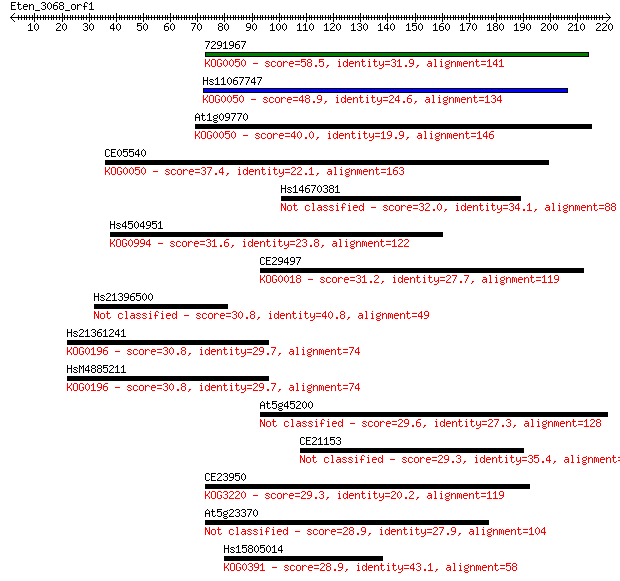
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_3068_orf1
Length=221
Score E
Sequences producing significant alignments: (Bits) Value
7291967 58.5 1e-08
Hs11067747 48.9 8e-06
At1g09770 40.0 0.003
CE05540 37.4 0.021
Hs14670381 32.0 0.91
Hs4504951 31.6 1.4
CE29497 31.2 1.7
Hs21396500 30.8 2.3
Hs21361241 30.8 2.4
HsM4885211 30.8 2.4
At5g45200 29.6 5.1
CE21153 29.3 7.0
CE23950 29.3 7.1
At5g23370 28.9 7.9
Hs15805014 28.9 8.3
> 7291967
Length=791
Score = 58.5 bits (140), Expect = 1e-08, Method: Compositional matrix adjust.
Identities = 45/155 (29%), Positives = 75/155 (48%), Gaps = 14/155 (9%)
Query 73 QLVFNPASKSYVSSSTLSPSERLQVYAHQVETLKTGLASAVQRTKKLEKKYKVLTTGYTN 132
Q+++ P+ Y ++ S +RL+ ++ET + +A +R K+EKK K+LT GY
Sbjct 637 QVLYLPSQHRYTRANLASKKDRLESAEKRLETNRRHMAKEAKRCGKIEKKLKILTGGYQA 696
Query 133 RQKELAKAISESSAELLQLDARIASFTKLHEAEQKAVKVRTEEKRVEVLRERAKHLFLQQ 192
R + L K + ++ ++ Q +++F L E E AV R E + +V R+ + LQQ
Sbjct 697 RAQVLIKQLQDTYGQIEQNSVSLSTFRFLGEQEAIAVPRRLESLQEDVRRQMDREKELQQ 756
Query 193 LYGRL--------------TAVKKHLQELLSDTGA 213
Y L T V+ Q+LL D A
Sbjct 757 KYASLVEERDSLYSQIEHITGVRPTAQQLLPDQEA 791
> Hs11067747
Length=802
Score = 48.9 bits (115), Expect = 8e-06, Method: Compositional matrix adjust.
Identities = 33/134 (24%), Positives = 68/134 (50%), Gaps = 0/134 (0%)
Query 72 TQLVFNPASKSYVSSSTLSPSERLQVYAHQVETLKTGLASAVQRTKKLEKKYKVLTTGYT 131
+Q+++ P Y ++ S +R++ ++E + + + +R K+EKK K+L GY
Sbjct 666 SQVLYLPGQSRYTRANLASKKDRIESLEKRLEINRGHMTTEAKRAAKMEKKMKILLGGYQ 725
Query 132 NRQKELAKAISESSAELLQLDARIASFTKLHEAEQKAVKVRTEEKRVEVLRERAKHLFLQ 191
+R L K +++ ++ Q + +F +L + E A+ R E + +V R++ + LQ
Sbjct 726 SRAMGLMKQLNDLWDQIEQAHLELRTFEELKKHEDSAIPRRLECLKEDVQRQQEREKELQ 785
Query 192 QLYGRLTAVKKHLQ 205
Y L K+ L+
Sbjct 786 HRYADLLLEKETLK 799
> At1g09770
Length=844
Score = 40.0 bits (92), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 29/146 (19%), Positives = 66/146 (45%), Gaps = 0/146 (0%)
Query 69 TLETQLVFNPASKSYVSSSTLSPSERLQVYAHQVETLKTGLASAVQRTKKLEKKYKVLTT 128
T L++ P +Y SS ++++ + ++E ++ + ++ + ++ KYK T
Sbjct 664 TCVNDLMYFPTRSAYELSSVAGNADKVAAFQEEMENVRKKMEEDEKKAEHMKAKYKTYTK 723
Query 129 GYTNRQKELAKAISESSAELLQLDARIASFTKLHEAEQKAVKVRTEEKRVEVLRERAKHL 188
G+ R + + I + + + F L E+ A R + + EV++++
Sbjct 724 GHERRAETVWTQIEATLKQAEIGGTEVECFKALKRQEEMAASFRKKNLQEEVIKQKETES 783
Query 189 FLQQLYGRLTAVKKHLQELLSDTGAQ 214
LQ YG + A+ + +E++ AQ
Sbjct 784 KLQTRYGNMLAMVEKAEEIMVGFRAQ 809
> CE05540
Length=755
Score = 37.4 bits (85), Expect = 0.021, Method: Compositional matrix adjust.
Identities = 36/165 (21%), Positives = 72/165 (43%), Gaps = 8/165 (4%)
Query 36 SLEQMQAVRELLETELASWNLDENYPRFDKLVLTLETQLVFNP--ASKSYVSSSTLSPSE 93
S E++ A +L++ E E+ P + L+ + Q + + + L E
Sbjct 573 SREELDAAADLIKQEA------ESGPELNSLMWKVVEQCTSEIILSKDKFTRIAILPREE 626
Query 94 RLQVYAHQVETLKTGLASAVQRTKKLEKKYKVLTTGYTNRQKELAKAISESSAELLQLDA 153
+++ + + + + +R K+EKK +V GY +L K E + E+ +
Sbjct 627 QMKALNDEFQMYRGWMNQRAKRAAKVEKKLRVKLGGYQAIHDKLCKKYQEVTTEIEMANI 686
Query 154 RIASFTKLHEAEQKAVKVRTEEKRVEVLRERAKHLFLQQLYGRLT 198
+F +L E E KA+ R + EV + + LQ++Y +L+
Sbjct 687 EKKTFERLGEHELKAINKRVGRLQQEVTTQETREKDLQKMYSKLS 731
> Hs14670381
Length=3113
Score = 32.0 bits (71), Expect = 0.91, Method: Compositional matrix adjust.
Identities = 30/90 (33%), Positives = 45/90 (50%), Gaps = 6/90 (6%)
Query 101 QVETLKTGLASAVQRTKKLEKKYKVLTTGYTNRQKELAKAISESSAELLQLDARIASFTK 160
+VETLKT + + K E L + N L K I E +L +LD ++SF
Sbjct 2182 EVETLKTQIEEMARSLKVFELDLVTLRSEKEN----LTKQIQEKQGQLSELDKLLSSFKS 2237
Query 161 -LHEAEQKAVKVRTEEKR-VEVLRERAKHL 188
L E EQ ++++ E K VE+L+ + K L
Sbjct 2238 LLEEKEQAEIQIKEESKTAVEMLQNQLKEL 2267
> Hs4504951
Length=1786
Score = 31.6 bits (70), Expect = 1.4, Method: Composition-based stats.
Identities = 29/122 (23%), Positives = 56/122 (45%), Gaps = 19/122 (15%)
Query 38 EQMQAVRELLETELASWNLDENYPRFDKLVLTLETQLVFNPASKSYVSSSTLSPSERLQV 97
+ + V++ L+ EL DE Y + + L+ A K+ S+ +E LQ
Sbjct 1681 QSAEDVKKTLDGEL-----DEKYKKVENLI-----------AKKTEESADARRKAEMLQ- 1723
Query 98 YAHQVETLKTGLASAVQRTKKLEKKYKVLTTGYTNRQKELAKAISESSAELLQLDARIAS 157
++ +TL S +Q K LE+KY+ ++ +ELA+ E + L + ++A
Sbjct 1724 --NEAKTLLAQANSKLQLLKDLERKYEDNQRYLEDKAQELARLEGEVRSLLKDISQKVAV 1781
Query 158 FT 159
++
Sbjct 1782 YS 1783
> CE29497
Length=1281
Score = 31.2 bits (69), Expect = 1.7, Method: Compositional matrix adjust.
Identities = 33/138 (23%), Positives = 61/138 (44%), Gaps = 26/138 (18%)
Query 93 ERLQVYAHQVETLKTGLASAVQRTKKLEKKYKVLTTGYTNRQKELA---KAISESSAELL 149
ER++ V ++T +A+ QR K+ E++ K+L + ++ A +E + EL
Sbjct 429 ERVKAKEGDVRRIETQIATLAQRIKETEEETKILKADLKKIENDVVIDKSAAAEYNKEL- 487
Query 150 QLDARIASFTKLHEAEQKAVKVRTEEKRVEVLRERAKHLFLQQLYGRLTA---------- 199
+A +L EA + + ++R E L E K F + +YGRL
Sbjct 488 -----VAVVRQLSEASGDSAEGERNQRRTEAL-EGLKKNFPESVYGRLVDLCQPSHKRFN 541
Query 200 ------VKKHLQELLSDT 211
++KH+ ++ DT
Sbjct 542 IATTKILQKHMNSIVCDT 559
> Hs21396500
Length=556
Score = 30.8 bits (68), Expect = 2.3, Method: Compositional matrix adjust.
Identities = 20/49 (40%), Positives = 27/49 (55%), Gaps = 3/49 (6%)
Query 32 LEPCSLEQMQAVRELLETELASWNLDENYPRFDKLVLTLETQLVFNPAS 80
LEP E+ QA++ L+E EL +DE R DKL LT + + P S
Sbjct 42 LEP---EEKQALKRLVEEELLKMQVDEAASREDKLDLTKKGKRPPTPCS 87
> Hs21361241
Length=983
Score = 30.8 bits (68), Expect = 2.4, Method: Composition-based stats.
Identities = 22/75 (29%), Positives = 33/75 (44%), Gaps = 13/75 (17%)
Query 22 GISPPPPTGELEPCSLEQMQAVRELLETELASWNLDEN-YPRFDKLVLTLETQLVFNPAS 80
G PPP M L + L W D N P+F+++V L+ +L+ NP S
Sbjct 840 GYRLPPP-----------MDCPAALYQLMLDCWQKDRNNRPKFEQIVSILD-KLIRNPGS 887
Query 81 KSYVSSSTLSPSERL 95
++S+ PS L
Sbjct 888 LKIITSAAARPSNLL 902
> HsM4885211
Length=983
Score = 30.8 bits (68), Expect = 2.4, Method: Composition-based stats.
Identities = 22/75 (29%), Positives = 33/75 (44%), Gaps = 13/75 (17%)
Query 22 GISPPPPTGELEPCSLEQMQAVRELLETELASWNLDEN-YPRFDKLVLTLETQLVFNPAS 80
G PPP M L + L W D N P+F+++V L+ +L+ NP S
Sbjct 840 GYRLPPP-----------MDCPAALYQLMLDCWQKDRNNRPKFEQIVSILD-KLIRNPGS 887
Query 81 KSYVSSSTLSPSERL 95
++S+ PS L
Sbjct 888 LKIITSAAARPSNLL 902
> At5g45200
Length=1261
Score = 29.6 bits (65), Expect = 5.1, Method: Compositional matrix adjust.
Identities = 35/135 (25%), Positives = 59/135 (43%), Gaps = 13/135 (9%)
Query 93 ERLQVYAHQVETLKTGLASAVQRTKKLEKKYKVLTTGYTNRQ---KELAKAISESSAELL 149
ER+ V+ ET+ TGL + QR ++ + V+++ YT Q EL K A
Sbjct 45 ERINVFIDTRETMGTGLENLFQRIQESKIAIVVISSRYTESQWCLNELVKIKECVEA--- 101
Query 150 QLDARIASFTKLHEAEQKAVKVRT----EEKRVEVLRERAKHLFLQQLYGRLTAVKKHLQ 205
+ F ++ + K V+ T E+ VLR ++ +Q +T+
Sbjct 102 ---GTLVVFPVFYKVDVKIVRFLTGSFGEKLETLVLRHSERYEPWKQALEFVTSKTGKRV 158
Query 206 ELLSDTGAQQEQQIQ 220
E SD GA+ EQ ++
Sbjct 159 EENSDEGAEVEQIVE 173
> CE21153
Length=705
Score = 29.3 bits (64), Expect = 7.0, Method: Compositional matrix adjust.
Identities = 29/87 (33%), Positives = 44/87 (50%), Gaps = 12/87 (13%)
Query 108 GLASAVQRTKKLEKKYKVLTTGYTNRQKELAKAISE---SSAELLQ-LDARIASFTKLHE 163
LAS+ + ++LEK KVL + T +K AI E SS++L+Q LD + +
Sbjct 374 DLASSRDKAERLEKTLKVLKSELTESEKAHTTAIDELQSSSSKLIQRLDEELRLMRSSRD 433
Query 164 -AEQKAVKVRTEEKRVEVLRERAKHLF 189
AEQK K +E+ +E+ HL
Sbjct 434 TAEQKI-------KDIEIAKEKVDHLL 453
> CE23950
Length=237
Score = 29.3 bits (64), Expect = 7.1, Method: Compositional matrix adjust.
Identities = 24/124 (19%), Positives = 54/124 (43%), Gaps = 5/124 (4%)
Query 73 QLVFNPASKSYVSSSTLSPSERLQVYAHQVETLKTGLASAVQRTKKL-----EKKYKVLT 127
+L+F+ K + P+ R +++ + L TG V T L +K
Sbjct 83 KLIFSNPEKRKALNGITHPAIRWEMFKQFLTLLITGTKYIVFDTPLLFESGYDKWIGTTI 142
Query 128 TGYTNRQKELAKAISESSAELLQLDARIASFTKLHEAEQKAVKVRTEEKRVEVLRERAKH 187
+ + ++E+ + ++ + ++RI + + E +++A V ++ LRE+ KH
Sbjct 143 VVWCDFEQEVERMVTRDNISRADAESRIHAQMDIEEKKKRAKIVIDNNGNIDELREKVKH 202
Query 188 LFLQ 191
+ Q
Sbjct 203 VIAQ 206
> At5g23370
Length=219
Score = 28.9 bits (63), Expect = 7.9, Method: Compositional matrix adjust.
Identities = 29/106 (27%), Positives = 47/106 (44%), Gaps = 7/106 (6%)
Query 73 QLVFNPASKSYVSSSTLSPS--ERLQVYAHQVETLKTGLASAVQRTKKLEKKYKVLTTGY 130
QL+ PA K+ + P+ +LQ+ + TG ++ R KK + T G
Sbjct 9 QLITFPAVKTSPAGYLPDPASINKLQIPTSSKFSFLTGKGKSMLRKKKNDS----FTNGV 64
Query 131 TNRQKELAKAISESSAELLQLDARIASFTKLHEAEQKAVKVRTEEK 176
+ Q +L ++E+ L L ARI L + ++ KV EEK
Sbjct 65 RD-QDKLGPKLTETVKRKLSLGARILQMGGLEKIYKRLFKVSDEEK 109
> Hs15805014
Length=3124
Score = 28.9 bits (63), Expect = 8.3, Method: Compositional matrix adjust.
Identities = 25/66 (37%), Positives = 35/66 (53%), Gaps = 8/66 (12%)
Query 80 SKSYVSSSTLSPSERLQVYAHQV--ETLKTGLASAVQRTKK-----LEKKYK-VLTTGYT 131
S+ Y+SS +PSE Q Y H+V + ++RTK+ L KKY+ VL +
Sbjct 1234 SRPYLSSPLRAPSEESQDYYHKVVIRLHRVTQPFILRRTKRDVEKQLTKKYEHVLKCRLS 1293
Query 132 NRQKEL 137
NRQK L
Sbjct 1294 NRQKAL 1299
Lambda K H
0.312 0.127 0.340
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 4183546302
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40