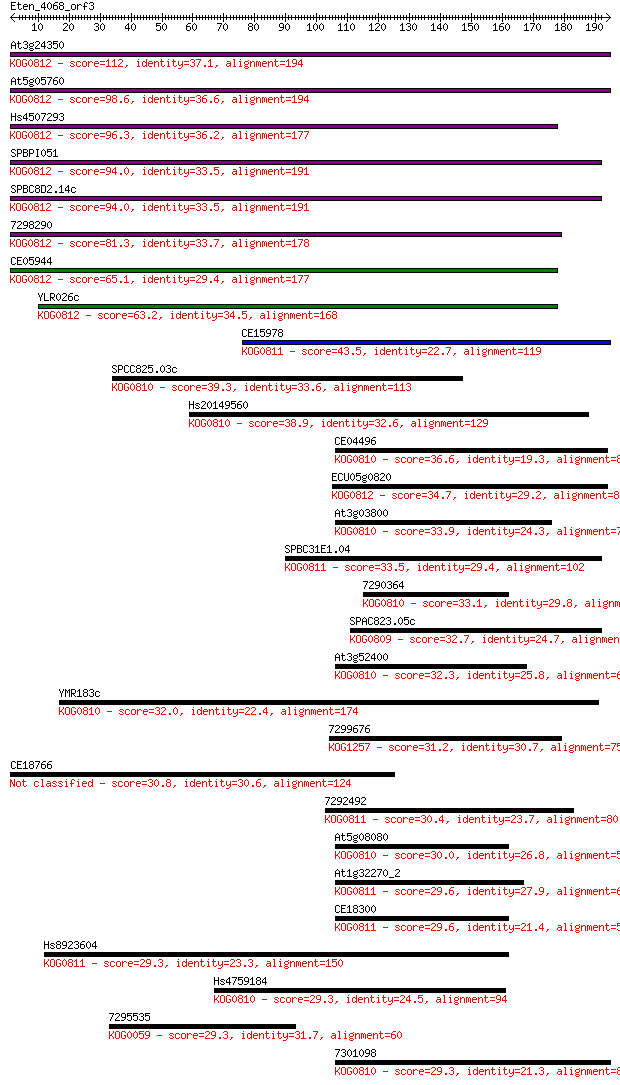
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_4068_orf3
Length=194
Score E
Sequences producing significant alignments: (Bits) Value
At3g24350 112 7e-25
At5g05760 98.6 8e-21
Hs4507293 96.3 4e-20
SPBPI051 94.0 2e-19
SPBC8D2.14c 94.0 2e-19
7298290 81.3 1e-15
CE05944 65.1 8e-11
YLR026c 63.2 4e-10
CE15978 43.5 2e-04
SPCC825.03c 39.3 0.005
Hs20149560 38.9 0.006
CE04496 36.6 0.034
ECU05g0820 34.7 0.12
At3g03800 33.9 0.19
SPBC31E1.04 33.5 0.25
7290364 33.1 0.37
SPAC823.05c 32.7 0.44
At3g52400 32.3 0.61
YMR183c 32.0 0.73
7299676 31.2 1.3
CE18766 30.8 2.0
7292492 30.4 2.1
At5g08080 30.0 2.7
At1g32270_2 29.6 3.8
CE18300 29.6 4.0
Hs8923604 29.3 5.0
Hs4759184 29.3 5.3
7295535 29.3 5.4
7301098 29.3 5.6
> At3g24350
Length=377
Score = 112 bits (279), Expect = 7e-25, Method: Compositional matrix adjust.
Identities = 72/245 (29%), Positives = 124/245 (50%), Gaps = 51/245 (20%)
Query 1 IRKTINTLNCKVDTLEKAAAAAGVSGQSKQ------HYITVTEVLKTRLLDVTKEFKDIL 54
I++ I+ LN + L+ ++ G + + H TV + LK RL+D TKEFKD+L
Sbjct 133 IKQEISALNSALVDLQLFRSSQNDEGNNSRDRDKSTHSATVVDDLKYRLMDTTKEFKDVL 192
Query 55 MLRTENLKRQDKRRNLYS----------------FGGDSGTAEAEEMYDIEG-------- 90
+RTEN+K + RR L+S + +E+ + G
Sbjct 193 TMRTENMKVHESRRQLFSSNASKESTNPFVRQRPLAAKAAASESVPLPWANGSSSSSSQL 252
Query 91 -------GQRTTLVSRPAT--------------AYSQARAEAVENVQRVIGELASIFQRV 129
G+ + L+ + Y Q RAEA+ V+ I EL+SIF ++
Sbjct 253 VPWKPGEGESSPLLQQSQQQQQQQQQQMVPLQDTYMQGRAEALHTVESTIHELSSIFTQL 312
Query 130 ATMIAHQEEMIQRIGQDIDTSMYHIREGQNELLNFYNRISSNRSLIIKVFLLLMAFILFF 189
ATM++ Q E+ RI Q+++ ++ ++ Q++L + N ISSNR L++K+F +L+AF++ F
Sbjct 313 ATMVSQQGEIAIRIDQNMEDTLANVEGAQSQLARYLNSISSNRWLMMKIFFVLIAFLMIF 372
Query 190 VFFLS 194
+FF++
Sbjct 373 LFFVA 377
> At5g05760
Length=336
Score = 98.6 bits (244), Expect = 8e-21, Method: Compositional matrix adjust.
Identities = 71/245 (28%), Positives = 119/245 (48%), Gaps = 51/245 (20%)
Query 1 IRKTINTLN---CKVDTLEKAAAAAGVSGQSK-QHYITVTEVLKTRLLDVTKEFKDILML 56
IR I LN + TL+ A G Q + HY V + LKTRL+ TK+ +D+L
Sbjct 92 IRNDITGLNMALSDLQTLQNMELADGNYSQDQVGHYTAVCDDLKTRLMGATKQLQDVLTT 151
Query 57 RTENLKRQDKRRNLYSFGG--DS----------------------GTAEAEEMYDIEGG- 91
R+EN+K + R+ L+S DS G + + + G
Sbjct 152 RSENMKAHENRKQLFSTKNAVDSPPQNNAKSVPEPPPWSSSSNPFGNLQQPLLPPLNTGA 211
Query 92 -------QRTTLVSRPATA---------------YSQARAEAVENVQRVIGELASIFQRV 129
+R+ + + P+ YSQ+RA A+ +V+ I EL+ IF ++
Sbjct 212 PPGSQLRRRSAIENAPSQQMEMSLLQQTVPKQENYSQSRAVALHSVESRITELSGIFPQL 271
Query 130 ATMIAHQEEMIQRIGQDIDTSMYHIREGQNELLNFYNRISSNRSLIIKVFLLLMAFILFF 189
ATM+ Q E+ RI ++D S+ ++ ++ LL RISSNR L++K+F +++ F++ F
Sbjct 272 ATMVTQQGELAIRIDDNMDESLVNVEGARSALLQHLTRISSNRWLMMKIFAVIILFLIVF 331
Query 190 VFFLS 194
+FF++
Sbjct 332 LFFVA 336
> Hs4507293
Length=301
Score = 96.3 bits (238), Expect = 4e-20, Method: Compositional matrix adjust.
Identities = 64/201 (31%), Positives = 109/201 (54%), Gaps = 25/201 (12%)
Query 1 IRKTINTLNCKVDTLEKAAAAAGV-SGQSKQ-HYITVTEVLKTRLLDVTKEFKDILMLRT 58
I++ IN+LN ++ L+ A G SG+ Q H T+ L+++L ++ +FK +L +RT
Sbjct 85 IKQDINSLNKQIAQLQDFVRAKGSQSGRHLQTHSNTIVVSLQSKLASMSNDFKSVLEVRT 144
Query 59 ENLKRQDKRRNLYS-------------FGGDSGTAEAE---------EMYDIEGGQRTTL 96
ENLK+Q RR +S GG + AE +M D Q+ L
Sbjct 145 ENLKQQRSRREQFSRAPVSALPLAPNHLGGGAVVLGAESHASKDVAIDMMDSRTSQQLQL 204
Query 97 VSRPATAYSQARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIRE 156
+ +Y Q+RA+ ++N++ I EL SIFQ++A M+ QEE IQRI +++ + +
Sbjct 205 IDE-QDSYIQSRADTMQNIESTIVELGSIFQQLAHMVKEQEETIQRIDENVLGAQLDVEA 263
Query 157 GQNELLNFYNRISSNRSLIIK 177
+E+L ++ ++SNR L++K
Sbjct 264 AHSEILKYFQSVTSNRWLMVK 284
> SPBPI051
Length=309
Score = 94.0 bits (232), Expect = 2e-19, Method: Compositional matrix adjust.
Identities = 64/220 (29%), Positives = 106/220 (48%), Gaps = 29/220 (13%)
Query 1 IRKTINTLNCKVDTLEKAAAAA-GVSGQSKQHYITVTEVLKTRLLDVTKEFKDILMLRTE 59
I++++++LN + +L++ Q QH V L+ L + + FKDIL +RT+
Sbjct 88 IKQSLSSLNSDIASLQQVVKGNRNKPAQMNQHSENVVVSLQNSLANTSMTFKDILEIRTQ 147
Query 60 NLKRQDKRR-----------NLYSFGGDS---------GTAEAEEMY---DIEGGQRT-- 94
N+K R N G+S EA E Y ++ G T
Sbjct 148 NMKASQNRTEKFVASSSMNANPLINSGNSISPFADYNDPKPEANEDYLSLNLGDGANTRY 207
Query 95 ---TLVSRPATAYSQARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSM 151
L+ YSQ R +++N++ I EL IF ++A M++ Q E +QRI D +
Sbjct 208 EQMALLESQTDTYSQQRMSSIQNIESTITELGGIFSQLAQMVSEQRETVQRIDMHTDDIV 267
Query 152 YHIREGQNELLNFYNRISSNRSLIIKVFLLLMAFILFFVF 191
+I Q E++ FY R+SSNR+L+ K+F +++ F L +V
Sbjct 268 SNIGSAQREIVKFYERMSSNRALLFKIFGIVIIFFLLWVL 307
> SPBC8D2.14c
Length=309
Score = 94.0 bits (232), Expect = 2e-19, Method: Compositional matrix adjust.
Identities = 64/220 (29%), Positives = 106/220 (48%), Gaps = 29/220 (13%)
Query 1 IRKTINTLNCKVDTLEKAAAAA-GVSGQSKQHYITVTEVLKTRLLDVTKEFKDILMLRTE 59
I++++++LN + +L++ Q QH V L+ L + + FKDIL +RT+
Sbjct 88 IKQSLSSLNSDIASLQQVVKGNRNKPAQMNQHSENVVVSLQNSLANTSMTFKDILEIRTQ 147
Query 60 NLKRQDKRR-----------NLYSFGGDS---------GTAEAEEMY---DIEGGQRT-- 94
N+K R N G+S EA E Y ++ G T
Sbjct 148 NMKASQNRTEKFVASSSMNANPLINSGNSISPFADYNDPKPEANEDYLSLNLGDGANTRY 207
Query 95 ---TLVSRPATAYSQARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSM 151
L+ YSQ R +++N++ I EL IF ++A M++ Q E +QRI D +
Sbjct 208 EQMALLESQTDTYSQQRMSSIQNIESTITELGGIFSQLAQMVSEQRETVQRIDMHTDDIV 267
Query 152 YHIREGQNELLNFYNRISSNRSLIIKVFLLLMAFILFFVF 191
+I Q E++ FY R+SSNR+L+ K+F +++ F L +V
Sbjct 268 SNIGSAQREIVKFYERMSSNRALLFKIFGIVIIFFLLWVL 307
> 7298290
Length=310
Score = 81.3 bits (199), Expect = 1e-15, Method: Compositional matrix adjust.
Identities = 60/207 (28%), Positives = 102/207 (49%), Gaps = 29/207 (14%)
Query 1 IRKTINTLNCKVDTLEKAAAAAGVSGQSKQ---HYITVTEVLKTRLLDVTKEFKDILMLR 57
I+ +N LN ++ L+ + K H + L+++L ++ +FK IL +R
Sbjct 88 IKGDLNALNQQIARLQDISKDQRRHTNGKHLVSHSSNMVLALQSKLASMSTDFKQILEVR 147
Query 58 TENLKRQDKRRNLYSFGG--------DSGTAE------AEE----MYDIEGGQRTTLVSR 99
TENLK+Q RR+ +S G TA+ +EE D+ T L+S
Sbjct 148 TENLKQQKTRRDQFSQGPGPLAAHTVSPSTAKQGSLLLSEENQAVSIDMGSSDTTPLLST 207
Query 100 --------PATAYSQARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSM 151
+ Y Q RAE ++N++ I EL IFQ++A M+ QEE+++RI ++ +
Sbjct 208 QTQMAIYDDSDNYVQQRAETMQNIESTIVELGGIFQQLAHMVKEQEEIVERIDTNVADAE 267
Query 152 YHIREGQNELLNFYNRISSNRSLIIKV 178
+I E+L ++ +S NR L+IK+
Sbjct 268 LNIEAAHGEILKYFQSVSKNRWLMIKI 294
> CE05944
Length=413
Score = 65.1 bits (157), Expect = 8e-11, Method: Compositional matrix adjust.
Identities = 52/206 (25%), Positives = 103/206 (50%), Gaps = 29/206 (14%)
Query 1 IRKTINTLNCKVDTLEKAAA--AAGVSGQSKQHYITVTEVLKTRLLDVTKEFKDILMLRT 58
++ I LN ++ L++ + A + Q+ H V L+++L +V K+++ +L + T
Sbjct 191 VKSDITGLNKQIGQLQEFSKRRAGNMKNQNSGHIQLVVVGLQSKLANVGKDYQSVLEIST 250
Query 59 ENLKRQDKRRNLYSFGG---------DSG----------------TAEAEEMYDIEGGQ- 92
E +K + RR+ +S G SG ++ A +M + Q
Sbjct 251 ETMKAEKNRRDKFSSGAAVPMGLPSSSSGANVRSKLLQDDEQHGSSSIALDMGALSNMQS 310
Query 93 RTTLVSRPAT-AYSQARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSM 151
+ T+ R ++ Y+QAR+ + ++ I EL IF ++A++++ Q EMI RI +++ +
Sbjct 311 QQTMQQRDSSLEYAQARSNTMATIEGSISELGQIFSQLASLVSEQGEMITRIDSNVEDTA 370
Query 152 YHIREGQNELLNFYNRISSNRSLIIK 177
+I +EL+ + IS NR L+I+
Sbjct 371 LNIDMAHSELVRYLQNISKNRWLMIQ 396
> YLR026c
Length=340
Score = 63.2 bits (152), Expect = 4e-10, Method: Compositional matrix adjust.
Identities = 58/216 (26%), Positives = 93/216 (43%), Gaps = 50/216 (23%)
Query 10 CKVDTLEKAAAAAGVSGQSK------QHYITVTEVLKTRLLDVTKEFKDILMLRTENLKR 63
++ L+K S QS QH V +L T++ +++ FKD+L R + L+
Sbjct 111 VQLSQLKKTDVNGNTSNQSSKQPSAVQHSKNVVNLLNTQMKNISGSFKDVLEER-QRLEM 169
Query 64 QDKRRNLYSFGGDSGTAEAEEM-----------YDIEGGQRTTLV--------------- 97
+K R D+G A A++ Y+ T+L+
Sbjct 170 ANKDR-WQKLTTDTGHAPADDQTQSNHAADLTTYNNSNPFMTSLLDESSEKNNNSSNQGE 228
Query 98 -SRPAT---------------AYSQARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQ 141
S P Y Q R AVE ++ I E+ ++FQ++A+M+ Q E+IQ
Sbjct 229 LSFPQNDSQLMLMEEGQLSNNVYLQERNRAVETIESTIQEVGNLFQQLASMVQEQGEVIQ 288
Query 142 RIGQDIDTSMYHIREGQNELLNFYNRISSNRSLIIK 177
RI ++D +I Q ELL +++RI SNR L K
Sbjct 289 RIDANVDDIDLNISGAQRELLKYFDRIKSNRWLAAK 324
> CE15978
Length=275
Score = 43.5 bits (101), Expect = 2e-04, Method: Compositional matrix adjust.
Identities = 27/128 (21%), Positives = 62/128 (48%), Gaps = 9/128 (7%)
Query 76 DSGTAEAEEMYDIEGGQRTTLVSRPATAYSQA-------RAEAVENVQRVIGELASIFQR 128
D+ A YD+ G + TA Q R A++ ++R IG++ +IF
Sbjct 147 DAQAARDAAEYDMYGNNGRSGGQMQMTAQQQGNLQDMKERQNALQQLERDIGDVNAIFAE 206
Query 129 VATMIAHQEEMIQRIGQDIDTSMYHIREGQNELLN--FYNRISSNRSLIIKVFLLLMAFI 186
+A ++ Q +M+ I +++ + ++ +G + +YN+ + + L++ F +++ FI
Sbjct 207 LANIVHEQGDMVDSIEANVEHAQIYVEQGAQNVQQAVYYNQKARQKKLLLLCFFVILIFI 266
Query 187 LFFVFFLS 194
+ +L+
Sbjct 267 IGLTLYLA 274
> SPCC825.03c
Length=284
Score = 39.3 bits (90), Expect = 0.005, Method: Compositional matrix adjust.
Identities = 38/126 (30%), Positives = 57/126 (45%), Gaps = 17/126 (13%)
Query 34 TVTEVLKTRLLDVTKEF----KDILMLRTENLKRQDKRRNLYSFGGDSGTAEAEEMYDIE 89
T TE +K + +D + F K + ++RQ + N + D TA EE
Sbjct 104 TQTEAVKKKFMDQIRHFLQIEKTYRAQYEQRMRRQLEIANPRATEDDFQTAINEE----N 159
Query 90 GGQ-------RTTLVSRPATAYS--QARAEAVENVQRVIGELASIFQRVATMIAHQEEMI 140
GGQ R+ TA Q R ++ ++R I ELA +FQ +ATM+ QE M+
Sbjct 160 GGQVFAQALLRSNRSGEARTALREVQERHADIKRIERTIAELAQLFQDMATMVQEQEPMV 219
Query 141 QRIGQD 146
+I D
Sbjct 220 DKIVTD 225
> Hs20149560
Length=297
Score = 38.9 bits (89), Expect = 0.006, Method: Compositional matrix adjust.
Identities = 42/144 (29%), Positives = 67/144 (46%), Gaps = 21/144 (14%)
Query 59 ENLKRQDKRRNLYSFGGDSGTAEAEEMYDIEGGQ---------RTTLVSRPATAYSQARA 109
E ++RQ K N G E E+M D GQ + T V+R A AR
Sbjct 154 ERIRRQLKITN----AGMVSDEELEQMLD--SGQSEVFVSNILKDTQVTRQALNEISARH 207
Query 110 EAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNEL---LNFYN 166
++ ++R I EL IF +AT + Q EMI RI ++I +S ++ GQ + L
Sbjct 208 SEIQQLERSIRELHDIFTFLATEVEMQGEMINRIEKNILSSADYVERGQEHVKTALENQK 267
Query 167 RISSNRSLI---IKVFLLLMAFIL 187
+ + LI + + ++L+A I+
Sbjct 268 KARKKKVLIAICVSITVVLLAVII 291
> CE04496
Length=299
Score = 36.6 bits (83), Expect = 0.034, Method: Compositional matrix adjust.
Identities = 17/88 (19%), Positives = 46/88 (52%), Gaps = 0/88 (0%)
Query 106 QARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNELLNFY 165
++RA+ ++N++R +GELA +F + M+ Q +M+ I ++ + + ++ + +
Sbjct 201 KSRADELKNLERQMGELAQMFHDLHIMVVSQAKMVDSIVNSVENATEYAKQARGNVEEAR 260
Query 166 NRISSNRSLIIKVFLLLMAFILFFVFFL 193
N R + + + + + +L + F+
Sbjct 261 NLQKRARKMKVCIIIGSIIAVLILILFI 288
> ECU05g0820
Length=240
Score = 34.7 bits (78), Expect = 0.12, Method: Compositional matrix adjust.
Identities = 26/92 (28%), Positives = 45/92 (48%), Gaps = 5/92 (5%)
Query 105 SQARAEAVENVQRV---IGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNEL 161
S+ E V+ QR+ I E+ I + ++ I+ QEE +RI + TS I + +
Sbjct 148 SEVVTERVKERQRISMQISEIGQIMEEISMHISLQEESFKRIDDLMGTSDTLISGSLDLM 207
Query 162 LNFYNRISSNRSLIIKVFLLLMAFILFFVFFL 193
+ +SS R I++ + M +L VF+L
Sbjct 208 RKTWENVSSTRPAIVRFVMFWM--VLALVFWL 237
> At3g03800
Length=306
Score = 33.9 bits (76), Expect = 0.19, Method: Compositional matrix adjust.
Identities = 17/70 (24%), Positives = 38/70 (54%), Gaps = 0/70 (0%)
Query 106 QARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNELLNFY 165
Q R +AV ++++ + +L +F +A ++ Q EM+ I + +++ H++ G N+L
Sbjct 209 QERHDAVRDLEKKLLDLQQVFLDMAVLVDAQGEMLDNIENMVSSAVDHVQSGNNQLTKAV 268
Query 166 NRISSNRSLI 175
S+R +
Sbjct 269 KSQKSSRKWM 278
> SPBC31E1.04
Length=317
Score = 33.5 bits (75), Expect = 0.25, Method: Compositional matrix adjust.
Identities = 30/112 (26%), Positives = 51/112 (45%), Gaps = 15/112 (13%)
Query 90 GGQRTTL----VSRPATAYSQ----ARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQ 141
GQR L +S Y Q R +EN+ + I EL IF+ ++T+I Q E++
Sbjct 149 SGQRQPLTESKISNSQLEYQQRLINERQGEIENLTQGINELNEIFRDLSTIINEQGELVT 208
Query 142 RIGQDIDTSMYHIREGQNEL--LNFYNRISSNRSLIIKVFLLLMAFILFFVF 191
I ++ + + + +L N ++R + RS F L +F +F F
Sbjct 209 NIEYNVGNTSTNTKNASRQLQIANEHSRKARKRS-----FCFLKSFAMFSSF 255
> 7290364
Length=315
Score = 33.1 bits (74), Expect = 0.37, Method: Compositional matrix adjust.
Identities = 14/47 (29%), Positives = 29/47 (61%), Gaps = 0/47 (0%)
Query 115 VQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNEL 161
+++ I E+ ++F R+ T++ Q E+IQR+ + H+ +G +EL
Sbjct 245 LEKSIEEVHALFMRIQTLVMEQSEVIQRVEFHAQQATLHVDKGADEL 291
> SPAC823.05c
Length=301
Score = 32.7 bits (73), Expect = 0.44, Method: Compositional matrix adjust.
Identities = 20/81 (24%), Positives = 38/81 (46%), Gaps = 0/81 (0%)
Query 111 AVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNELLNFYNRISS 170
AV + I ELA +FQ + ++ Q ++ RI +I+ + H + + EL+ + +
Sbjct 215 AVAKIAEGIIELAQMFQDLQVLVIEQGALVDRIDFNIEQTQVHAKSAEKELIKAESHQKN 274
Query 171 NRSLIIKVFLLLMAFILFFVF 191
L FL+L+ L +
Sbjct 275 TGRLRFICFLILLIVALIVIL 295
> At3g52400
Length=341
Score = 32.3 bits (72), Expect = 0.61, Method: Compositional matrix adjust.
Identities = 16/64 (25%), Positives = 35/64 (54%), Gaps = 2/64 (3%)
Query 106 QARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNELLN-- 163
Q R +AV+++++ + EL +F +A ++ HQ + I ++ + +R G + L+
Sbjct 217 QERHDAVKDIEKSLNELHQVFLDMAVLVEHQGAQLDDIEGNVKRANSLVRSGADRLVKAR 276
Query 164 FYNR 167
FY +
Sbjct 277 FYQK 280
> YMR183c
Length=295
Score = 32.0 bits (71), Expect = 0.73, Method: Compositional matrix adjust.
Identities = 39/187 (20%), Positives = 84/187 (44%), Gaps = 22/187 (11%)
Query 17 KAAAAAGVSGQSKQHYITVTEVLKTRLLDVTKEFKDILMLRTENLKRQDKRRNLYSFGGD 76
K A G+ +KQ E + + L + ++++ I E K Q KR+ Y+
Sbjct 103 KDAQRDGLHDSNKQ---AQAENCRQKFLKLIQDYRIIDSNYKEESKEQAKRQ--YTIIQP 157
Query 77 SGTAEAEE--MYDIEGGQRTTLV---------SRPATAYSQARAEAVENVQRVIGELASI 125
T E E + D+ G Q + ++ A A QAR + + +++ + EL +
Sbjct 158 EATDEEVEAAINDVNGQQIFSQALLNANRRGEAKTALAEVQARHQELLKLEKTMAELTQL 217
Query 126 FQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNELLNFYNRI--SSNRSLIIKVFLLLM 183
F + ++ Q+E + I ++++ + + +G + N+ S+ ++ K+ L++
Sbjct 218 FNDMEELVIEQQENVDVIDKNVEDAQQDVEQG----VGHTNKAVKSARKARKNKIRCLII 273
Query 184 AFILFFV 190
FI+F +
Sbjct 274 CFIIFAI 280
> 7299676
Length=763
Score = 31.2 bits (69), Expect = 1.3, Method: Compositional matrix adjust.
Identities = 23/78 (29%), Positives = 41/78 (52%), Gaps = 4/78 (5%)
Query 104 YSQARAEAVENVQR--VIGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQ-NE 160
YS+ A E Q+ + G L + + + + H ++ R+ QD+D MY I + NE
Sbjct 227 YSKGLAFTHEERQQLGIHGMLPYVVREPSEQVEHCRALLARLDQDLDKYMYLISLSERNE 286
Query 161 LLNFYNRISSNRSLIIKV 178
L FYN +SS+ + ++ +
Sbjct 287 RL-FYNVLSSDIAYMMPL 303
> CE18766
Length=838
Score = 30.8 bits (68), Expect = 2.0, Method: Compositional matrix adjust.
Identities = 38/147 (25%), Positives = 57/147 (38%), Gaps = 23/147 (15%)
Query 1 IRKTINTLNCKVDTLEKAAAAAGVSGQSK-QHYITVTEVLKTRLLD-------------V 46
+ K +TL+ K L+ A GQS QH I+V E L + LD +
Sbjct 112 LEKLKSTLSTKNPALKVNELAYSFIGQSNVQHAISVIETLNSEALDTIHFRTDVDEEINL 171
Query 47 TK-----EFKDILMLRTENLKRQDKRRNLYSFGGDSGTAEAEEMYDIEGGQRTTLVSRP- 100
TK + +I L+ E + N T + + D++ ++ L S+
Sbjct 172 TKIMELPHWPNITCLKIEGFIVSETLENFLHLTNAKVTKHSITIQDLQILKKNFLGSQSE 231
Query 101 ---ATAYSQARAEAVENVQRVIGELAS 124
Y E ENVQ V GEL+S
Sbjct 232 KKFVLVYKLDTGEIAENVQVVFGELSS 258
> 7292492
Length=301
Score = 30.4 bits (67), Expect = 2.1, Method: Compositional matrix adjust.
Identities = 19/80 (23%), Positives = 44/80 (55%), Gaps = 3/80 (3%)
Query 103 AYSQARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNELL 162
A QA + +EN+Q+ I +L +FQ + + A Q +++I + + ++ ++++G+ L
Sbjct 157 AQRQACLDQMENLQQEIYDLHGMFQGMRQLTAEQSVAVEKIADNAEEALENVQQGE---L 213
Query 163 NFYNRISSNRSLIIKVFLLL 182
N ++ +++ V LL
Sbjct 214 NLRRALTYKKAMYPVVGALL 233
> At5g08080
Length=307
Score = 30.0 bits (66), Expect = 2.7, Method: Compositional matrix adjust.
Identities = 15/56 (26%), Positives = 32/56 (57%), Gaps = 0/56 (0%)
Query 106 QARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNEL 161
Q R +AV ++++ + +L IF +A ++ Q EM+ I + +++ H++ G L
Sbjct 211 QERHDAVRDLEKKLLDLQQIFLDMAVLVDAQGEMLDNIESQVSSAVDHVQSGNTAL 266
> At1g32270_2
Length=279
Score = 29.6 bits (65), Expect = 3.8, Method: Compositional matrix adjust.
Identities = 17/61 (27%), Positives = 32/61 (52%), Gaps = 1/61 (1%)
Query 106 QARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNELLNFY 165
+AR + ++ V+ I E+ +F+ +A M+ HQ I I + ID +G++ L+
Sbjct 132 EAREQGIQEVKHQISEVMEMFKDLAVMVDHQ-GTIDDIDEKIDNLRSAAAQGKSHLVKAS 190
Query 166 N 166
N
Sbjct 191 N 191
> CE18300
Length=250
Score = 29.6 bits (65), Expect = 4.0, Method: Compositional matrix adjust.
Identities = 12/56 (21%), Positives = 30/56 (53%), Gaps = 0/56 (0%)
Query 106 QARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNEL 161
+ RAEA +++ + +L IFQ + ++ Q +++ I + ++ + ++ G L
Sbjct 133 KERAEATVKIEKDMADLEKIFQELGRIVHEQHDVVDSIEEQVERATEDVKRGNENL 188
> Hs8923604
Length=302
Score = 29.3 bits (64), Expect = 5.0, Method: Compositional matrix adjust.
Identities = 35/150 (23%), Positives = 64/150 (42%), Gaps = 19/150 (12%)
Query 12 VDTLEKAAAAAGVSGQSKQHYITVTEVLKTRLLDVTKEFKDILMLRTENLKRQDKRRNLY 71
+D +++ A+AA + + Q ++ E LK K+F D L L R
Sbjct 91 IDPVKEEASAA--TAEFLQLHLESVEELK-------KQFNDEETLLQPPLTRS------M 135
Query 72 SFGGDSGTAEAEEMYDIEGGQRTTLVSRPATAYSQARAEAVENVQRVIGELASIFQRVAT 131
+ GG T EAE T + + P Q AE+ E ++ + EL+ + +
Sbjct 136 TVGGAFHTTEAE----ASSQSLTQIYALPEIPQDQNAAESRETLEADLIELSQLVTDFSL 191
Query 132 MIAHQEEMIQRIGQDIDTSMYHIREGQNEL 161
++ Q+E I I ++++ ++ EG L
Sbjct 192 LVNSQQEKIDSIADHVNSAAVNVEEGTKNL 221
> Hs4759184
Length=289
Score = 29.3 bits (64), Expect = 5.3, Method: Compositional matrix adjust.
Identities = 23/102 (22%), Positives = 49/102 (48%), Gaps = 10/102 (9%)
Query 67 RRNLYSFGGDSGTAEAEEMYDIEGGQRT--------TLVSRPATAYSQARAEAVENVQRV 118
+R L G + E EEM +E G + +S+ A + + R + + ++
Sbjct 150 QRQLEITGKKTTDEELEEM--LESGNPAIFTSGIIDSQISKQALSEIEGRHKDIVRLESS 207
Query 119 IGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNE 160
I EL +F +A ++ +Q EM+ I ++ ++ H+ + ++E
Sbjct 208 IKELHDMFMDIAMLVENQGEMLDNIELNVMHTVDHVEKARDE 249
> 7295535
Length=1500
Score = 29.3 bits (64), Expect = 5.4, Method: Compositional matrix adjust.
Identities = 19/64 (29%), Positives = 34/64 (53%), Gaps = 5/64 (7%)
Query 33 ITVTEVLKTRLLDVTKEFKDILMLRTENLKRQDKRRNLY----SFGGDSGTAEAEEMYDI 88
+T EV T V D+L+L+ E++ ++ KR+ L+ FGG SG A + +++
Sbjct 176 VTTEEVRVTTEQTVVSGDPDLLLLKNEDVLKRSKRQGLFDLLGGFGG-SGDANKKNKFEV 234
Query 89 EGGQ 92
+ Q
Sbjct 235 DNMQ 238
> 7301098
Length=291
Score = 29.3 bits (64), Expect = 5.6, Method: Compositional matrix adjust.
Identities = 19/89 (21%), Positives = 40/89 (44%), Gaps = 0/89 (0%)
Query 106 QARAEAVENVQRVIGELASIFQRVATMIAHQEEMIQRIGQDIDTSMYHIREGQNELLNFY 165
+AR + + ++ I EL +F +A ++ Q EMI RI ++ +M +++ +
Sbjct 199 EARHQDIMKLETSIKELHDMFMDMAMLVESQGEMIDRIEYHVEHAMDYVQTATQDTKKAL 258
Query 166 NRISSNRSLIIKVFLLLMAFILFFVFFLS 194
S R I + + L + ++S
Sbjct 259 KYQSKARRKKIMILICLTVLGILAASYVS 287
Lambda K H
0.321 0.134 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3264145066
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40