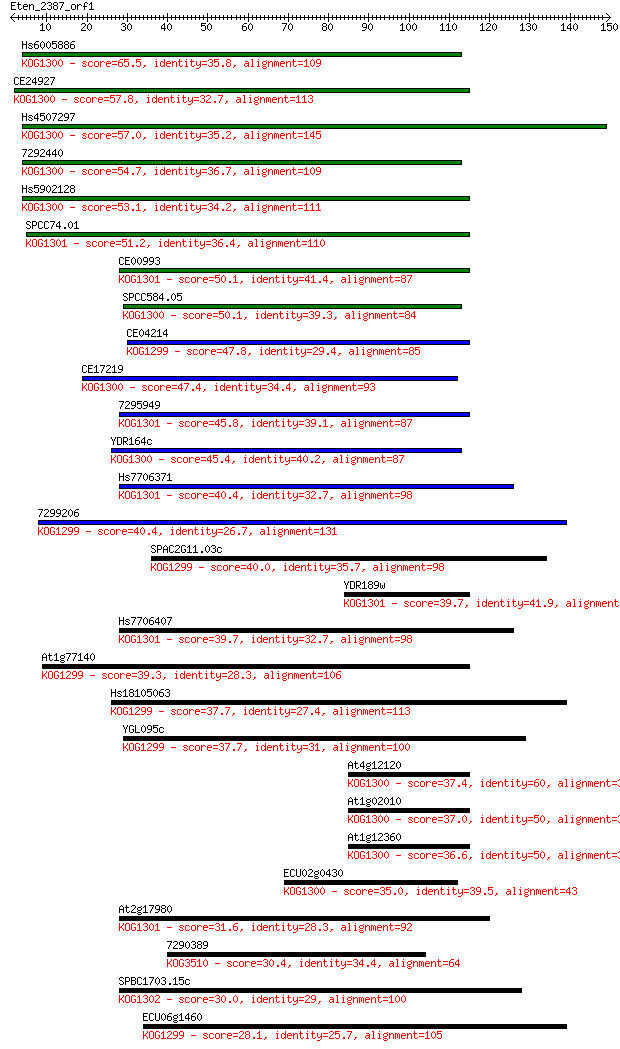
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_2387_orf1
Length=149
Score E
Sequences producing significant alignments: (Bits) Value
Hs6005886 65.5 3e-11
CE24927 57.8 8e-09
Hs4507297 57.0 1e-08
7292440 54.7 6e-08
Hs5902128 53.1 2e-07
SPCC74.01 51.2 7e-07
CE00993 50.1 1e-06
SPCC584.05 50.1 2e-06
CE04214 47.8 7e-06
CE17219 47.4 1e-05
7295949 45.8 3e-05
YDR164c 45.4 4e-05
Hs7706371 40.4 0.001
7299206 40.4 0.001
SPAC2G11.03c 40.0 0.002
YDR189w 39.7 0.002
Hs7706407 39.7 0.002
At1g77140 39.3 0.003
Hs18105063 37.7 0.008
YGL095c 37.7 0.009
At4g12120 37.4 0.011
At1g02010 37.0 0.015
At1g12360 36.6 0.021
ECU02g0430 35.0 0.058
At2g17980 31.6 0.63
7290389 30.4 1.4
SPBC1703.15c 30.0 1.7
ECU06g1460 28.1 5.7
> Hs6005886
Length=592
Score = 65.5 bits (158), Expect = 3e-11, Method: Compositional matrix adjust.
Identities = 39/121 (32%), Positives = 64/121 (52%), Gaps = 13/121 (10%)
Query 4 YEQRVFHFGMPDALFQLFPLPSP-------YLLQKIADDLVSLCVTMGQKPAIRYHRNQL 56
+E +V+ +PDA + + P P +++ +AD +V++C T+ + P +RY L
Sbjct 143 HESQVYTLDVPDAFYYCYS-PDPGNAKGKDAIMETMADQIVTVCATLDENPGVRYKSKPL 201
Query 57 PWCEQLATLVHKGLK-----SEKIPPSDERDTILLILDRSVDLAPLFVHEYTYQALAYDV 111
+LA LV K L+ EK + + LLI+DR D +HE T+QA+AYD+
Sbjct 202 DNASKLAQLVEKKLEDYYKIDEKSLIKGKTHSQLLIIDRGFDPVSTVLHELTFQAMAYDL 261
Query 112 L 112
L
Sbjct 262 L 262
> CE24927
Length=591
Score = 57.8 bits (138), Expect = 8e-09, Method: Compositional matrix adjust.
Identities = 37/123 (30%), Positives = 62/123 (50%), Gaps = 11/123 (8%)
Query 2 TAYEQRVFHFGMPDALFQLFPLPS----PYLLQKIADDLVSLCVTMGQKPAIRYHRNQLP 57
T YE +VF+ PD F + L++IA+ + ++C T+G+ P++RY R
Sbjct 136 TPYESQVFNLDSPDTFFLYYNAQKQGGLTSNLERIAEQIATVCATLGEYPSLRY-RADFE 194
Query 58 WCEQLATLVHKGLKSEKI------PPSDERDTILLILDRSVDLAPLFVHEYTYQALAYDV 111
+L LV + L + K +D+ + L+I+DR D +HE T QA+ YD+
Sbjct 195 RNVELGHLVEQKLDAYKADDPSMGEGADKARSQLIIIDRGYDAITPLLHELTLQAMCYDL 254
Query 112 LEL 114
L +
Sbjct 255 LGI 257
> Hs4507297
Length=603
Score = 57.0 bits (136), Expect = 1e-08, Method: Compositional matrix adjust.
Identities = 51/158 (32%), Positives = 79/158 (50%), Gaps = 15/158 (9%)
Query 4 YEQRVFHFGMPDALFQLFPLPSPY-----LLQKIADDLVSLCVTMGQKPAIRYHRNQLPW 58
YE +V+ D+ FQ F P +L+++A+ + +LC T+ + PA+RY R +
Sbjct 140 YESQVYSLDSADS-FQSFYSPHKAQMKNPILERLAEQIATLCATLKEYPAVRY-RGEYKD 197
Query 59 CEQLATLVHKGLKSEKIP-PS-----DERDTILLILDRSVDLAPLFVHEYTYQALAYDVL 112
LA L+ L + K P+ D+ + LLILDR D + +HE T+QA++YD+L
Sbjct 198 NALLAQLIQDKLDAYKADDPTMGEGPDKARSQLLILDRGFDPSSPVLHELTFQAMSYDLL 257
Query 113 --ELPVCCLNSSKASHSDTPEVLEDVFEYAGIARMLKH 148
E V +S + EVL D + IA KH
Sbjct 258 PIENDVYKYETSGIGEARVKEVLLDEDDDLWIALRHKH 295
> 7292440
Length=597
Score = 54.7 bits (130), Expect = 6e-08, Method: Compositional matrix adjust.
Identities = 40/120 (33%), Positives = 61/120 (50%), Gaps = 13/120 (10%)
Query 4 YEQRVFHFGMPDALFQLFPLPS-----PYLLQKIADDLVSLCVTMGQKPAIRYHRNQLPW 58
YE +VF PD FQ P+ +++IA+ + +LC T+G+ P +RY R+
Sbjct 151 YECQVFSLDSPDT-FQCLYSPAFASIRSKHIERIAEQIATLCATLGEYPNVRY-RSDWDR 208
Query 59 CEQLATLVHKGLKSEKIPPS------DERDTILLILDRSVDLAPLFVHEYTYQALAYDVL 112
LA V + L + K + ++ + LLILDR D +HE T QA+AYD+L
Sbjct 209 NIDLAASVQQKLDAYKADDATMGEGPEKARSQLLILDRGFDCVSPLLHELTLQAMAYDLL 268
> Hs5902128
Length=593
Score = 53.1 bits (126), Expect = 2e-07, Method: Compositional matrix adjust.
Identities = 38/124 (30%), Positives = 61/124 (49%), Gaps = 17/124 (13%)
Query 4 YEQRVFHFGMPDALFQLF-PLPS---PYLLQKIADDLVSLCVTMGQKPAIRYHRNQLPWC 59
YE +VF P + + L+ P + L+ +A + +LC T+ + PAIRY +
Sbjct 140 YEAQVFSLDAPHSTYNLYCPFRAEERTRQLEVLAQQIATLCATLQEYPAIRYRKGP---- 195
Query 60 EQLATLVHKGLKSEKIPPSD---------ERDTILLILDRSVDLAPLFVHEYTYQALAYD 110
E A L H L +D + + LLI+DR+ D +HE T+QA+AYD
Sbjct 196 EDTAQLAHAVLAKLNAFKADTPSLGEGPEKTRSQLLIMDRAADPVSPLLHELTFQAMAYD 255
Query 111 VLEL 114
+L++
Sbjct 256 LLDI 259
> SPCC74.01
Length=639
Score = 51.2 bits (121), Expect = 7e-07, Method: Compositional matrix adjust.
Identities = 40/123 (32%), Positives = 59/123 (47%), Gaps = 16/123 (13%)
Query 5 EQRVFHFGMPDALFQLFPLPSP------YLLQKIADDLVSLCVTMGQKPAIRYHRNQLPW 58
E F +P +F F PS +Q I + L S+ VT+G P IR Q
Sbjct 160 ESDFFSLQLP-KIFHTFHNPSSDEALINSRVQDIVNGLFSVIVTLGTIPIIRCP--QGSA 216
Query 59 CEQLATLVHKGLKSEKIPPSDERDT-------ILLILDRSVDLAPLFVHEYTYQALAYDV 111
E +A +++ LK + D + IL++LDR+VDL P+ H +TYQAL +D
Sbjct 217 AEMVAQKLNQRLKDHLMNTKDAFVSVNPKPRPILILLDRTVDLIPMINHSWTYQALIHDT 276
Query 112 LEL 114
L +
Sbjct 277 LNM 279
> CE00993
Length=707
Score = 50.1 bits (118), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 36/104 (34%), Positives = 50/104 (48%), Gaps = 22/104 (21%)
Query 28 LLQKIADDLVSLCVTMGQKPAIRYHRNQLPWCEQLATLVHKGLKSEKIPPSDERDTI--- 84
+L IAD L ++C TMG P IR + E +A + + L+ D R+ +
Sbjct 211 MLDSIADGLFAVCATMGIVPIIRCPKGNA--AEMVAKKLDQKLRDNL---RDSRNNLFTM 265
Query 85 --------------LLILDRSVDLAPLFVHEYTYQALAYDVLEL 114
L+I DRS DLA + H +TYQAL +DVLEL
Sbjct 266 DGVRMGLLQTSRPLLVIGDRSADLATMLHHTWTYQALMHDVLEL 309
> SPCC584.05
Length=693
Score = 50.1 bits (118), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 33/94 (35%), Positives = 50/94 (53%), Gaps = 10/94 (10%)
Query 29 LQKIADDLVSLCVTMGQKPAIR-YHRNQLPWCEQLATLVHKGLKSE-------KIPPSDE 80
L K+A + S+CV++G P IR Y+ P + + + SE K P E
Sbjct 163 LSKVAHGIFSVCVSLGISPNIRCYYPKNAPHASKTMSFILANQLSEIVEEYCSKHPGYHE 222
Query 81 --RDTILLILDRSVDLAPLFVHEYTYQALAYDVL 112
+ LI+DRS+D A F+HE+TYQA+ +D+L
Sbjct 223 AASKSTCLIVDRSLDTAAPFLHEFTYQAMIHDLL 256
> CE04214
Length=547
Score = 47.8 bits (112), Expect = 7e-06, Method: Composition-based stats.
Identities = 25/86 (29%), Positives = 54/86 (62%), Gaps = 2/86 (2%)
Query 30 QKIADDLVSLCVTMGQKPAIRYHRNQLPWCEQLATLVHKGLKSEKIPPSDER-DTILLIL 88
++I +++L + + + PA+RY ++ P C+++A V + ++ E + R DT LLI+
Sbjct 158 ERIKCGIIALLLQLKKAPAVRYQKSS-PSCKKVADDVAQFIRRENGLFENSRADTTLLII 216
Query 89 DRSVDLAPLFVHEYTYQALAYDVLEL 114
+RS D ++++TY+A+ +++L L
Sbjct 217 ERSQDAVTPLLNQWTYEAMIHEMLTL 242
> CE17219
Length=562
Score = 47.4 bits (111), Expect = 1e-05, Method: Compositional matrix adjust.
Identities = 32/95 (33%), Positives = 48/95 (50%), Gaps = 4/95 (4%)
Query 19 QLFPLPSPYL--LQKIADDLVSLCVTMGQKPAIRYHRNQLPWCEQLATLVHKGLKSEKIP 76
Q+F + S + + K AD +VSLC T+ P +R+ + ++ V + LK E
Sbjct 148 QIFTVNSQFRGDMTKTADGIVSLCATLNIHPTLRFQ-SDFAQSSEICQRVEQKLK-EFGN 205
Query 77 PSDERDTILLILDRSVDLAPLFVHEYTYQALAYDV 111
D L++LDRS DL +HE T QA+ DV
Sbjct 206 EGMGTDAELVVLDRSFDLVSPLLHEVTLQAMVVDV 240
> 7295949
Length=641
Score = 45.8 bits (107), Expect = 3e-05, Method: Compositional matrix adjust.
Identities = 34/103 (33%), Positives = 49/103 (47%), Gaps = 18/103 (17%)
Query 28 LLQKIADDLVSLCVTMGQKPAIRYHRNQLPWCEQLATLVHKGLKSEKIPPSDE------- 80
L+ I D L +L VT+G P IR RN E +A + K L+
Sbjct 164 LMDSIVDSLFALFVTLGNVPIIRCPRNSA--AEMVARKLEKKLRENLWDARANLFHMDAT 221
Query 81 ---------RDTILLILDRSVDLAPLFVHEYTYQALAYDVLEL 114
+ +LL+LDR++DLA H ++YQAL +DVL+L
Sbjct 222 QAGGGVFSFQRPVLLLLDRNMDLATPLHHTWSYQALVHDVLDL 264
> YDR164c
Length=724
Score = 45.4 bits (106), Expect = 4e-05, Method: Compositional matrix adjust.
Identities = 35/111 (31%), Positives = 56/111 (50%), Gaps = 25/111 (22%)
Query 26 PYLLQKIADDLVSLCVTMGQKPAIRYH----------RN------------QLPWCEQLA 63
P ++KI LVSLCV G+ P +RY RN + Q+A
Sbjct 165 PTNVRKIVGSLVSLCVITGEYPIVRYSVSNPVEEEDARNGNAVVNANSLTRSIANAFQIA 224
Query 64 TLVHKGLKSEKIPPSDER-DTILLILDRSVD-LAPLFVHEYTYQALAYDVL 112
+ + P + ER +IL+I DR++D AP+ +H+++YQA+AYD++
Sbjct 225 IDTYARNNPDFPPQNTERPRSILIITDRTLDPFAPI-LHDFSYQAMAYDLV 274
> Hs7706371
Length=640
Score = 40.4 bits (93), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 32/115 (27%), Positives = 53/115 (46%), Gaps = 22/115 (19%)
Query 28 LLQKIADDLVSLCVTMGQKPAIRYHRNQLPWCEQLATLVHKGLKSEKIPPSDERDTI--- 84
++ I D L T+G P IR R E +A + K L+ D R+++
Sbjct 187 VMDTIVDSLFCFYGTLGAVPIIRCSRGTA--AEMVAVKLDKKLREN---LRDARNSLFTG 241
Query 85 --------------LLILDRSVDLAPLFVHEYTYQALAYDVLELPVCCLNSSKAS 125
L+++DR++DLA H +TYQAL +DVL+ + +N ++S
Sbjct 242 DTLGAGQFSFQRPLLVLVDRNIDLATPLHHTWTYQALVHDVLDFHLNRVNLEESS 296
> 7299206
Length=574
Score = 40.4 bits (93), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 35/143 (24%), Positives = 66/143 (46%), Gaps = 26/143 (18%)
Query 8 VFHFGMPDALFQLFPLPSPYLLQKIADDLVSLCVTMGQKPAIRYHRNQLPWCEQLATLVH 67
+F G+P+ + L LP L + + ++ +++ P IRY R + LA L++
Sbjct 137 LFSLGIPNCMANLNWLPDA--LNRSMQGITAVLLSLKLNPVIRY-RAGSQAAQLLAKLIY 193
Query 68 KGLKSEK------------IPPSDERDTILLILDRSVDLAPLFVHEYTYQALAYDVLELP 115
+ + E PP +LL+LDR D +H++TYQA+ +++L +
Sbjct 194 EQITKESSLFDFRSNMDGAAPP------LLLVLDRRDDPVTPLLHQWTYQAMVHELLHIK 247
Query 116 VCCLNSSKASHSDTPEVLEDVFE 138
+++ S+ P V +D E
Sbjct 248 -----NNRLDLSNCPNVPKDFKE 265
> SPAC2G11.03c
Length=558
Score = 40.0 bits (92), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 35/105 (33%), Positives = 58/105 (55%), Gaps = 9/105 (8%)
Query 36 LVSLCVTMGQKPAIRYHRNQLPWCEQLATLVHKGLKSE-KIPPSDERDT--ILLILDRSV 92
++SL +++ +KP IRY N L C +LA V ++ E ++ + DT ILL+LDR
Sbjct 162 IISLLLSLKKKPVIRYDNNSL-LCLKLAEEVSYTIQHESQLFNFRKPDTAPILLLLDRKN 220
Query 93 D-LAPLFVHEYTYQALAYDVLELP---VCCLNSSKASHSDTPEVL 133
D + PL ++TYQA+ +++ + V NS+ + T VL
Sbjct 221 DPITPLLT-QWTYQAMVHELFGIDNGRVSFSNSTSDNEKSTEIVL 264
> YDR189w
Length=666
Score = 39.7 bits (91), Expect = 0.002, Method: Composition-based stats.
Identities = 13/31 (41%), Positives = 23/31 (74%), Gaps = 0/31 (0%)
Query 84 ILLILDRSVDLAPLFVHEYTYQALAYDVLEL 114
+L+ILDR++D A +F H + YQ + +D+ +L
Sbjct 260 VLIILDRNIDFASMFSHSWIYQCMVFDIFKL 290
> Hs7706407
Length=638
Score = 39.7 bits (91), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 32/115 (27%), Positives = 52/115 (45%), Gaps = 22/115 (19%)
Query 28 LLQKIADDLVSLCVTMGQKPAIRYHRNQLPWCEQLATLVHKGLKSEKIPPSDERDTI--- 84
++ I D L T G P IR R E +A + K L+ D R+++
Sbjct 186 VMDTIVDSLFCFYGTRGDVPIIRCSRGTA--AEMVAVKLDKKLREN---LRDARNSLFTG 240
Query 85 --------------LLILDRSVDLAPLFVHEYTYQALAYDVLELPVCCLNSSKAS 125
L+++DR++DLA H +TYQAL +DVL+ + +N ++S
Sbjct 241 DTLGAGQFSFQRPLLVLVDRNIDLATPLHHTWTYQALVHDVLDFHLNRVNLEESS 295
> At1g77140
Length=550
Score = 39.3 bits (90), Expect = 0.003, Method: Composition-based stats.
Identities = 30/117 (25%), Positives = 61/117 (52%), Gaps = 12/117 (10%)
Query 9 FHFGMPDALFQLFPLPS---PYLLQKIADDLV----SLCVTMGQKPAIRYHRNQLPWCEQ 61
+HF + A L+ +P+ P LQ+ +D +V ++ + + ++P IRY R ++
Sbjct 136 YHFTLNMASNHLYMIPAVVDPSGLQRFSDRVVDGIAAVFLALKRRPVIRYQRTS-DTAKR 194
Query 62 LATLVHKGLKSEKIPPSDERDT----ILLILDRSVDLAPLFVHEYTYQALAYDVLEL 114
+A K + + D R T +LL++DR D ++++TYQA+ ++++ L
Sbjct 195 IAHETAKLMYQHESALFDFRRTESSPLLLVIDRRDDPVTPLLNQWTYQAMVHELIGL 251
> Hs18105063
Length=570
Score = 37.7 bits (86), Expect = 0.008, Method: Compositional matrix adjust.
Identities = 31/122 (25%), Positives = 60/122 (49%), Gaps = 21/122 (17%)
Query 26 PYLLQKIADDLVSLCVTMGQKPAIRYHRNQLPWCEQLATLVHKGLKSE---------KIP 76
P L + L +L +++ + P IRY + ++LA V + + E ++P
Sbjct 152 PAQLSRTTQGLTALLLSLKKCPMIRYQLSS-EAAKRLAECVKQVITKEYELFEFRRTEVP 210
Query 77 PSDERDTILLILDRSVDLAPLFVHEYTYQALAYDVLELPVCCLNSSKASHSDTPEVLEDV 136
P +LLILDR D ++++TYQA+ +++L +N+++ S P + +D+
Sbjct 211 P------LLLILDRCDDAITPLLNQWTYQAMVHELL-----GINNNRIDLSRVPGISKDL 259
Query 137 FE 138
E
Sbjct 260 RE 261
> YGL095c
Length=577
Score = 37.7 bits (86), Expect = 0.009, Method: Composition-based stats.
Identities = 31/104 (29%), Positives = 52/104 (50%), Gaps = 5/104 (4%)
Query 29 LQKIADDLVSLCVTMGQKPAIRYHRNQLPWCEQLATLVHKGL-KSEKIP---PSDERDTI 84
L K + LVS+ +++ KP IRY CE+LA V + K+E+ P + +
Sbjct 168 LTKCTNSLVSVLLSLKIKPDIRYE-GASKICERLAKEVSYEIGKNERTFFDFPVMDSTPV 226
Query 85 LLILDRSVDLAPLFVHEYTYQALAYDVLELPVCCLNSSKASHSD 128
LLILDR+ D + +TYQ++ + + + ++ SK D
Sbjct 227 LLILDRNTDPITPLLQPWTYQSMINEYIGIKRNIVDLSKVPRID 270
> At4g12120
Length=635
Score = 37.4 bits (85), Expect = 0.011, Method: Compositional matrix adjust.
Identities = 18/31 (58%), Positives = 26/31 (83%), Gaps = 2/31 (6%)
Query 85 LLILDRSVD-LAPLFVHEYTYQALAYDVLEL 114
LLILDRS+D +APL +HE+TY A+ +D+L +
Sbjct 233 LLILDRSIDQIAPL-IHEWTYDAMCHDLLNM 262
> At1g02010
Length=426
Score = 37.0 bits (84), Expect = 0.015, Method: Compositional matrix adjust.
Identities = 15/30 (50%), Positives = 23/30 (76%), Gaps = 0/30 (0%)
Query 85 LLILDRSVDLAPLFVHEYTYQALAYDVLEL 114
LLI+DRSVD +HE+TY A+ +D+L++
Sbjct 27 LLIVDRSVDQIAPIIHEWTYDAMCHDLLDM 56
> At1g12360
Length=637
Score = 36.6 bits (83), Expect = 0.021, Method: Compositional matrix adjust.
Identities = 15/30 (50%), Positives = 22/30 (73%), Gaps = 0/30 (0%)
Query 85 LLILDRSVDLAPLFVHEYTYQALAYDVLEL 114
LLILDRS+D +HE+TY A+ +D+L +
Sbjct 226 LLILDRSIDQIAPVIHEWTYDAMCHDLLNM 255
> ECU02g0430
Length=498
Score = 35.0 bits (79), Expect = 0.058, Method: Composition-based stats.
Identities = 17/43 (39%), Positives = 27/43 (62%), Gaps = 0/43 (0%)
Query 69 GLKSEKIPPSDERDTILLILDRSVDLAPLFVHEYTYQALAYDV 111
G +EK+ +ER L+ILDRS+DL +H +T++ L D+
Sbjct 165 GEMAEKMMGDEERKGELIILDRSIDLFTPLLHFFTFRTLLEDL 207
> At2g17980
Length=627
Score = 31.6 bits (70), Expect = 0.63, Method: Compositional matrix adjust.
Identities = 26/103 (25%), Positives = 49/103 (47%), Gaps = 13/103 (12%)
Query 28 LLQKIADDLVSLCVTMGQKPAIRYHRNQLPWCEQLATLVHKGLKSEKIPP---------- 77
+++++A L + VT+G P IR E +A+L+ + L+ +
Sbjct 176 IIERVASGLFCVLVTLGVVPVIRCPSG--GPAEMVASLLDQKLRDHLLSKNNLFTEGGGF 233
Query 78 -SDERDTILLILDRSVDLAPLFVHEYTYQALAYDVLELPVCCL 119
S + +L I DR+ +L+ H++ Y+ L +DVL L + L
Sbjct 234 MSSFQRPLLCIFDRNFELSVGIQHDFRYRPLVHDVLGLKLNQL 276
> 7290389
Length=968
Score = 30.4 bits (67), Expect = 1.4, Method: Composition-based stats.
Identities = 22/65 (33%), Positives = 32/65 (49%), Gaps = 10/65 (15%)
Query 40 CVTMGQKPAIRYHRNQLPWCEQLATLVHKGLKSEKIPPSDERDTILLILDRSV-DLAPLF 98
CV+ G KPA ++ W + L ++ KG++ K P +D R I RS+ LAP
Sbjct 209 CVSQGGKPAA-----EITWIDGLGNVLTKGIEYVKEPLADSRR----ITARSILKLAPKK 259
Query 99 VHEYT 103
H T
Sbjct 260 EHHNT 264
> SPBC1703.15c
Length=592
Score = 30.0 bits (66), Expect = 1.7, Method: Compositional matrix adjust.
Identities = 29/115 (25%), Positives = 53/115 (46%), Gaps = 19/115 (16%)
Query 28 LLQKIADDLVSLCVTMGQKPAIRYHRNQLPWCEQLATLVHKGLKSEKI--------PPSD 79
LLQ+ D L+ T G+ P + + P+ ++ L+ K + E S
Sbjct 153 LLQRSTDALIDFERTHGRFPRVS---GRGPYAAKMLELLEKTYQEEATINFGKVEGEISA 209
Query 80 ERDTILLILDRSVDLAPLFVHEYTYQALAYDVL-------ELPVCCLNSSKASHS 127
D++LL+ DRS+D F+ + TY ++L +LP +N ++AS++
Sbjct 210 LYDSVLLV-DRSLDRITPFLTQLTYFGFLDEILGIQQMNVKLPSSLVNRNEASNT 263
> ECU06g1460
Length=490
Score = 28.1 bits (61), Expect = 5.7, Method: Composition-based stats.
Identities = 27/113 (23%), Positives = 51/113 (45%), Gaps = 15/113 (13%)
Query 34 DDLVSLCVTMGQKPAIRYHRNQLPWCEQLATLVHK--GLKSEKIPPSDERDTILLILDRS 91
D + +L + +G+ P I+ E TL + GL ++ L++LDR+
Sbjct 154 DGIFALVMNLGRIPTIKIQTEDRHLMEDSETLSTRLAGLNLKQ-------GGTLIMLDRA 206
Query 92 VDLAPLFVHEYTYQALAYDVLELPVCCLNSSKASHS--DTP----EVLEDVFE 138
DL ++E+ YQ+L ++ + + K S+S D P +D++E
Sbjct 207 FDLYTPLLYEWRYQSLLHEHADYANGIVRIGKKSYSVADDPFFNASKFKDIYE 259
Lambda K H
0.322 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1858150626
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40