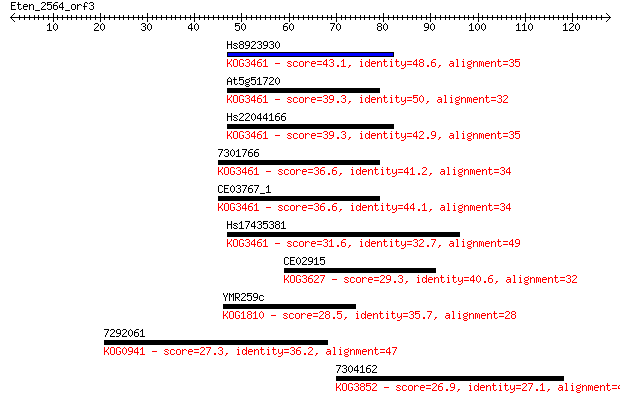
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_2564_orf3
Length=127
Score E
Sequences producing significant alignments: (Bits) Value
Hs8923930 43.1 1e-04
At5g51720 39.3 0.002
Hs22044166 39.3 0.002
7301766 36.6 0.011
CE03767_1 36.6 0.011
Hs17435381 31.6 0.37
CE02915 29.3 1.8
YMR259c 28.5 2.8
7292061 27.3 7.3
7304162 26.9 9.8
> Hs8923930
Length=108
Score = 43.1 bits (100), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 17/36 (47%), Positives = 25/36 (69%), Gaps = 1/36 (2%)
Query 47 CRCWQSTKFPICDNAH-KVLQKQGCNCGPTMLEVRQ 81
CRCW+S KFP CD AH K ++ G N GP +++ ++
Sbjct 72 CRCWRSKKFPFCDGAHTKHNEETGDNVGPLIIKKKE 107
> At5g51720
Length=108
Score = 39.3 bits (90), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 16/33 (48%), Positives = 22/33 (66%), Gaps = 1/33 (3%)
Query 47 CRCWQSTKFPICDNAH-KVLQKQGCNCGPTMLE 78
CRCW+S FP+CD +H K + G N GP +L+
Sbjct 74 CRCWRSGTFPLCDGSHVKHNKANGDNVGPLLLK 106
> Hs22044166
Length=547
Score = 39.3 bits (90), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 15/36 (41%), Positives = 23/36 (63%), Gaps = 1/36 (2%)
Query 47 CRCWQSTKFPICDNAH-KVLQKQGCNCGPTMLEVRQ 81
C CW+ KFP CD +H K ++ G N GP ++E ++
Sbjct 511 CHCWRYKKFPFCDGSHTKHNEQTGDNVGPLIIEKKE 546
> 7301766
Length=133
Score = 36.6 bits (83), Expect = 0.011, Method: Compositional matrix adjust.
Identities = 14/35 (40%), Positives = 22/35 (62%), Gaps = 1/35 (2%)
Query 45 SICRCWQSTKFPICDNAHKVLQKQ-GCNCGPTMLE 78
+ CRCW++ +P CD +H KQ G N GP +++
Sbjct 98 AFCRCWKTKNWPYCDGSHGEHNKQTGDNVGPIVIK 132
> CE03767_1
Length=148
Score = 36.6 bits (83), Expect = 0.011, Method: Compositional matrix adjust.
Identities = 15/35 (42%), Positives = 23/35 (65%), Gaps = 1/35 (2%)
Query 45 SICRCWQSTKFPICDNAHKVLQKQ-GCNCGPTMLE 78
+ CRCW+S K+P CD +H K+ G N GP +++
Sbjct 96 AFCRCWKSEKWPYCDGSHGKHNKETGDNVGPLIVK 130
> Hs17435381
Length=141
Score = 31.6 bits (70), Expect = 0.37, Method: Compositional matrix adjust.
Identities = 16/50 (32%), Positives = 23/50 (46%), Gaps = 9/50 (18%)
Query 47 CRCWQSTKFPICDN-AHKVLQKQGCNCGPTMLEVRQAPSLTEPRDMNSSF 95
C CW+S KFP+CD +HK + C + E R++N F
Sbjct 99 CHCWRSRKFPLCDGVSHKTQGRDWRQC--------ETSDHQEKRNLNGHF 140
> CE02915
Length=331
Score = 29.3 bits (64), Expect = 1.8, Method: Compositional matrix adjust.
Identities = 13/32 (40%), Positives = 18/32 (56%), Gaps = 0/32 (0%)
Query 59 DNAHKVLQKQGCNCGPTMLEVRQAPSLTEPRD 90
DN + V++K GC T +E+ Q S EP D
Sbjct 19 DNKNDVIEKVGCGLHSTNVELAQTRSAQEPAD 50
> YMR259c
Length=1420
Score = 28.5 bits (62), Expect = 2.8, Method: Composition-based stats.
Identities = 10/29 (34%), Positives = 20/29 (68%), Gaps = 1/29 (3%)
Query 46 ICRCWQSTKFPIC-DNAHKVLQKQGCNCG 73
+ + W++T+ +C D+AH +L ++ NCG
Sbjct 623 VLKSWEATRNVVCHDSAHGILPEKYANCG 651
> 7292061
Length=1057
Score = 27.3 bits (59), Expect = 7.3, Method: Composition-based stats.
Identities = 17/51 (33%), Positives = 24/51 (47%), Gaps = 4/51 (7%)
Query 21 CSPADLPSVHTEVLYPTAKRTVNFSICRCWQ----STKFPICDNAHKVLQK 67
CSPAD+ S+ +L P VN + Q + F + +N KVL K
Sbjct 505 CSPADVESMRLYLLLPLYHEFVNSKHYKSLQVPFANAIFKLAENPRKVLNK 555
> 7304162
Length=568
Score = 26.9 bits (58), Expect = 9.8, Method: Composition-based stats.
Identities = 13/48 (27%), Positives = 20/48 (41%), Gaps = 0/48 (0%)
Query 70 CNCGPTMLEVRQAPSLTEPRDMNSSFESKERMSAGALFGGAASVAAAV 117
C C T L Q + T +NSS K+ + GGA + ++
Sbjct 447 CTCNSTWLSTTQVQNQTRSTTINSSNRKKKNKAKTQTKGGAGGITGSL 494
Lambda K H
0.321 0.131 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1176738752
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40