014
Length=3124
Score = 28.1 bits (61), Expect = 4.5, Method: Composition-based stats.
Identities = 12/22 (54%), Positives = 17/22 (77%), Gaps = 0/22 (0%)
Query 39 VSSLLFRLRRICNHSLLMQGRY 60
V S+L RL+RICNH L++ R+
Sbjct 1321 VLSILVRLQRICNHPGLVEPRH 1342
> At3g12810
Length=1048
Score = 27.7 bits (60), Expect = 4.8, Method: Composition-based stats.
Identities = 9/19 (47%), Positives = 15/19 (78%), Gaps = 0/19 (0%)
Query 41 SLLFRLRRICNHSLLMQGR 59
S++ +LR++CNH L +GR
Sbjct 395 SIIMQLRKVCNHPDLFEGR 413
> 7298963
Length=1061
Score = 27.7 bits (60), Expect = 5.0, Method: Composition-based stats.
Identities = 17/44 (38%), Positives = 21/44 (47%), Gaps = 5/44 (11%)
Query 33 AGGKRFVSS-----LLFRLRRICNHSLLMQGRYTSEQKDRMARH 71
AG K+ V S LL RLR+IC H L+ E+ M H
Sbjct 777 AGSKKEVKSHDILVLLLRLRQICCHPGLIDAMLDGEESQTMGDH 820
Lambda K H
0.314 0.131 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1167934574
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40