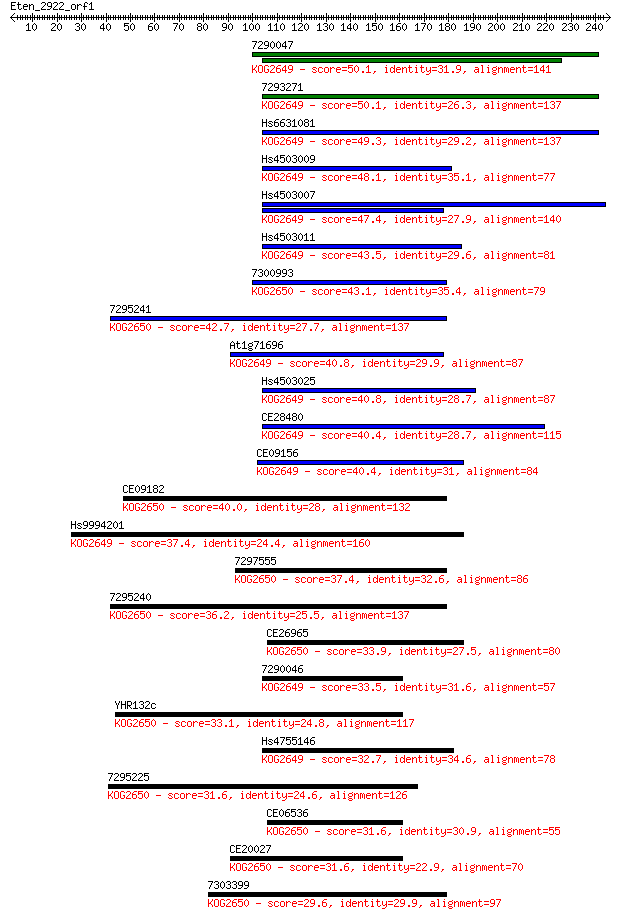
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_2922_orf1
Length=245
Score E
Sequences producing significant alignments: (Bits) Value
7290047 50.1 5e-06
7293271 50.1 5e-06
Hs6631081 49.3 7e-06
Hs4503009 48.1 1e-05
Hs4503007 47.4 3e-05
Hs4503011 43.5 4e-04
7300993 43.1 5e-04
7295241 42.7 7e-04
At1g71696 40.8 0.003
Hs4503025 40.8 0.003
CE28480 40.4 0.003
CE09156 40.4 0.004
CE09182 40.0 0.004
Hs9994201 37.4 0.027
7297555 37.4 0.031
7295240 36.2 0.056
CE26965 33.9 0.31
7290046 33.5 0.41
YHR132c 33.1 0.57
Hs4755146 32.7 0.65
7295225 31.6 1.6
CE06536 31.6 1.8
CE20027 31.6 1.8
7303399 29.6 5.7
> 7290047
Length=1412
Score = 50.1 bits (118), Expect = 5e-06, Method: Compositional matrix adjust.
Identities = 45/163 (27%), Positives = 66/163 (40%), Gaps = 27/163 (16%)
Query 100 NRSTPTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWG 159
N TP + Y +HGDE VG + VYM L I ++ L++S + + P +NP G
Sbjct 88 NLLTPPVKYIANMHGDETVGRQLLVYMAQYLLGNHERISDLGQLVNSTDIYLVPTMNPDG 147
Query 160 F-------------FSNNREEKGVDPNRDFP-YLNSSCFQTL---SSKLVGAAFERYL-- 200
+ + +D NRDFP L S L S + AA ++
Sbjct 148 YALSQEGNCESLPNYVGRGNAANIDLNRDFPDRLEQSHVHQLRAQSRQPETAALVNWIVS 207
Query 201 --FIGGITWHGGMRAIARPW-GSFDHSEPVGRGSFRSRLAPDD 240
F+ +HGG + P+ S H+E L PDD
Sbjct 208 KPFVLSANFHGGAVVASYPYDNSLAHNE-----CCEESLTPDD 245
Score = 39.7 bits (91), Expect = 0.006, Method: Compositional matrix adjust.
Identities = 33/136 (24%), Positives = 55/136 (40%), Gaps = 17/136 (12%)
Query 104 PTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGF--- 160
P Y +HG+EVVG + + + + ++ D + L++ R+ +NP G+
Sbjct 508 PEFKYVANMHGNEVVGKELLLILTKYMLERYGNDDRITKLVNGTRMHFLYSMNPDGYEIS 567
Query 161 FSNNR-------EEKGVDPNRDFP-YLNSSCFQTLSSKLVGAAFERYL---FIGGITWHG 209
+R G+D NR+FP + F ++ V A L F+ HG
Sbjct 568 IEGDRTGGVGRANAHGIDLNRNFPDQYGTDRFNKVTEPEVAAVMNWTLSLPFVLSANLHG 627
Query 210 GMRAIARPWGSFDHSE 225
G P FD +E
Sbjct 628 GSLVANYP---FDDNE 640
> 7293271
Length=518
Score = 50.1 bits (118), Expect = 5e-06, Method: Compositional matrix adjust.
Identities = 36/153 (23%), Positives = 68/153 (44%), Gaps = 16/153 (10%)
Query 104 PTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGFFSN 163
P + Y G +HG+E VG + ++++ + ++ V +LL + R+ + P +NP G+ +
Sbjct 124 PDVKYVGNIHGNEPVGREMLLHLIQYFVTSYNTDQYVKWLLDNTRIHILPTMNPDGYAVS 183
Query 164 NR----------EEKGVDPNRDFP--YLNSSCFQTLSSKLVGAAFERYLFIGGITWHGGM 211
+G D NR+FP + ++ + V + F+ + HGG
Sbjct 184 KEGTCDGGQGRYNARGFDLNRNFPDYFKQNNKRGQPETDSVKDWISKIQFVLSGSLHGGA 243
Query 212 RAIARPWGSFDHSEPVG----RGSFRSRLAPDD 240
+ P+ + +S P+G S L PDD
Sbjct 244 LVASYPYDNTPNSSPLGAVFQTYSAAPSLTPDD 276
> Hs6631081
Length=443
Score = 49.3 bits (116), Expect = 7e-06, Method: Compositional matrix adjust.
Identities = 40/151 (26%), Positives = 67/151 (44%), Gaps = 19/151 (12%)
Query 104 PTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGF--- 160
P Y +HGDE VG + ++++ L + + E+ L++S R+ + P +NP GF
Sbjct 74 PEFKYVANMHGDETVGRELLLHLIDYLVTSDGKDPEITNLINSTRIHIMPSMNPDGFEAV 133
Query 161 ------FSNNREE-KGVDPNRDFPYLNSSCFQTLSSKLVGAAFERYL----FIGGITWHG 209
+S RE D NR+FP ++ + +S + A ++L F+ HG
Sbjct 134 KKPDCYYSIGRENYNQYDLNRNFP--DAFEYNNVSRQPETVAVMKWLKTETFVLSANLHG 191
Query 210 GMRAIARPWGSFDHSEPVGRGSFRSRLAPDD 240
G + P FD+ + L PDD
Sbjct 192 GALVASYP---FDNGVQATGALYSRSLTPDD 219
> Hs4503009
Length=476
Score = 48.1 bits (113), Expect = 1e-05, Method: Compositional matrix adjust.
Identities = 27/91 (29%), Positives = 46/91 (50%), Gaps = 14/91 (15%)
Query 104 PTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDE-VFYLLSSRRLVVFPFLNPWGF-- 160
P Y G +HG+E VG + +++ L +++ + +E + L+ S R+ + P LNP GF
Sbjct 105 PEFKYIGNMHGNEAVGRELLIFLAQYLCNEYQKGNETIVNLIHSTRIHIMPSLNPDGFEK 164
Query 161 -----------FSNNREEKGVDPNRDFPYLN 180
F +G+D NR+FP L+
Sbjct 165 AASQPGELKDWFVGRSNAQGIDLNRNFPDLD 195
> Hs4503007
Length=1377
Score = 47.4 bits (111), Expect = 3e-05, Method: Compositional matrix adjust.
Identities = 39/152 (25%), Positives = 63/152 (41%), Gaps = 18/152 (11%)
Query 104 PTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGFFSN 163
P Y G +HG+EVVG + + ++ L F EV L+ + R+ + P +NP G+ +
Sbjct 552 PEFKYIGNMHGNEVVGRELLLNLIEYLCKNFGTDPEVTDLVHNTRIHLMPSMNPDGYEKS 611
Query 164 N----------REEKGVDPNRDFP--YLNSSCFQTLSSKLVGAAFERYLFIGGITWHGGM 211
D NR+FP ++ + + V + + Y F+ HGG
Sbjct 612 QEGDSISVIGRNNSNNFDLNRNFPDQFVQITDPTQPETIAVMSWMKSYPFVLSANLHGGS 671
Query 212 RAIARPWGSFDHSEPVGRGSFRSRLAPDDAAL 243
+ P FD E +G +PDDA
Sbjct 672 LVVNYP---FDDDE---QGLATYSKSPDDAVF 697
Score = 36.2 bits (82), Expect = 0.061, Method: Compositional matrix adjust.
Identities = 26/91 (28%), Positives = 37/91 (40%), Gaps = 17/91 (18%)
Query 104 PTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGF--- 160
P + G +HGDE V V +Y+ LA+ + LL++ + + P LNP GF
Sbjct 128 PQVKLVGNMHGDETVSRQVLIYLARELAALPPGDPRLVRLLNTTDVYLLPSLNPDGFERA 187
Query 161 --------------FSNNREEKGVDPNRDFP 177
S +G D NR FP
Sbjct 188 REGDCGFGDGGPSGASGRDNSRGRDLNRSFP 218
> Hs4503011
Length=458
Score = 43.5 bits (101), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 24/95 (25%), Positives = 43/95 (45%), Gaps = 14/95 (14%)
Query 104 PTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDE-VFYLLSSRRLVVFPFLNPWGF-- 160
P + Y G +HG+E +G + + + L +F ++ + L+ R+ + P +NP G+
Sbjct 77 PEVKYVGNMHGNEALGRELMLQLSEFLCEEFRNRNQRIVQLIQDTRIHILPSMNPDGYEV 136
Query 161 -----------FSNNREEKGVDPNRDFPYLNSSCF 184
GVD NR+FP LN+ +
Sbjct 137 AAAQGPNKPGYLVGRNNANGVDLNRNFPDLNTYIY 171
> 7300993
Length=467
Score = 43.1 bits (100), Expect = 5e-04, Method: Compositional matrix adjust.
Identities = 28/92 (30%), Positives = 44/92 (47%), Gaps = 15/92 (16%)
Query 100 NRSTPTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWG 159
N + T+F +H E + P A Y++ L + S+ V L S+R +FP +NP G
Sbjct 210 NENRTTVFVESGIHAREWIAPATATYIIDQLVN--SKDSAVQALARSQRWYIFPTVNPDG 267
Query 160 F---FSNNREE----------KGVDPNRDFPY 178
+ F +R +GVD NR+FP+
Sbjct 268 YQYTFKGDRMWRKNRALFGICRGVDLNRNFPF 299
> 7295241
Length=344
Score = 42.7 bits (99), Expect = 7e-04, Method: Compositional matrix adjust.
Identities = 38/153 (24%), Positives = 65/153 (42%), Gaps = 34/153 (22%)
Query 42 TYSSILSLLPLLQQQFPHLIEVKDAVESFGLQD-EVVELTCGRDTCRVPYLELGLRSELN 100
T+S I L L ++FP ++VK S+ + +V+ +T G +
Sbjct 46 THSEINDYLDSLLERFPKRVQVKQFGWSYERRPLKVLTITNG---------------DGR 90
Query 101 RSTPTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGF 160
R+ P I G +H E + P++A+Y++ L + + E LL V+ P +N G+
Sbjct 91 RNKPVILIDGTVHAREWISPSMALYIIQQLLDNYGDNQE---LLQDYDWVIMPVVNADGY 147
Query 161 F---------------SNNREEKGVDPNRDFPY 178
++N E G D NR+F Y
Sbjct 148 EYTHTDSRYWRKSRRPTSNPECIGTDINRNFGY 180
> At1g71696
Length=367
Score = 40.8 bits (94), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 26/88 (29%), Positives = 40/88 (45%), Gaps = 1/88 (1%)
Query 91 LELGLRSELNRSTPTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLV 150
+E+ R + P Y G +HGDE VG + + + + + + ++ + L
Sbjct 103 IEISDRPGEIEAEPAFKYIGNVHGDEPVGRELLLRLANWICDNYKKDPLAQMIVENVHLH 162
Query 151 VFPFLNPWGFFSNNREE-KGVDPNRDFP 177
+ P LNP GF R VD NRDFP
Sbjct 163 IMPSLNPDGFSIRKRNNANNVDLNRDFP 190
> Hs4503025
Length=641
Score = 40.8 bits (94), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 25/101 (24%), Positives = 49/101 (48%), Gaps = 14/101 (13%)
Query 104 PTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEID-EVFYLLSSRRLVVFPFLNPWGF-- 160
P + G +HG+EV G + +Y+ L S++ + + LL++ R+ + P +NP G+
Sbjct 228 PEVKLIGNIHGNEVAGREMLIYLAQYLCSEYLLGNPRIQRLLNTTRIHLLPSINPDGYEV 287
Query 161 -----------FSNNREEKGVDPNRDFPYLNSSCFQTLSSK 190
S + + +D NR+FP L S ++ ++
Sbjct 288 AAAEGAGYNGWTSGRQNAQNLDLNRNFPDLTSEYYRLAETR 328
> CE28480
Length=1013
Score = 40.4 bits (93), Expect = 0.003, Method: Composition-based stats.
Identities = 33/132 (25%), Positives = 55/132 (41%), Gaps = 17/132 (12%)
Query 104 PTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGF--- 160
P + G +HG+EVVG +Y+ +L + + + L+++ R+ + P +NP G+
Sbjct 130 PELKIVGNMHGNEVVGREAVLYLAEILCLNYGKNKYLTDLVNNARIHLMPSMNPDGYEKG 189
Query 161 FSNNR-------EEKGVDPNRDFPYLNSSCFQTLSSK-------LVGAAFERYLFIGGIT 206
F +R VD NR+FP S +T V + Y F+
Sbjct 190 FPGDRISAMGRANANDVDLNRNFPTKFESHRETSGGSEPEKENIAVMKWLQAYPFVLSTN 249
Query 207 WHGGMRAIARPW 218
HGG P+
Sbjct 250 LHGGSLVANYPY 261
> CE09156
Length=472
Score = 40.4 bits (93), Expect = 0.004, Method: Compositional matrix adjust.
Identities = 26/98 (26%), Positives = 44/98 (44%), Gaps = 14/98 (14%)
Query 102 STPTIFYSGLLHGDEVVGPTVAV-YMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGF 160
+ P + G +HG+E +G + + + L + E+ LL+S + + P +NP GF
Sbjct 90 TKPEVKLIGNMHGNEPIGRELLLRFAETLCNGAINNDKEIVQLLNSTSIHILPSMNPDGF 149
Query 161 -------------FSNNREEKGVDPNRDFPYLNSSCFQ 185
+ GVD NRDFP L+S ++
Sbjct 150 ELALGTEPAQRQWLTGRSNINGVDLNRDFPDLDSIFYE 187
> CE09182
Length=323
Score = 40.0 bits (92), Expect = 0.004, Method: Compositional matrix adjust.
Identities = 37/148 (25%), Positives = 64/148 (43%), Gaps = 37/148 (25%)
Query 47 LSLLPLLQQQFPHLIEVKDAVESFGLQDEVVELTCGRDTCRVPYLELGLRSELNRSTPTI 106
+ L L QQ+P+ +++++ S+ GR V + G S P +
Sbjct 1 MKFLNSLAQQYPNDVKLQNIGNSYE----------GRSITAVRIADDG------SSKPIV 44
Query 107 FYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGF------ 160
+ +H E + VA+Y++ + SQ + + LL S +LVV P NP G+
Sbjct 45 WIDAGIHAREWISYNVALYLIYTIVSQPAYRN----LLDSVQLVVVPNTNPDGYEYSRTN 100
Query 161 ----------FSNNREEKGVDPNRDFPY 178
F+N+R G D NR++P+
Sbjct 101 DRMWRKTRSRFTNSR-CAGADANRNYPF 127
> Hs9994201
Length=734
Score = 37.4 bits (85), Expect = 0.027, Method: Compositional matrix adjust.
Identities = 39/175 (22%), Positives = 80/175 (45%), Gaps = 31/175 (17%)
Query 26 AAAVARLINGQHEIPLTYSSILSLLPLLQQQFPHLIEVKDAVESF-GLQDEVVELTCGRD 84
A+ + ++ QH Y ++ L+ +Q+Q P++ + +S+ GL+ V+E++
Sbjct 288 ASGSSDPLDFQHH---NYKAMRKLMKQVQEQCPNITRIYSIGKSYQGLKLYVMEMSD--- 341
Query 85 TCRVPYLELGLRSELNRSTPTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEID-EVFYL 143
+ ELG P + Y +HG+E +G + + ++ L +F + V L
Sbjct 342 --KPGEHELG--------EPEVRYVAGMHGNEALGRELLLLLMQFLCHEFLRGNPRVTRL 391
Query 144 LSSRRLVVFPFLNPWGF------------FSNNR-EEKGVDPNRDFPYLNSSCFQ 185
LS R+ + P +NP G+ ++ R + +D N +F LN+ ++
Sbjct 392 LSEMRIHLLPSMNPDGYEIAYHRGSELVGWAEGRWNNQSIDLNHNFADLNTPLWE 446
> 7297555
Length=424
Score = 37.4 bits (85), Expect = 0.031, Method: Compositional matrix adjust.
Identities = 28/103 (27%), Positives = 41/103 (39%), Gaps = 19/103 (18%)
Query 93 LGLRSELNR---STPTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRL 149
LGLR N +IF +H E +GP A Y L S S+ E+ L S
Sbjct 155 LGLRINHNDGRAEKQSIFLEAGMHAREWIGPATATYFANELLS--SQQQEIMNLARSYVW 212
Query 150 VVFPFLNPWGFFSNNREEK--------------GVDPNRDFPY 178
+ P NP G+ ++ + G DPNR++ +
Sbjct 213 YILPHANPDGYVYTHKTNRMWRKTRSPQDKNCVGTDPNRNWDF 255
> 7295240
Length=424
Score = 36.2 bits (82), Expect = 0.056, Method: Compositional matrix adjust.
Identities = 35/157 (22%), Positives = 60/157 (38%), Gaps = 38/157 (24%)
Query 42 TYSSILSLLPLLQQQFPHLIEVKDAVESFGLQD-EVVELTCGRDTCRVPYLELGLRSELN 100
T+ I++ + L Q+FP + VK S+ + + + +T G
Sbjct 126 THEEIINYIDDLAQRFPSRVFVKTVGWSYEQRVLKTITITNGDGKA-------------- 171
Query 101 RSTPTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGF 160
+ IF G H E + P +Y++ L QF +E +LL V+ P +N G+
Sbjct 172 -NKKVIFMDGGFHAREWISPAAVLYVIDQLVEQF---EENAHLLKDYDWVILPLVNADGY 227
Query 161 -----------FSNNREE--------KGVDPNRDFPY 178
+ R+ G DPNR+F +
Sbjct 228 EHTQTGTLARMWRKTRQPYTYAGQTCYGADPNRNFDF 264
> CE26965
Length=291
Score = 33.9 bits (76), Expect = 0.31, Method: Compositional matrix adjust.
Identities = 22/84 (26%), Positives = 37/84 (44%), Gaps = 8/84 (9%)
Query 106 IFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGF----F 161
I+ G +H E V P+ +Y + L +Q+ + ++ + + P LNP G+
Sbjct 177 IWVDGGIHAREWVSPSTVLYFIHQLVTQYDKDVQIKQFVDQLEWYIVPLLNPDGYEYSRS 236
Query 162 SNNREEKGVDPNRDFPYLNSSCFQ 185
SN+ E + NR P C Q
Sbjct 237 SNDPEIRLWRKNRSPP----KCIQ 256
> 7290046
Length=175
Score = 33.5 bits (75), Expect = 0.41, Method: Compositional matrix adjust.
Identities = 18/57 (31%), Positives = 27/57 (47%), Gaps = 0/57 (0%)
Query 104 PTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGF 160
P + + GDE VG + +YM LA+ + +V LL+ + P NP GF
Sbjct 90 PMVKLVANIQGDEAVGRQMVLYMAEYLATHYDGDPKVQALLNLTEIHFLPTCNPDGF 146
> YHR132c
Length=430
Score = 33.1 bits (74), Expect = 0.57, Method: Compositional matrix adjust.
Identities = 29/117 (24%), Positives = 47/117 (40%), Gaps = 12/117 (10%)
Query 44 SSILSLLPLLQQQFPHLIEVKDAVESFGLQDEVVELTCGRDTCRVPYLELGLRSELNRST 103
+I L LL++ FP L+ AVE G E EL + G + E N
Sbjct 125 DTIYMWLDLLERSFPSLV----AVEHLGRTFEGRELKALHIS--------GNKPESNPEK 172
Query 104 PTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGF 160
TI +G +H E + + + + L +++ + L +V P NP G+
Sbjct 173 KTIVITGGIHAREWISVSTVCWALYQLLNRYGSSKKETKYLDDLDFLVIPVFNPDGY 229
> Hs4755146
Length=1158
Score = 32.7 bits (73), Expect = 0.65, Method: Composition-based stats.
Identities = 27/92 (29%), Positives = 44/92 (47%), Gaps = 14/92 (15%)
Query 104 PTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEID-EVFYLLSSRRLVVFPFLNPWGF-- 160
P Y+ +HG+EV+G + + ++ L ++ + + V L+ R+ + P LNP G+
Sbjct 616 PEFRYTAGIHGNEVLGRELLLLLMQYLCREYRDGNPRVRSLVQDTRIHLVPSLNPDGYEV 675
Query 161 -------FSNNR----EEKGVDPNRDFPYLNS 181
F N E+G D DFP LNS
Sbjct 676 AAQMGSEFGNWALGLWTEEGFDIFEDFPDLNS 707
> 7295225
Length=385
Score = 31.6 bits (70), Expect = 1.6, Method: Compositional matrix adjust.
Identities = 31/130 (23%), Positives = 51/130 (39%), Gaps = 24/130 (18%)
Query 41 LTYSSILSLLPLLQQQFPHLIEVKDAVESF---GLQDEVVELTCGRDTCRVPYLELGLRS 97
L+Y I+ L L + + +KD ++ L+ ++ GR RV
Sbjct 39 LSYDGIMQYLDELALSHSNRVTLKDVARTYENRALKMAIITNGDGRPGKRV--------- 89
Query 98 ELNRSTPTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNP 157
IF LH E + P A+ + L +F+E + LL+ + P NP
Sbjct 90 --------IFLDAALHSREWMTPAAALLTIHKLVVEFAENSD---LLTDYDWHIMPLANP 138
Query 158 WGF-FSNNRE 166
G+ +S N E
Sbjct 139 DGYEYSRNTE 148
> CE06536
Length=373
Score = 31.6 bits (70), Expect = 1.8, Method: Compositional matrix adjust.
Identities = 17/55 (30%), Positives = 27/55 (49%), Gaps = 0/55 (0%)
Query 106 IFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLVVFPFLNPWGF 160
++ G H E VAVY + L + + D++ + + + VFP LNP GF
Sbjct 81 VWLDGGNHAREWPAFHVAVYFIEKLVNGYLSDDKITKYVETLDIYVFPVLNPDGF 135
> CE20027
Length=666
Score = 31.6 bits (70), Expect = 1.8, Method: Compositional matrix adjust.
Identities = 16/70 (22%), Positives = 33/70 (47%), Gaps = 2/70 (2%)
Query 91 LELGLRSELNRSTPTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVFYLLSSRRLV 150
L++G RS + ++ G +H E A+Y + L S++ + ++ + +
Sbjct 175 LKIGARSSHKKRA--VWVDGNIHAREWASSHTALYFINQLVSEYGKDAQITNYVDTLDFY 232
Query 151 VFPFLNPWGF 160
+ P LNP G+
Sbjct 233 IVPCLNPDGY 242
> 7303399
Length=422
Score = 29.6 bits (65), Expect = 5.7, Method: Compositional matrix adjust.
Identities = 29/116 (25%), Positives = 44/116 (37%), Gaps = 28/116 (24%)
Query 82 GRDTCRVPYLELGLRSELNRSTPTIFYSGLLHGDEVVGPTVAVYMVALLASQFSEIDEVF 141
GRD + + L RS IF G +H E + +++ L + SE E+
Sbjct 149 GRD---IKSIRLSKRS----GNKAIFLEGNIHAMEWISSATVTFLLNQLIN--SEDPEMQ 199
Query 142 YLLSSRRLVVFPFLNPWGFFSNNREEK-------------------GVDPNRDFPY 178
L +V P +NP GF + E+ G+D NR+F Y
Sbjct 200 RLSEEYDWIVVPMVNPDGFVYTHEVERLWRKNRRPNGYRNESGDCYGIDMNRNFDY 255
Lambda K H
0.322 0.137 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 4938804274
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40