bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_3211_orf1
Length=209
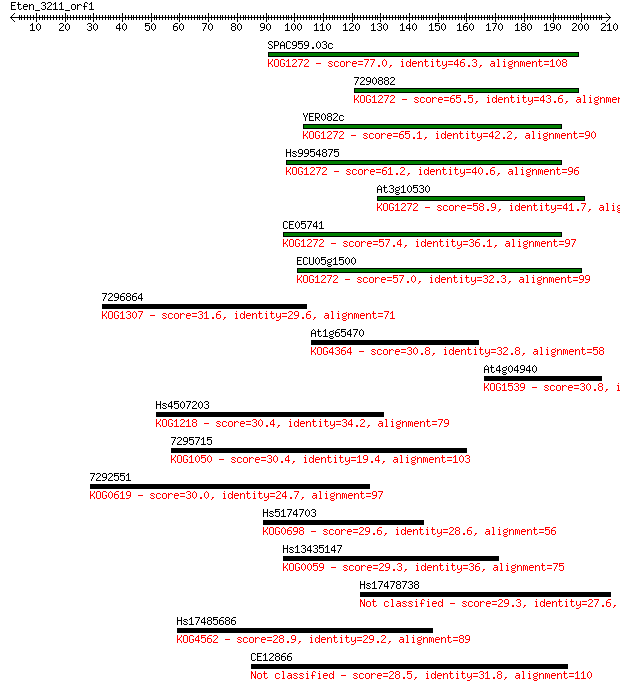
Score E
Sequences producing significant alignments: (Bits) Value
SPAC959.03c 77.0 3e-14
7290882 65.5 7e-11
YER082c 65.1 9e-11
Hs9954875 61.2 1e-09
At3g10530 58.9 8e-09
CE05741 57.4 2e-08
ECU05g1500 57.0 3e-08
7296864 31.6 1.2
At1g65470 30.8 2.0
At4g04940 30.8 2.1
Hs4507203 30.4 2.5
7295715 30.4 2.8
7292551 30.0 3.4
Hs5174703 29.6 4.9
Hs13435147 29.3 5.4
Hs17478738 29.3 6.5
Hs17485686 28.9 6.9
CE12866 28.5 9.1
> SPAC959.03c
Length=520
Score = 77.0 bits (188), Expect = 3e-14, Method: Compositional matrix adjust.
Identities = 50/124 (40%), Positives = 68/124 (54%), Gaps = 16/124 (12%)
Query 91 QLPMDDPQSRTAKRSRG--------TIKRQRALEKKVQESVSAVTALLTETE-------G 135
+L ++D + T K SRG K+ R+ +K++E + V + L +TE G
Sbjct 5 KLNLEDVNASTKKYSRGRKLNPKKIKDKKLRSNIQKIEERIENVESSLAKTEILHEDNPG 64
Query 136 YLEAEGLEKTYKFTQADILENVSEEIGKKAFRLGLS-FGPYFVDYTRGGRHLLLGGQKRE 194
LEAEGLE+TYKF Q + NV+ E K+F L L FG Y DYTR GR +LLGG+K
Sbjct 65 LLEAEGLERTYKFRQDQLAPNVALETATKSFSLDLDKFGGYSFDYTRDGRMILLGGRKGH 124
Query 195 FGCF 198
F
Sbjct 125 ISAF 128
> 7290882
Length=609
Score = 65.5 bits (158), Expect = 7e-11, Method: Compositional matrix adjust.
Identities = 34/78 (43%), Positives = 45/78 (57%), Gaps = 0/78 (0%)
Query 121 ESVSAVTALLTETEGYLEAEGLEKTYKFTQADILENVSEEIGKKAFRLGLSFGPYFVDYT 180
E + LL ET G L+A+ E T +F Q+ I ENV + K F L + FGPY + YT
Sbjct 123 EQAARTELLLQETAGQLQADEGETTAEFRQSQIAENVDIQSSAKHFNLNMEFGPYRMRYT 182
Query 181 RGGRHLLLGGQKREFGCF 198
+ GRHLLLGG++ F
Sbjct 183 KNGRHLLLGGKRGHVAAF 200
> YER082c
Length=554
Score = 65.1 bits (157), Expect = 9e-11, Method: Compositional matrix adjust.
Identities = 38/92 (41%), Positives = 51/92 (55%), Gaps = 2/92 (2%)
Query 103 KRSRGTIKRQRALEKKVQESVSAVTALLTETEGYLEAEG-LEKTYKFTQADILENVSEEI 161
K+ R +K+ KK S +A LL E+ GYLE E LEKT+K Q++I +V
Sbjct 40 KKLRAGLKKIDEQYKKAVSSAAATDYLLPESNGYLEPENELEKTFKVQQSEIKSSVDVST 99
Query 162 GKKAFRLGLS-FGPYFVDYTRGGRHLLLGGQK 192
KA L L FGPY + Y + G HLL+ G+K
Sbjct 100 ANKALDLSLKEFGPYHIKYAKNGTHLLITGRK 131
> Hs9954875
Length=610
Score = 61.2 bits (147), Expect = 1e-09, Method: Compositional matrix adjust.
Identities = 39/97 (40%), Positives = 52/97 (53%), Gaps = 3/97 (3%)
Query 97 PQSRTAKRSRGTIKRQRALEKKVQESVSAVTALLTETEGYLEAEGLEKTYKFTQADILEN 156
P S+ RSR + E ++ + S + LL E G+LE E E T K QADI+E
Sbjct 122 PHSKAKTRSRLEVAEAEEEETSIKAARSEL--LLAEEPGFLEGEDGEDTAKICQADIVEA 179
Query 157 VSEEIGKKAFRLGL-SFGPYFVDYTRGGRHLLLGGQK 192
V K F L L FGPY ++Y+R GRHL GG++
Sbjct 180 VDIASAAKHFDLNLRQFGPYRLNYSRTGRHLAFGGRR 216
> At3g10530
Length=548
Score = 58.9 bits (141), Expect = 8e-09, Method: Compositional matrix adjust.
Identities = 30/73 (41%), Positives = 40/73 (54%), Gaps = 1/73 (1%)
Query 129 LLTETEGYLEAEGLEKTYKFTQADILENVSEEIGKKAFRLGL-SFGPYFVDYTRGGRHLL 187
LL GYLE EGLEKT++ Q DI V + + + L FGPY +D+T GRH+L
Sbjct 86 LLPAEAGYLETEGLEKTWRVKQTDIANEVDILSSRNQYDIVLPDFGPYKLDFTASGRHML 145
Query 188 LGGQKREFGCFGL 200
GG+K +
Sbjct 146 AGGRKGHLALLDM 158
> CE05741
Length=580
Score = 57.4 bits (137), Expect = 2e-08, Method: Compositional matrix adjust.
Identities = 35/100 (35%), Positives = 51/100 (51%), Gaps = 3/100 (3%)
Query 96 DPQSRTAKRSRGTIKRQRALEKKVQESVSAVTALLTETEGYL--EAEGLEKTYKFTQADI 153
DP S +K + +K + ++ E+ + L E G++ + E TY Q DI
Sbjct 90 DPSSMRSKLQQEKMKVTKDKFQRRIEATAKAEKRLVENTGFVAPDEEDHSLTYSIRQKDI 149
Query 154 LENVSEEIGKKAFRLGLS-FGPYFVDYTRGGRHLLLGGQK 192
E+V K F L L FGPY +DYT GRHL++GG+K
Sbjct 150 AESVDLAAATKHFELKLPRFGPYHIDYTDNGRHLVIGGRK 189
> ECU05g1500
Length=435
Score = 57.0 bits (136), Expect = 3e-08, Method: Compositional matrix adjust.
Identities = 32/99 (32%), Positives = 51/99 (51%), Gaps = 0/99 (0%)
Query 101 TAKRSRGTIKRQRALEKKVQESVSAVTALLTETEGYLEAEGLEKTYKFTQADILENVSEE 160
+ K R IK + + ++ V+ L TET GY+EA+ EKTY+ TQ I +VS +
Sbjct 2 SKKIVRERIKNSKKIAEEAHRDVTNYGILRTETCGYIEADSDEKTYEVTQDKIRRHVSIK 61
Query 161 IGKKAFRLGLSFGPYFVDYTRGGRHLLLGGQKREFGCFG 199
++A+ L GP + ++R G ++LL K F
Sbjct 62 ACEQAYNLSFENGPIYAKHSRNGAYMLLRNSKGYLASFN 100
> 7296864
Length=658
Score = 31.6 bits (70), Expect = 1.2, Method: Composition-based stats.
Identities = 21/75 (28%), Positives = 39/75 (52%), Gaps = 7/75 (9%)
Query 33 FCDLGVHCLA---QNEAATW-LALAVGLQQPNQLQRGALHQARDMGGPQGAQASTAPGAK 88
C +G +C+A E W L L + + P++ ++ AL +++ P+G+ + G
Sbjct 261 LCYVG-YCVALHFNTELERWALGLNLPFKLPSKEEQSALVTYKNV--PEGSYTQESVGQT 317
Query 89 QSQLPMDDPQSRTAK 103
Q Q DD ++R+AK
Sbjct 318 QGQKATDDSETRSAK 332
> At1g65470
Length=815
Score = 30.8 bits (68), Expect = 2.0, Method: Composition-based stats.
Identities = 19/58 (32%), Positives = 33/58 (56%), Gaps = 8/58 (13%)
Query 106 RGTIKRQRALEKKVQESVSAVTALLTETEGYLEAEGLEKTYKFTQADILENVSEEIGK 163
RG +K +R KK+ E ++AV+A+L + E+T K ++D L +E++GK
Sbjct 165 RGVLKLRRTCRKKIHERITAVSAMLAALQ-------REETEKLWRSD-LSKAAEKLGK 214
> At4g04940
Length=931
Score = 30.8 bits (68), Expect = 2.1, Method: Composition-based stats.
Identities = 14/41 (34%), Positives = 21/41 (51%), Gaps = 0/41 (0%)
Query 166 FRLGLSFGPYFVDYTRGGRHLLLGGQKREFGCFGLPRNENN 206
FR G S P + + GRH+L GQ R F F + + + +
Sbjct 316 FRSGHSAPPLCIRFYSNGRHILSAGQDRAFRLFSVIQEQQS 356
> Hs4507203
Length=830
Score = 30.4 bits (67), Expect = 2.5, Method: Compositional matrix adjust.
Identities = 27/90 (30%), Positives = 40/90 (44%), Gaps = 17/90 (18%)
Query 52 LAVGLQQPNQLQRGALHQARDMGGPQGAQASTAPGAKQSQLPMDDP------QSRTAKR- 104
L+ L++P +L RGA GP+G +A + G +++ P P S T R
Sbjct 609 LSSPLRKPKRLSRGA------QSGPEGREAEESTGPDEAEAPESFPAAASPGDSATGHRR 662
Query 105 ----SRGTIKRQRALEKKVQESVSAVTALL 130
SR + A+E VQES VT +
Sbjct 663 PPLGSRTVAEHVEAIEGSVQESSGPVTTIY 692
> 7295715
Length=809
Score = 30.4 bits (67), Expect = 2.8, Method: Composition-based stats.
Identities = 20/103 (19%), Positives = 51/103 (49%), Gaps = 3/103 (2%)
Query 57 QQPNQLQRGALHQARDMGGPQGAQASTAPGAKQSQLPMDDPQSRTAKRSRGTIKRQRALE 116
+ PN L+ + +AR + G +A+ A A +++ P+ Q + S ++ ++
Sbjct 653 ETPNHLRGAMVDKARSLIEKYGFKATEAHCALEARPPV---QWNKGRASIYILRTSFGVD 709
Query 117 KKVQESVSAVTALLTETEGYLEAEGLEKTYKFTQADILENVSE 159
+ + V LT+ + + +G+ +T++ T +DI++ ++
Sbjct 710 WNERIKIIYVGDDLTDEDAMVALKGMARTFRVTSSDIVKTAAD 752
> 7292551
Length=691
Score = 30.0 bits (66), Expect = 3.4, Method: Composition-based stats.
Identities = 24/97 (24%), Positives = 34/97 (35%), Gaps = 2/97 (2%)
Query 29 LRGSFCDLGVHCLAQNEAATWLALAVGLQQPNQLQRGALHQARDMGGPQGAQASTAPGAK 88
LR S DL HC Q + V + L G G ++ G+
Sbjct 437 LRDSLEDL--HCTLQEPKLSHFVDPVPPTILDVLNIGGFTAIGSNSASMGGVGNSVVGSS 494
Query 89 QSQLPMDDPQSRTAKRSRGTIKRQRALEKKVQESVSA 125
S L MDD + RSR ++ QR + V +
Sbjct 495 YSGLTMDDSRKHLGSRSRQALRGQRQFASSAENVVES 531
> Hs5174703
Length=504
Score = 29.6 bits (65), Expect = 4.9, Method: Compositional matrix adjust.
Identities = 16/56 (28%), Positives = 29/56 (51%), Gaps = 0/56 (0%)
Query 89 QSQLPMDDPQSRTAKRSRGTIKRQRALEKKVQESVSAVTALLTETEGYLEAEGLEK 144
QSQLP PQ + + + ++R + LE+++ AV A+L + Y+ G +
Sbjct 132 QSQLPEGVPQHQLPPQYQKILERLKTLEREISGGAMAVVAVLLNNKLYVANVGTNR 187
> Hs13435147
Length=156
Score = 29.3 bits (64), Expect = 5.4, Method: Compositional matrix adjust.
Identities = 27/79 (34%), Positives = 34/79 (43%), Gaps = 4/79 (5%)
Query 96 DPQSRTAKRSRGTIKRQRALEKKVQ-ESVSAVTALLT---ETEGYLEAEGLEKTYKFTQA 151
DP R + RSRGT E+ VQ E V A AL T E E + A L K Y T+
Sbjct 11 DPVFRISPRSRGTHTNPEEPEEDVQAERVQAANALTTPNLEEEPVITASCLHKEYYETKK 70
Query 152 DILENVSEEIGKKAFRLGL 170
+ + + FR L
Sbjct 71 VAFQQQGRKQPSEMFRFVL 89
> Hs17478738
Length=896
Score = 29.3 bits (64), Expect = 6.5, Method: Composition-based stats.
Identities = 24/87 (27%), Positives = 41/87 (47%), Gaps = 4/87 (4%)
Query 123 VSAVTALLTETEGYLEAEGLEKTYKFTQADILENVSEEIGKKAFRLGLSFGPYFVDYTRG 182
+ A+ ALL G ++AE +KTY + + +++ K L S ++DY +
Sbjct 46 IRAICALLNSGGGVIKAEIDDKTYSYQCHGLGQDLETSFQK----LLPSGSQKYLDYMQQ 101
Query 183 GRHLLLGGQKREFGCFGLPRNENNLRS 209
G +LL+ + F LP +LRS
Sbjct 102 GHNLLIFVKSWSPDVFSLPLRICSLRS 128
> Hs17485686
Length=341
Score = 28.9 bits (63), Expect = 6.9, Method: Compositional matrix adjust.
Identities = 26/94 (27%), Positives = 40/94 (42%), Gaps = 5/94 (5%)
Query 59 PNQLQRGALHQAR---DMGGPQGAQASTAPGAKQSQLPMD--DPQSRTAKRSRGTIKRQR 113
P+ L +G L + G PQ +Q +P P + D SR+ ++ +
Sbjct 44 PSTLIQGTLEKVSASGTPGTPQSSQRVCSPCTTIKATPWNQSDESSRSQEKKDPGASQAL 103
Query 114 ALEKKVQESVSAVTALLTETEGYLEAEGLEKTYK 147
LEKKV E V ++ T + EAE L+ K
Sbjct 104 MLEKKVDELVKFLSVKYTTKQPITEAEMLKGVIK 137
> CE12866
Length=1165
Score = 28.5 bits (62), Expect = 9.1, Method: Composition-based stats.
Identities = 35/123 (28%), Positives = 55/123 (44%), Gaps = 18/123 (14%)
Query 85 PGAKQ----SQLPMDDPQSRTAKRSRGTI--KRQRALEKKVQESVSAVTALLTE-----T 133
P A++ +LPM++ + K+ + K +R LEK ES+ V L+ E
Sbjct 695 PAARKCFAIRELPMEEVKKPVKKKDVSKMDNKPERELEKTQDESIKDVPKLILELDMSSA 754
Query 134 EGYLEA--EGLEKTYKFTQADILENVSEEIGKKAFRLGLSFGPYFVDYTRGGRHLLLGGQ 191
E Y++ E L+ +K T L SEE+ KK + G+ F Y GR L Q
Sbjct 755 EKYVDCRYEALQDMHKETCIPFL---SEELIKKFY--GVKFDAMNEKYEVPGREALKSEQ 809
Query 192 KRE 194
++
Sbjct 810 MKQ 812
Lambda K H
0.316 0.133 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3754464630
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40