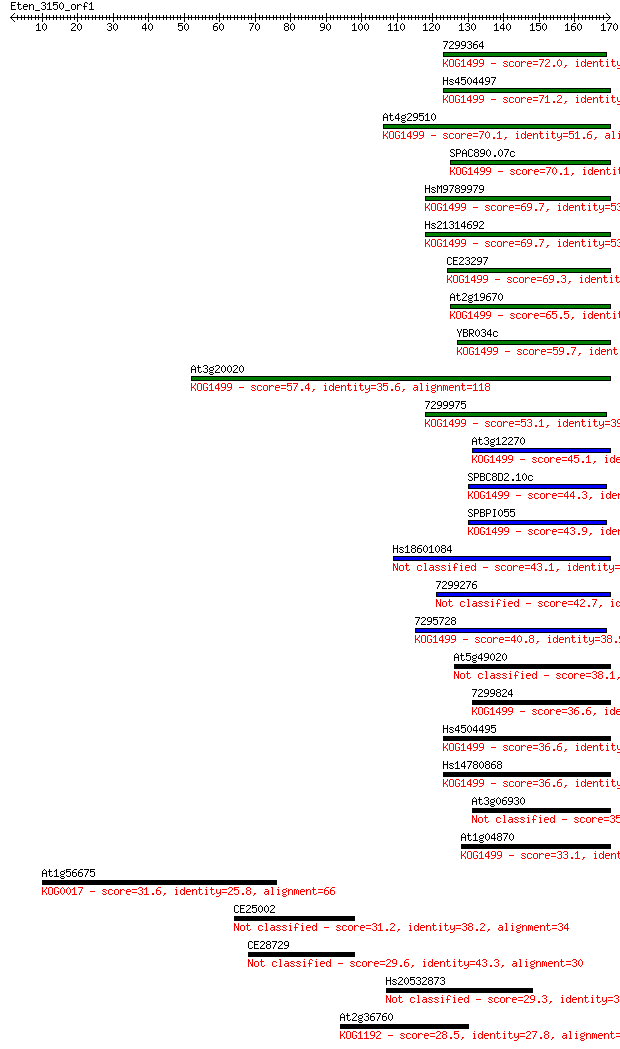
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_3150_orf1
Length=169
Score E
Sequences producing significant alignments: (Bits) Value
7299364 72.0 5e-13
Hs4504497 71.2 9e-13
At4g29510 70.1 2e-12
SPAC890.07c 70.1 2e-12
HsM9789979 69.7 3e-12
Hs21314692 69.7 3e-12
CE23297 69.3 3e-12
At2g19670 65.5 5e-11
YBR034c 59.7 2e-09
At3g20020 57.4 1e-08
7299975 53.1 2e-07
At3g12270 45.1 7e-05
SPBC8D2.10c 44.3 1e-04
SPBPI055 43.9 1e-04
Hs18601084 43.1 3e-04
7299276 42.7 4e-04
7295728 40.8 0.001
At5g49020 38.1 0.009
7299824 36.6 0.021
Hs4504495 36.6 0.022
Hs14780868 36.6 0.024
At3g06930 35.0 0.065
At1g04870 33.1 0.25
At1g56675 31.6 0.71
CE25002 31.2 1.1
CE28729 29.6 3.3
Hs20532873 29.3 4.4
At2g36760 28.5 6.5
> 7299364
Length=376
Score = 72.0 bits (175), Expect = 5e-13, Method: Compositional matrix adjust.
Identities = 29/46 (63%), Positives = 39/46 (84%), Gaps = 0/46 (0%)
Query 123 DDISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGK 168
D+++S DYYF+SYAHF IHEEM+KD VRT +Y+ A+ N+HLF+GK
Sbjct 48 DEMTSRDYYFDSYAHFGIHEEMLKDEVRTVTYRNAMYHNKHLFQGK 93
> Hs4504497
Length=360
Score = 71.2 bits (173), Expect = 9e-13, Method: Compositional matrix adjust.
Identities = 28/47 (59%), Positives = 39/47 (82%), Gaps = 0/47 (0%)
Query 123 DDISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
+D++S DYYF+SYAHF IHEEM+KD VRT +Y+ ++ N+HLF+ KV
Sbjct 35 EDMTSKDYYFDSYAHFGIHEEMLKDEVRTLTYRNSMFHNRHLFKDKV 81
> At4g29510
Length=390
Score = 70.1 bits (170), Expect = 2e-12, Method: Compositional matrix adjust.
Identities = 33/68 (48%), Positives = 48/68 (70%), Gaps = 5/68 (7%)
Query 106 SPEEKAAF----SADWKQLQQDDISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDN 161
+P++++ F SAD ++ DD +SADYYF+SY+HF IHEEM+KD VRT +YQ I N
Sbjct 44 TPQDESMFDAGESADTAEVT-DDTTSADYYFDSYSHFGIHEEMLKDVVRTKTYQNVIYQN 102
Query 162 QHLFRGKV 169
+ L + K+
Sbjct 103 KFLIKDKI 110
> SPAC890.07c
Length=339
Score = 70.1 bits (170), Expect = 2e-12, Method: Compositional matrix adjust.
Identities = 28/45 (62%), Positives = 37/45 (82%), Gaps = 0/45 (0%)
Query 125 ISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
+++ DYYF+SY+H+ IHEEM+KD VRT SY+ AIM N HLFR K+
Sbjct 13 LTAKDYYFDSYSHWGIHEEMLKDDVRTLSYRDAIMQNPHLFRDKI 57
> HsM9789979
Length=334
Score = 69.7 bits (169), Expect = 3e-12, Method: Compositional matrix adjust.
Identities = 28/52 (53%), Positives = 41/52 (78%), Gaps = 0/52 (0%)
Query 118 KQLQQDDISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
K L ++++S DYYF+SYAHF IHEEM+KD VRT +Y+ ++ N+H+F+ KV
Sbjct 3 KLLNPEEMTSRDYYFDSYAHFGIHEEMLKDEVRTLTYRNSMYHNKHVFKDKV 54
> Hs21314692
Length=334
Score = 69.7 bits (169), Expect = 3e-12, Method: Compositional matrix adjust.
Identities = 28/52 (53%), Positives = 41/52 (78%), Gaps = 0/52 (0%)
Query 118 KQLQQDDISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
K L ++++S DYYF+SYAHF IHEEM+KD VRT +Y+ ++ N+H+F+ KV
Sbjct 3 KLLNPEEMTSRDYYFDSYAHFGIHEEMLKDEVRTLTYRNSMYHNKHVFKDKV 54
> CE23297
Length=348
Score = 69.3 bits (168), Expect = 3e-12, Method: Compositional matrix adjust.
Identities = 28/46 (60%), Positives = 37/46 (80%), Gaps = 0/46 (0%)
Query 124 DISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
+++S DYYF+SYAHF IHEEM+KD VRT +Y+ +I N HLF+ KV
Sbjct 20 ELTSKDYYFDSYAHFGIHEEMLKDEVRTTTYRNSIYHNSHLFKDKV 65
> At2g19670
Length=366
Score = 65.5 bits (158), Expect = 5e-11, Method: Compositional matrix adjust.
Identities = 28/45 (62%), Positives = 35/45 (77%), Gaps = 0/45 (0%)
Query 125 ISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
I+SADYYF+SY+HF IHEEM+KD VRT SYQ I N+ L + K+
Sbjct 42 ITSADYYFDSYSHFGIHEEMLKDVVRTKSYQDVIYKNKFLIKDKI 86
> YBR034c
Length=348
Score = 59.7 bits (143), Expect = 2e-09, Method: Compositional matrix adjust.
Identities = 24/43 (55%), Positives = 34/43 (79%), Gaps = 0/43 (0%)
Query 127 SADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
S +YFNSY H+ IHEEM++D+VRT SY+ AI+ N+ LF+ K+
Sbjct 19 SEQHYFNSYDHYGIHEEMLQDTVRTLSYRNAIIQNKDLFKDKI 61
> At3g20020
Length=409
Score = 57.4 bits (137), Expect = 1e-08, Method: Compositional matrix adjust.
Identities = 42/123 (34%), Positives = 57/123 (46%), Gaps = 7/123 (5%)
Query 52 MESGQNHQKGALPSHAKGNGISANNGGSQTGNGSVR-----GGRGGRPAYAQSLQCQKLS 106
M+SG + G H + + G S + G + G R R A L+
Sbjct 1 MQSGGDFSNGFHGDHHRELELEDKQGPSLSSFGRAKKRSHAGARDPRGGLANVLRVSDQL 60
Query 107 PEEKAAFSADWKQLQQDDISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFR 166
E K+ +++ D A YF+SYAH IHEEMIKD RT +Y+ AIM +Q L
Sbjct 61 GEHKSLETSESSPPPCTDFDVA--YFHSYAHVGIHEEMIKDRARTETYREAIMQHQSLIE 118
Query 167 GKV 169
GKV
Sbjct 119 GKV 121
> 7299975
Length=516
Score = 53.1 bits (126), Expect = 2e-07, Method: Composition-based stats.
Identities = 20/51 (39%), Positives = 35/51 (68%), Gaps = 0/51 (0%)
Query 118 KQLQQDDISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGK 168
+ + ++++ + YF SYAHF IH EM+ D VRT +Y+ +++ N+ + RGK
Sbjct 195 QSVPRNNVCLDNEYFKSYAHFGIHHEMLSDKVRTSTYRASLLQNEAVVRGK 245
> At3g12270
Length=590
Score = 45.1 bits (105), Expect = 7e-05, Method: Composition-based stats.
Identities = 19/39 (48%), Positives = 25/39 (64%), Gaps = 0/39 (0%)
Query 131 YFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
YF SY+ F IH EM+ D VRT +Y+ A++ N L G V
Sbjct 241 YFGSYSSFGIHREMLSDKVRTEAYRDALLKNPTLLNGSV 279
> SPBC8D2.10c
Length=543
Score = 44.3 bits (103), Expect = 1e-04, Method: Composition-based stats.
Identities = 20/39 (51%), Positives = 25/39 (64%), Gaps = 0/39 (0%)
Query 130 YYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGK 168
YYF SYA IH M+ DSVRT Y+ + N+H+F GK
Sbjct 219 YYFESYAGNDIHFLMLNDSVRTEGYRDFVYHNKHIFAGK 257
> SPBPI055
Length=348
Score = 43.9 bits (102), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 20/39 (51%), Positives = 25/39 (64%), Gaps = 0/39 (0%)
Query 130 YYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGK 168
YYF SYA IH M+ DSVRT Y+ + N+H+F GK
Sbjct 24 YYFESYAGNDIHFLMLNDSVRTEGYRDFVYHNKHIFAGK 62
> Hs18601084
Length=608
Score = 43.1 bits (100), Expect = 3e-04, Method: Compositional matrix adjust.
Identities = 23/61 (37%), Positives = 36/61 (59%), Gaps = 6/61 (9%)
Query 109 EKAAFSADWKQLQQDDISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGK 168
E++ FS ++ + SSA YF Y + S + M++D VRT +YQRAI+ N F+ K
Sbjct 133 ERSVFS------ERTEESSAVQYFQFYGYLSQQQNMMQDYVRTGTYQRAILQNHTDFKDK 186
Query 169 V 169
+
Sbjct 187 I 187
> 7299276
Length=530
Score = 42.7 bits (99), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 21/49 (42%), Positives = 31/49 (63%), Gaps = 0/49 (0%)
Query 121 QQDDISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
Q+ + SSA YF Y + S + M++D VRT +YQRAI+ N F+ K+
Sbjct 134 QRTEESSASQYFQFYGYLSQQQNMMQDYVRTSTYQRAILGNAVDFQDKI 182
> 7295728
Length=355
Score = 40.8 bits (94), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 21/54 (38%), Positives = 30/54 (55%), Gaps = 1/54 (1%)
Query 115 ADWKQLQQDDISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGK 168
D Q+ +D ++YF Y IHE ++KDSVR +Y+ AI N+ FR K
Sbjct 17 TDANQIIKDRRRQEEHYFKLYGRIEIHEWLLKDSVRIKAYREAIQHNE-FFRHK 69
> At5g49020
Length=577
Score = 38.1 bits (87), Expect = 0.009, Method: Compositional matrix adjust.
Identities = 17/44 (38%), Positives = 27/44 (61%), Gaps = 0/44 (0%)
Query 126 SSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
+SA YF+ Y + M++D VRT +Y A+M+N+ F G+V
Sbjct 146 ASAKMYFHYYGQLLHQQNMLQDYVRTGTYHAAVMENRSDFSGRV 189
> 7299824
Length=341
Score = 36.6 bits (83), Expect = 0.021, Method: Compositional matrix adjust.
Identities = 15/39 (38%), Positives = 25/39 (64%), Gaps = 0/39 (0%)
Query 131 YFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
YF SY+ H M++DSVR +++ AI+ + LF+ K+
Sbjct 19 YFQSYSRLETHMNMLRDSVRMQAFRDAIVQDGGLFQDKI 57
> Hs4504495
Length=433
Score = 36.6 bits (83), Expect = 0.022, Method: Compositional matrix adjust.
Identities = 16/47 (34%), Positives = 22/47 (46%), Gaps = 0/47 (0%)
Query 123 DDISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
+D + YF SY +H EM+ D RT Y I+ N+ KV
Sbjct 94 EDTWQDEEYFGSYGTLKLHLEMLADQPRTTKYHSVILQNKESLTDKV 140
> Hs14780868
Length=375
Score = 36.6 bits (83), Expect = 0.024, Method: Compositional matrix adjust.
Identities = 16/47 (34%), Positives = 22/47 (46%), Gaps = 0/47 (0%)
Query 123 DDISSADYYFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
+D + YF SY +H EM+ D RT Y I+ N+ KV
Sbjct 94 EDTWQDEEYFGSYGTLKLHLEMLADQPRTTKYHSVILQNKESLTDKV 140
> At3g06930
Length=388
Score = 35.0 bits (79), Expect = 0.065, Method: Compositional matrix adjust.
Identities = 15/39 (38%), Positives = 23/39 (58%), Gaps = 0/39 (0%)
Query 131 YFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
YF+ Y + M++D VRT +Y A+M+N F G+V
Sbjct 2 YFHYYGQLLHQQNMLQDYVRTGTYYAAVMENHSDFAGRV 40
> At1g04870
Length=383
Score = 33.1 bits (74), Expect = 0.25, Method: Compositional matrix adjust.
Identities = 16/44 (36%), Positives = 24/44 (54%), Gaps = 2/44 (4%)
Query 128 ADY--YFNSYAHFSIHEEMIKDSVRTCSYQRAIMDNQHLFRGKV 169
DY YF +Y+ ++M+ D VR +Y A+ N+H F GK
Sbjct 30 VDYAQYFCTYSFLYHQKDMLSDRVRMDAYFNAVFQNKHHFEGKT 73
> At1g56675
Length=1486
Score = 31.6 bits (70), Expect = 0.71, Method: Composition-based stats.
Identities = 17/70 (24%), Positives = 35/70 (50%), Gaps = 4/70 (5%)
Query 10 DRVCFSC----EFLDTSLILFGRFPRAQASVDLRSVKAASVSILAQMESGQNHQKGALPS 65
+RVC +C + L G P + + L++ ++S L+ + Q+H +G+ +
Sbjct 255 NRVCSNCGRVGHLAEQCFKLIGYPPWLEEKLRLKNTASSSRGGLSSFKGKQSHGRGSSIN 314
Query 66 HAKGNGISAN 75
H +G++AN
Sbjct 315 HVASSGMAAN 324
> CE25002
Length=4927
Score = 31.2 bits (69), Expect = 1.1, Method: Composition-based stats.
Identities = 13/34 (38%), Positives = 17/34 (50%), Gaps = 0/34 (0%)
Query 64 PSHAKGNGISANNGGSQTGNGSVRGGRGGRPAYA 97
P+ GI A NG ++T NGS G G +A
Sbjct 686 PTQVTNGGIKAANGSAKTANGSANGSANGSAVHA 719
> CE28729
Length=4203
Score = 29.6 bits (65), Expect = 3.3, Method: Composition-based stats.
Identities = 13/30 (43%), Positives = 16/30 (53%), Gaps = 0/30 (0%)
Query 68 KGNGISANNGGSQTGNGSVRGGRGGRPAYA 97
K GI A NG ++T NGS G G +A
Sbjct 14 KNGGIKAANGSAKTANGSANGSANGSAVHA 43
> Hs20532873
Length=981
Score = 29.3 bits (64), Expect = 4.4, Method: Composition-based stats.
Identities = 13/41 (31%), Positives = 23/41 (56%), Gaps = 0/41 (0%)
Query 107 PEEKAAFSADWKQLQQDDISSADYYFNSYAHFSIHEEMIKD 147
P EK F++ + + D+IS AD+ F+ H+ + E K+
Sbjct 47 PLEKGIFTSGTWEEKSDEISFADFKFSVTHHYLVQESTDKE 87
> At2g36760
Length=496
Score = 28.5 bits (62), Expect = 6.5, Method: Compositional matrix adjust.
Identities = 10/36 (27%), Positives = 19/36 (52%), Gaps = 0/36 (0%)
Query 94 PAYAQSLQCQKLSPEEKAAFSADWKQLQQDDISSAD 129
P++ ++ KL K FS DWK++ + + + D
Sbjct 184 PSFPDRVEFTKLQVTVKTNFSGDWKEIMDEQVDADD 219
Lambda K H
0.318 0.130 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2454498488
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40