bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
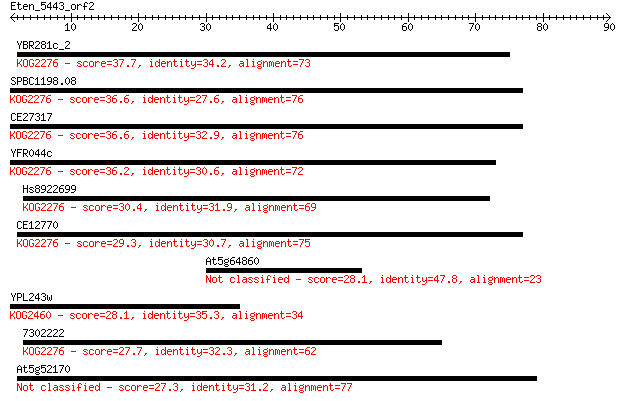
Query= Eten_5443_orf2
Length=89
Score E
Sequences producing significant alignments: (Bits) Value
YBR281c_2 37.7 0.005
SPBC1198.08 36.6 0.010
CE27317 36.6 0.011
YFR044c 36.2 0.016
Hs8922699 30.4 0.82
CE12770 29.3 1.7
At5g64860 28.1 4.4
YPL243w 28.1 4.7
7302222 27.7 4.8
At5g52170 27.3 7.4
> YBR281c_2
Length=452
Score = 37.7 bits (86), Expect = 0.005, Method: Composition-based stats.
Identities = 25/73 (34%), Positives = 34/73 (46%), Gaps = 14/73 (19%)
Query 2 VREGGSMPAIPFLADAFRRLPGPLGQCQGSDEVAVCQLPFGQASDNAHLPNERLGIRQVE 61
VREGGS+ + L F D AV Q+P GQ++DN HL NE L I+
Sbjct 394 VREGGSISCLRMLERIF-------------DAPAV-QIPCGQSTDNGHLANENLRIKNWS 439
Query 62 KGVDVLACTLESL 74
++L+ L
Sbjct 440 NLTEILSKVFNRL 452
> SPBC1198.08
Length=474
Score = 36.6 bits (83), Expect = 0.010, Method: Composition-based stats.
Identities = 21/76 (27%), Positives = 33/76 (43%), Gaps = 14/76 (18%)
Query 1 YVREGGSMPAIPFLADAFRRLPGPLGQCQGSDEVAVCQLPFGQASDNAHLPNERLGIRQV 60
+VREGGS+P + ++ V LP G+ D AH NE+L +
Sbjct 410 FVREGGSIPVTVTFEQSLKK--------------NVLLLPMGRGDDGAHSINEKLDLDNF 455
Query 61 EKGVDVLACTLESLAA 76
KG+ + + LA+
Sbjct 456 LKGIKLFCTYVHELAS 471
> CE27317
Length=461
Score = 36.6 bits (83), Expect = 0.011, Method: Composition-based stats.
Identities = 25/76 (32%), Positives = 33/76 (43%), Gaps = 14/76 (18%)
Query 1 YVREGGSMPAIPFLADAFRRLPGPLGQCQGSDEVAVCQLPFGQASDNAHLPNERLGIRQV 60
+ REGGS+P + D + V LP G + D AH NE++
Sbjct 399 FTREGGSIPVTLTIQDLTKS--------------PVMLLPIGASDDMAHSQNEKINRDNF 444
Query 61 EKGVDVLACTLESLAA 76
KG+ VLA L LAA
Sbjct 445 VKGMKVLAAYLFELAA 460
> YFR044c
Length=481
Score = 36.2 bits (82), Expect = 0.016, Method: Composition-based stats.
Identities = 22/72 (30%), Positives = 30/72 (41%), Gaps = 14/72 (19%)
Query 1 YVREGGSMPAIPFLADAFRRLPGPLGQCQGSDEVAVCQLPFGQASDNAHLPNERLGIRQV 60
+ REGGS+P DA +V LP G+ D AH NE+L I
Sbjct 416 FTREGGSIPITLTFQDAL--------------NTSVLLLPMGRGDDGAHSINEKLDISNF 461
Query 61 EKGVDVLACTLE 72
G+ +A L+
Sbjct 462 VGGMKTMAAYLQ 473
> Hs8922699
Length=475
Score = 30.4 bits (67), Expect = 0.82, Method: Composition-based stats.
Identities = 22/69 (31%), Positives = 28/69 (40%), Gaps = 14/69 (20%)
Query 3 REGGSMPAIPFLADAFRRLPGPLGQCQGSDEVAVCQLPFGQASDNAHLPNERLGIRQVEK 62
REGGS+P +A + V LP G A D AH NE+L +
Sbjct 413 REGGSIPVTLTFQEATGK--------------NVMLLPVGSADDGAHSQNEKLNRYNYIE 458
Query 63 GVDVLACTL 71
G +LA L
Sbjct 459 GTKMLAAYL 467
> CE12770
Length=473
Score = 29.3 bits (64), Expect = 1.7, Method: Composition-based stats.
Identities = 23/75 (30%), Positives = 32/75 (42%), Gaps = 14/75 (18%)
Query 2 VREGGSMPAIPFLADAFRRLPGPLGQCQGSDEVAVCQLPFGQASDNAHLPNERLGIRQVE 61
+REG S+P + F+ L G +V LP G A D AH NE+ I
Sbjct 412 IREGCSIP----ITLTFQELTGK----------SVLLLPIGAADDMAHSQNEKNNIWNYV 457
Query 62 KGVDVLACTLESLAA 76
+GV L + L +
Sbjct 458 EGVKTLLAYIMELGS 472
> At5g64860
Length=576
Score = 28.1 bits (61), Expect = 4.4, Method: Composition-based stats.
Identities = 11/23 (47%), Positives = 15/23 (65%), Gaps = 0/23 (0%)
Query 30 GSDEVAVCQLPFGQASDNAHLPN 52
G+ +AV Q FG +DN HLP+
Sbjct 437 GAPGMAVLQFAFGGGADNPHLPH 459
> YPL243w
Length=599
Score = 28.1 bits (61), Expect = 4.7, Method: Composition-based stats.
Identities = 12/34 (35%), Positives = 18/34 (52%), Gaps = 0/34 (0%)
Query 1 YVREGGSMPAIPFLADAFRRLPGPLGQCQGSDEV 34
Y +G M A+ DA+RRL L + + DE+
Sbjct 427 YQSKGRYMEALALYVDAYRRLENKLSEIESLDEI 460
> 7302222
Length=462
Score = 27.7 bits (60), Expect = 4.8, Method: Composition-based stats.
Identities = 20/62 (32%), Positives = 27/62 (43%), Gaps = 14/62 (22%)
Query 3 REGGSMPAIPFLADAFRRLPGPLGQCQGSDEVAVCQLPFGQASDNAHLPNERLGIRQVEK 62
REGGS+P L +A G + + V P G D AH NE++ I +
Sbjct 414 REGGSIPVTLTLQEA-----------TGKNVILV---PVGACDDGAHSQNEKIDIYNYIE 459
Query 63 GV 64
GV
Sbjct 460 GV 461
> At5g52170
Length=682
Score = 27.3 bits (59), Expect = 7.4, Method: Composition-based stats.
Identities = 24/88 (27%), Positives = 32/88 (36%), Gaps = 13/88 (14%)
Query 2 VREGGSMPAIPFLADAFRRLPGPLGQCQGSD-----------EVAVCQLPFGQASDNAHL 50
V GG + L F LP G SD E C L G L
Sbjct 593 VMSGGDSAYVALLPSGFSILPD--GSSSSSDQFDTDGGLVNQESKGCLLTVGFQILVNSL 650
Query 51 PNERLGIRQVEKGVDVLACTLESLAARL 78
P +L + VE +++ACT+ + A L
Sbjct 651 PTAKLNVESVETVNNLIACTIHKIRAAL 678
Lambda K H
0.320 0.136 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1181033294
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40