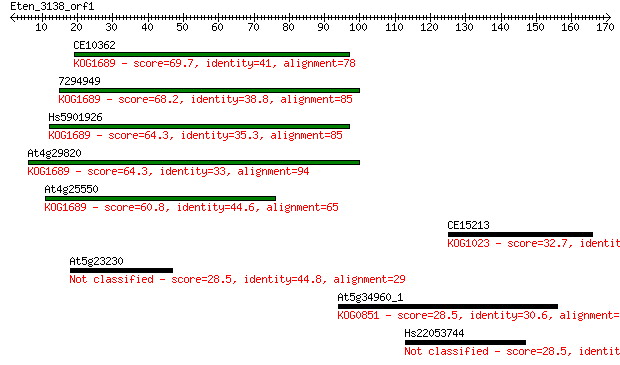
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_3138_orf1
Length=170
Score E
Sequences producing significant alignments: (Bits) Value
CE10362 69.7 2e-12
7294949 68.2 7e-12
Hs5901926 64.3 1e-10
At4g29820 64.3 1e-10
At4g25550 60.8 1e-09
CE15213 32.7 0.37
At5g23230 28.5 6.0
At5g34960_1 28.5 6.5
Hs22053744 28.5 7.5
> CE10362
Length=227
Score = 69.7 bits (169), Expect = 2e-12, Method: Compositional matrix adjust.
Identities = 32/80 (40%), Positives = 45/80 (56%), Gaps = 2/80 (2%)
Query 19 LGEWWRGEFDDDLVPYLPPHVTRPKERVRVHQVQLPHRCSFRIPVGFCLTVVPIFEL--S 76
+G WWR FD PY+P HVT+PKE ++ VQLP + +F +P F L P+FEL +
Sbjct 142 IGNWWRPNFDPPRYPYIPAHVTKPKEHTKLLLVQLPSKSTFCVPKNFKLVAAPLFELYDN 201
Query 77 PQRVGLAIGGLSHLIARFSI 96
G I L ++RF+
Sbjct 202 AAAYGPLISSLPTTLSRFNF 221
> 7294949
Length=237
Score = 68.2 bits (165), Expect = 7e-12, Method: Compositional matrix adjust.
Identities = 33/87 (37%), Positives = 47/87 (54%), Gaps = 2/87 (2%)
Query 15 VGEFLGEWWRGEFDDDLVPYLPPHVTRPKERVRVHQVQLPHRCSFRIPVGFCLTVVPIFE 74
V + +G WWR F+ PY+PPH+T+PKE R+ VQL + F +P + L P+FE
Sbjct 151 VEDTIGNWWRPNFEPPQYPYIPPHITKPKEHKRLFLVQLHEKALFAVPKNYKLVAAPLFE 210
Query 75 L--SPQRVGLAIGGLSHLIARFSIRLM 99
L + Q G I L + RF+ M
Sbjct 211 LYDNSQGYGPIISSLPQALCRFNFIYM 237
> Hs5901926
Length=227
Score = 64.3 bits (155), Expect = 1e-10, Method: Compositional matrix adjust.
Identities = 30/87 (34%), Positives = 46/87 (52%), Gaps = 2/87 (2%)
Query 12 DLEVGEFLGEWWRGEFDDDLVPYLPPHVTRPKERVRVHQVQLPHRCSFRIPVGFCLTVVP 71
D + + +G WWR F+ PY+P H+T+PKE ++ VQL + F +P + L P
Sbjct 138 DWVIDDCIGNWWRPNFEPPQYPYIPAHITKPKEHKKLFLVQLQEKALFAVPKNYKLVAAP 197
Query 72 IFELSPQRVGLA--IGGLSHLIARFSI 96
+FEL G I L L++RF+
Sbjct 198 LFELYDNAPGYGPIISSLPQLLSRFNF 224
> At4g29820
Length=185
Score = 64.3 bits (155), Expect = 1e-10, Method: Compositional matrix adjust.
Identities = 31/96 (32%), Positives = 55/96 (57%), Gaps = 2/96 (2%)
Query 6 LSTDGPDLEVGEFLGEWWRGEFDDDLVPYLPPHVTRPKERVRVHQVQLPHRCSFRIPVGF 65
L++ ++ VGE +G WWR F+ + P+LPP++ PKE ++ V+LP F +P F
Sbjct 88 LASLCINIAVGECIGMWWRPNFETLMYPFLPPNIKHPKECTKLFLVRLPVHQQFVVPKNF 147
Query 66 CLTVVPIFEL--SPQRVGLAIGGLSHLIARFSIRLM 99
L VP+ +L + + G + + L+++FS +M
Sbjct 148 KLLAVPLCQLHENEKTYGPIMSQIPKLLSKFSFNMM 183
> At4g25550
Length=210
Score = 60.8 bits (146), Expect = 1e-09, Method: Compositional matrix adjust.
Identities = 29/71 (40%), Positives = 39/71 (54%), Gaps = 6/71 (8%)
Query 11 PDLEVGEFLGEWWRGEFDDDLVPYLPPHVTRPK------ERVRVHQVQLPHRCSFRIPVG 64
PD VGE + WWR F+ + PY PPH+T+PK E R++ V L + F +P
Sbjct 123 PDWTVGECVATWWRPNFETMMYPYCPPHITKPKVVKKHNECKRLYIVHLSEKEYFAVPKN 182
Query 65 FCLTVVPIFEL 75
L VP+FEL
Sbjct 183 LKLLAVPLFEL 193
> CE15213
Length=1170
Score = 32.7 bits (73), Expect = 0.37, Method: Composition-based stats.
Identities = 17/47 (36%), Positives = 27/47 (57%), Gaps = 7/47 (14%)
Query 125 TSFDIHELEEQPDV------AGGHSEAKPALENDGELEMLPPELQHL 165
++FD+ +E+PDV GGH+ +P+L+ GE + P L HL
Sbjct 792 SAFDLENRKEKPDVIIYQVKKGGHNPMRPSLDT-GETVEINPALLHL 837
> At5g23230
Length=198
Score = 28.5 bits (62), Expect = 6.0, Method: Compositional matrix adjust.
Identities = 13/35 (37%), Positives = 18/35 (51%), Gaps = 6/35 (17%)
Query 18 FLGEWWRGEF------DDDLVPYLPPHVTRPKERV 46
LGEWW G+ D +++P + VT P E V
Sbjct 72 MLGEWWNGDLILDGTTDSEIIPEINRQVTGPDEIV 106
> At5g34960_1
Length=591
Score = 28.5 bits (62), Expect = 6.5, Method: Composition-based stats.
Identities = 19/62 (30%), Positives = 28/62 (45%), Gaps = 1/62 (1%)
Query 94 FSIRLMVPTLDKYEADTSLEHKGEAGLDDLGTSFDIHELEEQPDVAGGHSEAKPALENDG 153
FS + +P L SL+ AG+ L SFD ++ PD H ++ L NDG
Sbjct 162 FSFFVPIPFLSILHDLWSLKCYETAGVKSLSNSFDASQVHVNPDFPEAHHFSQ-TLPNDG 220
Query 154 EL 155
+
Sbjct 221 AI 222
> Hs22053744
Length=679
Score = 28.5 bits (62), Expect = 7.5, Method: Compositional matrix adjust.
Identities = 11/34 (32%), Positives = 18/34 (52%), Gaps = 0/34 (0%)
Query 113 EHKGEAGLDDLGTSFDIHELEEQPDVAGGHSEAK 146
+H+ + G+ G S+ +H QP G HSE +
Sbjct 54 DHRAQQGVQRQGVSYSVHAYTGQPSPRGLHSENR 87
Lambda K H
0.318 0.140 0.430
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2490594054
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40