bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_3062_orf1
Length=148
Score E
Sequences producing significant alignments: (Bits) Value
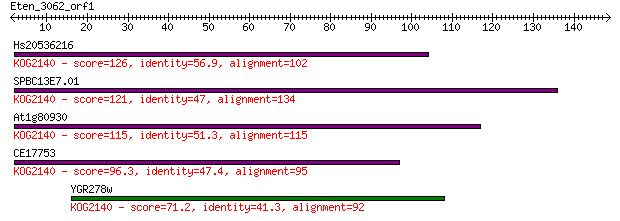
Hs20536216 126 1e-29
SPBC13E7.01 121 5e-28
At1g80930 115 3e-26
CE17753 96.3 2e-20
YGR278w 71.2 7e-13
> Hs20536216
Length=908
Score = 126 bits (317), Expect = 1e-29, Method: Compositional matrix adjust.
Identities = 58/106 (54%), Positives = 75/106 (70%), Gaps = 4/106 (3%)
Query 2 NKYRSIAAAAA----ADAIPWSVLEVFELSEEQTTSSGRIFLKVLLQEISEGLGIKTLNE 57
NK R++A A D++PWSVLE +LSEE TTSS RIF+K+ QE+ E +G+ LN
Sbjct 542 NKLRNVAKMFAHLLYTDSLPWSVLECIKLSEETTTSSSRIFVKIFFQELCEYMGLPKLNA 601
Query 58 RLRHPDLKPYVKGLFPDDHPRNVRFCINFFTAIGLGGLTDHQRQLL 103
RL+ L+P+ +GL P D+PRN RF INFFT+IGLGGLTD R+ L
Sbjct 602 RLKDETLQPFFEGLLPRDNPRNTRFAINFFTSIGLGGLTDELREHL 647
> SPBC13E7.01
Length=628
Score = 121 bits (303), Expect = 5e-28, Method: Compositional matrix adjust.
Identities = 63/138 (45%), Positives = 85/138 (61%), Gaps = 12/138 (8%)
Query 2 NKYRSIAAAAA----ADAIPWSVLEVFELSEEQTTSSGRIFLKVLLQEISEGLGIKTLNE 57
N+ R+IA A D+I W V + L+E+ TT+S RIFLK++ QEI E LG+K+L E
Sbjct 499 NRLRNIALFFANLLSTDSIGWEVYDCVRLTEDDTTASSRIFLKIMFQEIVEALGLKSLVE 558
Query 58 RLRHPDLKPYVKGLFPDDHPRNVRFCINFFTAIGLGGLTDHQRQLLAEMQQQAHLQQQQQ 117
RL P+L PY+ GLFP D RNVRF IN+FT+IGLG LT+ R+ L M +
Sbjct 559 RLHDPNLVPYLHGLFPVDEARNVRFSINYFTSIGLGALTEEMREYLLTM--------PKS 610
Query 118 QLQQQSSSSSSSSSSSSS 135
+ ++Q S SS S + S
Sbjct 611 EPKEQDSEGYSSGSETGS 628
> At1g80930
Length=900
Score = 115 bits (289), Expect = 3e-26, Method: Composition-based stats.
Identities = 59/119 (49%), Positives = 82/119 (68%), Gaps = 4/119 (3%)
Query 2 NKYRSIAAAAA----ADAIPWSVLEVFELSEEQTTSSGRIFLKVLLQEISEGLGIKTLNE 57
NK R++A A DA+PW VL L+EE TTSS RIF+K+L QE+SE LGI+ LNE
Sbjct 737 NKLRNVAKFFAHLLGTDALPWHVLAYIRLTEEDTTSSSRIFIKILFQELSEHLGIRLLNE 796
Query 58 RLRHPDLKPYVKGLFPDDHPRNVRFCINFFTAIGLGGLTDHQRQLLAEMQQQAHLQQQQ 116
RL+ P ++ ++ +FP D+P+N RF INFFT+IGLGG+T++ R+ L M +Q+Q
Sbjct 797 RLQDPTMQESLESIFPKDNPKNTRFAINFFTSIGLGGITENLREYLKNMPSLIMQRQKQ 855
> CE17753
Length=897
Score = 96.3 bits (238), Expect = 2e-20, Method: Compositional matrix adjust.
Identities = 45/95 (47%), Positives = 60/95 (63%), Gaps = 0/95 (0%)
Query 2 NKYRSIAAAAAADAIPWSVLEVFELSEEQTTSSGRIFLKVLLQEISEGLGIKTLNERLRH 61
N R IA + DAI W +L +++EE TTSSGRI++K + E+ E +G+ L+ R+
Sbjct 577 NLARLIAHLLSTDAIDWKILADMKMTEEDTTSSGRIYIKYIFNELVEAMGMVKLHSRVTD 636
Query 62 PDLKPYVKGLFPDDHPRNVRFCINFFTAIGLGGLT 96
P L GLFP +P + RF INFFT IGLGGLT
Sbjct 637 PTLAHCFVGLFPRTNPNSARFSINFFTMIGLGGLT 671
> YGR278w
Length=577
Score = 71.2 bits (173), Expect = 7e-13, Method: Compositional matrix adjust.
Identities = 38/93 (40%), Positives = 55/93 (59%), Gaps = 5/93 (5%)
Query 16 IPWSVLEVFELSEEQTTSSGRIFLKVLLQEISEGLGIKTLNERLRHPDLKPYVKGLFP-D 74
+P L++ +L+EE++ GRIF+K L QE+ LG+ L RL L G+FP +
Sbjct 397 LPMDCLKIIKLTEEESCPQGRIFIKFLFQELVNELGLDELQLRLNSSKLD----GMFPLE 452
Query 75 DHPRNVRFCINFFTAIGLGGLTDHQRQLLAEMQ 107
++R+ INFFTAIGLG LT+ R L +Q
Sbjct 453 GDAEHIRYSINFFTAIGLGLLTEDMRSRLTIIQ 485
Lambda K H
0.310 0.122 0.325
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1821716300
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40