bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_1737_orf1
Length=281
Score E
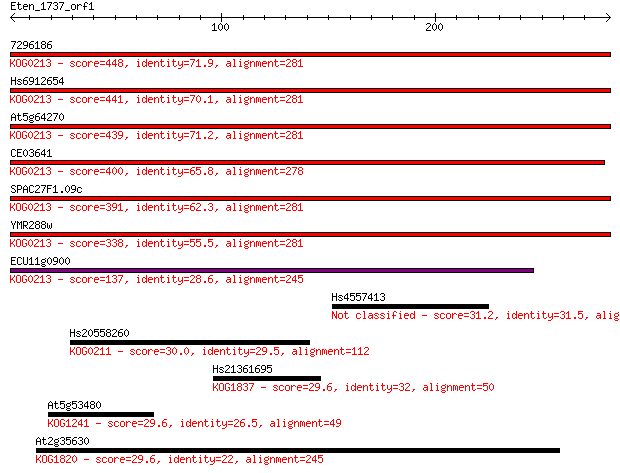
Sequences producing significant alignments: (Bits) Value
7296186 448 5e-126
Hs6912654 441 1e-123
At5g64270 439 2e-123
CE03641 400 2e-111
SPAC27F1.09c 391 9e-109
YMR288w 338 8e-93
ECU11g0900 137 3e-32
Hs4557413 31.2 2.5
Hs20558260 30.0 6.3
Hs21361695 29.6 6.6
At5g53480 29.6 7.0
At2g35630 29.6 7.5
> 7296186
Length=1349
Score = 448 bits (1153), Expect = 5e-126, Method: Compositional matrix adjust.
Identities = 202/281 (71%), Positives = 246/281 (87%), Gaps = 0/281 (0%)
Query 1 LKNRHEKVQENLIDLIGRIADRGGDLVSPKEWDRICFDLLDLLRAQKKSIRRATVNTFGY 60
LKNRHEKVQEN IDL+GRIADRG + VS +EW RICF+LL+LL+A KK+IRRATVNTFGY
Sbjct 1069 LKNRHEKVQENCIDLVGRIADRGPEYVSAREWMRICFELLELLKAHKKAIRRATVNTFGY 1128
Query 61 IARTIGPQDVLATLLNNLKMQERQLRLCTTIAIAIVAETCLPYSVLPALINEYRVPELNI 120
IA+ IGP DVLATLLNNLK+QERQ R+CTT+AIAIVAE+C P++VLPAL+NEYRVPELN+
Sbjct 1129 IAKAIGPHDVLATLLNNLKVQERQNRVCTTVAIAIVAESCRPFTVLPALMNEYRVPELNV 1188
Query 121 QNGVLKTLSFMFEYIGEMAKDYIYTVTPLLEDALMDRDLVHRQTAAWACKHLALGVHGLS 180
QNGVLK+LSF+FEYIGEM KDYIY V PLLEDALMDRDLVHRQTA A KH++LGV+G
Sbjct 1189 QNGVLKSLSFLFEYIGEMGKDYIYAVCPLLEDALMDRDLVHRQTACSAIKHMSLGVYGFG 1248
Query 181 CEDALTHLFNYVWPNIFEVSAHMVQAFFDAVDGMRVAIGPGVVFRYVILGLFHPARKVRE 240
CEDALTHL NYVWPNIFE S H+VQAF D+VDG+RV++GP + +Y + GLFHPARKVR+
Sbjct 1249 CEDALTHLLNYVWPNIFETSPHLVQAFMDSVDGLRVSLGPIKILQYTLQGLFHPARKVRD 1308
Query 241 VYWRVYNNVYIGHQDAMVAFFPRLPDDGKHSFARHELELVI 281
VYW++YN++YIG QDA++A +PR+ +D K+ + R+EL+ +
Sbjct 1309 VYWKIYNSLYIGGQDALIAGYPRITNDPKNQYERYELDYTL 1349
> Hs6912654
Length=1304
Score = 441 bits (1133), Expect = 1e-123, Method: Compositional matrix adjust.
Identities = 197/281 (70%), Positives = 246/281 (87%), Gaps = 0/281 (0%)
Query 1 LKNRHEKVQENLIDLIGRIADRGGDLVSPKEWDRICFDLLDLLRAQKKSIRRATVNTFGY 60
LKNRHEKVQEN IDL+GRIADRG + VS +EW RICF+LL+LL+A KK+IRRATVNTFGY
Sbjct 1024 LKNRHEKVQENCIDLVGRIADRGAEYVSAREWMRICFELLELLKAHKKAIRRATVNTFGY 1083
Query 61 IARTIGPQDVLATLLNNLKMQERQLRLCTTIAIAIVAETCLPYSVLPALINEYRVPELNI 120
IA+ IGP DVLATLLNNLK+QERQ R+CTT+AIAIVAETC P++VLPAL+NEYRVPELN+
Sbjct 1084 IAKAIGPHDVLATLLNNLKVQERQNRVCTTVAIAIVAETCSPFTVLPALMNEYRVPELNV 1143
Query 121 QNGVLKTLSFMFEYIGEMAKDYIYTVTPLLEDALMDRDLVHRQTAAWACKHLALGVHGLS 180
QNGVLK+LSF+FEYIGEM KDYIY VTPLLEDALMDRDLVHRQTA+ +H++LGV+G
Sbjct 1144 QNGVLKSLSFLFEYIGEMGKDYIYAVTPLLEDALMDRDLVHRQTASAVVQHMSLGVYGFG 1203
Query 181 CEDALTHLFNYVWPNIFEVSAHMVQAFFDAVDGMRVAIGPGVVFRYVILGLFHPARKVRE 240
CED+L HL NYVWPN+FE S H++QA A++G+RVAIGP + +Y + GLFHPARKVR+
Sbjct 1204 CEDSLNHLLNYVWPNVFETSPHVIQAVMGALEGLRVAIGPCRMLQYCLQGLFHPARKVRD 1263
Query 241 VYWRVYNNVYIGHQDAMVAFFPRLPDDGKHSFARHELELVI 281
VYW++YN++YIG QDA++A +PR+ +D K+++ R+EL+ ++
Sbjct 1264 VYWKIYNSIYIGSQDALIAHYPRIYNDDKNTYIRYELDYIL 1304
> At5g64270
Length=1269
Score = 439 bits (1130), Expect = 2e-123, Method: Compositional matrix adjust.
Identities = 200/281 (71%), Positives = 240/281 (85%), Gaps = 0/281 (0%)
Query 1 LKNRHEKVQENLIDLIGRIADRGGDLVSPKEWDRICFDLLDLLRAQKKSIRRATVNTFGY 60
LKNRHEKVQEN IDL+GRIADRG + V +EW RICF+LL++L+A KK IRRATVNTFGY
Sbjct 989 LKNRHEKVQENCIDLVGRIADRGAEFVPAREWMRICFELLEMLKAHKKGIRRATVNTFGY 1048
Query 61 IARTIGPQDVLATLLNNLKMQERQLRLCTTIAIAIVAETCLPYSVLPALINEYRVPELNI 120
IA+ IGPQDVLATLLNNLK+QERQ R+CTT+AIAIVAETC P++VLPAL+NEYRVPELN+
Sbjct 1049 IAKAIGPQDVLATLLNNLKVQERQNRVCTTVAIAIVAETCSPFTVLPALMNEYRVPELNV 1108
Query 121 QNGVLKTLSFMFEYIGEMAKDYIYTVTPLLEDALMDRDLVHRQTAAWACKHLALGVHGLS 180
QNGVLK+LSF+FEYIGEM KDYIY VTPLLEDALMDRDLVHRQTAA A KH+ALGV GL
Sbjct 1109 QNGVLKSLSFLFEYIGEMGKDYIYAVTPLLEDALMDRDLVHRQTAASAVKHMALGVAGLG 1168
Query 181 CEDALTHLFNYVWPNIFEVSAHMVQAFFDAVDGMRVAIGPGVVFRYVILGLFHPARKVRE 240
CEDAL HL N++WPNIFE S H++ A +A++GMRVA+G V+ Y + GLFHPARKVRE
Sbjct 1169 CEDALVHLLNFIWPNIFETSPHVINAVMEAIEGMRVALGAAVILNYCLQGLFHPARKVRE 1228
Query 241 VYWRVYNNVYIGHQDAMVAFFPRLPDDGKHSFARHELELVI 281
VYW++YN++YIG QD +VA +P L D+ + ++R EL + +
Sbjct 1229 VYWKIYNSLYIGAQDTLVAAYPVLEDEQNNVYSRPELTMFV 1269
> CE03641
Length=1322
Score = 400 bits (1028), Expect = 2e-111, Method: Compositional matrix adjust.
Identities = 183/278 (65%), Positives = 228/278 (82%), Gaps = 0/278 (0%)
Query 1 LKNRHEKVQENLIDLIGRIADRGGDLVSPKEWDRICFDLLDLLRAQKKSIRRATVNTFGY 60
LKNRHEKVQEN IDL+G IADRG + VS +EW RICF+LL+LL+A KKSIRRA +NTFG+
Sbjct 1042 LKNRHEKVQENCIDLVGAIADRGSEFVSAREWMRICFELLELLKAHKKSIRRAAINTFGF 1101
Query 61 IARTIGPQDVLATLLNNLKMQERQLRLCTTIAIAIVAETCLPYSVLPALINEYRVPELNI 120
IA+ IGP DVLATLLNNLK+QERQLR+CTT+AIAIV+ETC P++VLPA++NEYRVPE+N+
Sbjct 1102 IAKAIGPHDVLATLLNNLKVQERQLRVCTTVAIAIVSETCAPFTVLPAIMNEYRVPEINV 1161
Query 121 QNGVLKTLSFMFEYIGEMAKDYIYTVTPLLEDALMDRDLVHRQTAAWACKHLALGVHGLS 180
QNGVLK LSFMFEYIGEMAKDYIY V PLL DALM+RD VHRQ A A HLA+GV+G
Sbjct 1162 QNGVLKALSFMFEYIGEMAKDYIYAVVPLLIDALMERDQVHRQIAVDAVAHLAIGVYGFG 1221
Query 181 CEDALTHLFNYVWPNIFEVSAHMVQAFFDAVDGMRVAIGPGVVFRYVILGLFHPARKVRE 240
CEDAL HL NYVWPN+ E S H++Q + A +GMRV++GP V +Y + L+HPARKVRE
Sbjct 1222 CEDALIHLLNYVWPNMLENSPHLIQRWVFACEGMRVSLGPIKVLQYCLQALWHPARKVRE 1281
Query 241 VYWRVYNNVYIGHQDAMVAFFPRLPDDGKHSFARHELE 278
W+V+NN+ +G DA++A +PR+ + + + R+EL+
Sbjct 1282 PVWKVFNNLILGSADALIAAYPRIENTPTNQYVRYELD 1319
> SPAC27F1.09c
Length=1188
Score = 391 bits (1004), Expect = 9e-109, Method: Compositional matrix adjust.
Identities = 175/281 (62%), Positives = 223/281 (79%), Gaps = 0/281 (0%)
Query 1 LKNRHEKVQENLIDLIGRIADRGGDLVSPKEWDRICFDLLDLLRAQKKSIRRATVNTFGY 60
L+NRHEKVQEN IDL+G+IADRG + VS +EW RICF+L+D+L+A KKSIRRA VNTFGY
Sbjct 908 LRNRHEKVQENTIDLVGKIADRGSEYVSAREWMRICFELIDMLKAHKKSIRRAAVNTFGY 967
Query 61 IARTIGPQDVLATLLNNLKMQERQLRLCTTIAIAIVAETCLPYSVLPALINEYRVPELNI 120
I++ IGPQDVLATLLNNLK+QERQ R+CTT+AIAIVAETC+P++V+PAL+ +YR PE+N+
Sbjct 968 ISKAIGPQDVLATLLNNLKVQERQNRVCTTVAIAIVAETCMPFTVVPALMADYRTPEMNV 1027
Query 121 QNGVLKTLSFMFEYIGEMAKDYIYTVTPLLEDALMDRDLVHRQTAAWACKHLALGVHGLS 180
QNGVLK+L+FMFEYIGE A+DY+Y +TPLL DALMDRD VHRQTAA KHL+LG GL
Sbjct 1028 QNGVLKSLAFMFEYIGEQARDYVYAITPLLADALMDRDAVHRQTAASVIKHLSLGCVGLG 1087
Query 181 CEDALTHLFNYVWPNIFEVSAHMVQAFFDAVDGMRVAIGPGVVFRYVILGLFHPARKVRE 240
EDA+ HL N +WPNI E S H++ A + +DG+R IG G + Y++ GLFHP+RKVR
Sbjct 1088 VEDAMIHLLNILWPNILEESPHVINAVREGIDGIRNCIGVGPIMAYLVQGLFHPSRKVRN 1147
Query 241 VYWRVYNNVYIGHQDAMVAFFPRLPDDGKHSFARHELELVI 281
YW YN+ Y+ DAMV ++P + DD +++ L + I
Sbjct 1148 TYWTSYNSAYVQSADAMVPYYPHVDDDQFNNYDMKTLHICI 1188
> YMR288w
Length=971
Score = 338 bits (867), Expect = 8e-93, Method: Compositional matrix adjust.
Identities = 156/281 (55%), Positives = 198/281 (70%), Gaps = 2/281 (0%)
Query 1 LKNRHEKVQENLIDLIGRIADRGGDLVSPKEWDRICFDLLDLLRAQKKSIRRATVNTFGY 60
L+N+H KV+ N I +G I PKEW RICF+LL+LL++ K IRR+ TFG+
Sbjct 693 LRNKHRKVEVNTIKFVGLIGKLAPTYAPPKEWMRICFELLELLKSTNKEIRRSANATFGF 752
Query 61 IARTIGPQDVLATLLNNLKMQERQLRLCTTIAIAIVAETCLPYSVLPALINEYRVPELNI 120
IA IGP DVL LLNNLK+QERQLR+CT +AI IVA+ C PY+VLP ++NEY PE N+
Sbjct 753 IAEAIGPHDVLVALLNNLKVQERQLRVCTAVAIGIVAKVCGPYNVLPVIMNEYTTPETNV 812
Query 121 QNGVLKTLSFMFEYIGEMAKDYIYTVTPLLEDALMDRDLVHRQTAAWACKHLALGVHGLS 180
QNGVLK +SFMFEYIG M+KDYIY +TPLLEDAL DRDLVHRQTA+ HLAL G
Sbjct 813 QNGVLKAMSFMFEYIGNMSKDYIYFITPLLEDALTDRDLVHRQTASNVITHLALNCSGTG 872
Query 181 CEDALTHLFNYVWPNIFEVSAHMVQAFFDAVDGMRVAIGPGVVFRYVILGLFHPARKVRE 240
EDA HL N + PNIFE S H + + ++ + A+GPG+ Y+ GLFHPA+ VR+
Sbjct 873 HEDAFIHLMNLLIPNIFETSPHAIMRILEGLEALSQALGPGLFMNYIWAGLFHPAKNVRK 932
Query 241 VYWRVYNNVYIGHQDAMVAFFPRLPDDGKHSFARHELELVI 281
+WRVYNN+Y+ +QDAMV F+P PD+ + EL+LV+
Sbjct 933 AFWRVYNNMYVMYQDAMVPFYPVTPDNNEEYI--EELDLVL 971
> ECU11g0900
Length=903
Score = 137 bits (345), Expect = 3e-32, Method: Compositional matrix adjust.
Identities = 70/249 (28%), Positives = 136/249 (54%), Gaps = 4/249 (1%)
Query 1 LKNRHEKVQENLIDLIGRI---ADRGGDLVSPKEWDRICFDLLDLLRAQKKSIRRATVNT 57
LK++ +K+ + + L+ I + + + +EW RI + L+D L + K +RR +
Sbjct 628 LKSKEQKIVASGVALLHTICMNSPEECEKIGVREWMRISYGLVDSLVSWNKEMRRNATES 687
Query 58 FGYIARTIGPQDVLATLLNNLKMQERQLRLCTTIAIAIVAETCLPYSVLPALINEYRVPE 117
G I+R +GPQ++L L++ L+ ++R R +++ I++V E +SVLP L+++Y P
Sbjct 688 LGCISRIVGPQEILDILMDGLESEDRHQRTGSSLGISVVGEYNGLFSVLPTLLSDYETPN 747
Query 118 LNIQNGVLKTLSFMFEYIGEMAKDYIYTVTPLLEDALMDRDLVHRQTAAWACKHLALGVH 177
+Q G+L+ + F+ + + Y++++ P++EDA+ D D +R +H+ L
Sbjct 748 AFVQQGILRAMCHFFQRTHQASLKYVHSMLPMIEDAMTDEDPSYRSLGMNLIRHIVLNHP 807
Query 178 GLSCEDALT-HLFNYVWPNIFEVSAHMVQAFFDAVDGMRVAIGPGVVFRYVILGLFHPAR 236
+ + L HL N +W NI + + Q+F + ++ + +++YV GLFHP+
Sbjct 808 PATMDIELAIHLLNLIWANILDPIPTVQQSFDECMESFATVLSSQAMYKYVQQGLFHPSS 867
Query 237 KVREVYWRV 245
VR+ Y V
Sbjct 868 TVRKRYCTV 876
> Hs4557413
Length=162
Score = 31.2 bits (69), Expect = 2.5, Method: Compositional matrix adjust.
Identities = 23/76 (30%), Positives = 36/76 (47%), Gaps = 9/76 (11%)
Query 152 DALMDRDLVHRQTAAWACKHLALGVHGLSCEDALTHLFNYVW---PNIFEVSAHMVQAFF 208
D+ DRD HR + HL LG + E TH F+ VW +FE+S +++ F
Sbjct 35 DSDQDRD-PHRLNS-----HLKLGFEDVIAEPVTTHSFDKVWICSHALFEISKYVMYKFL 88
Query 209 DAVDGMRVAIGPGVVF 224
+ +A G++F
Sbjct 89 TVFLAIPLAFIAGILF 104
> Hs20558260
Length=860
Score = 30.0 bits (66), Expect = 6.3, Method: Compositional matrix adjust.
Identities = 33/123 (26%), Positives = 56/123 (45%), Gaps = 13/123 (10%)
Query 29 PKEWDRICFDLLDLLRAQKKSIRRATVNTFGYIARTIGPQDVLATLLN------NLKMQE 82
PKE D++ L +L++ R+ + AR +GP V A LL N K E
Sbjct 200 PKERDQLLHILFNLIKRPDDEQRQMILTGCVAFARHVGPTRVEAELLPQCWEQINHKYPE 259
Query 83 RQLRL---CTTIAIAIVAE--TCLPYSVLPALINEYRVPELNIQNGVLKTLSFMFEYIGE 137
R+L + C +A + E + L S+L ++ E + ++ V+K+L + YI +
Sbjct 260 RRLLVAESCGALAPYLPKEIRSSLVLSMLQQMLMEDKAD--LVREAVIKSLGIIMGYIDD 317
Query 138 MAK 140
K
Sbjct 318 PDK 320
> Hs21361695
Length=349
Score = 29.6 bits (65), Expect = 6.6, Method: Compositional matrix adjust.
Identities = 16/50 (32%), Positives = 28/50 (56%), Gaps = 0/50 (0%)
Query 96 VAETCLPYSVLPALINEYRVPELNIQNGVLKTLSFMFEYIGEMAKDYIYT 145
+A T P +LPA+ Y+ E N +N + +S + E+IG M K+ + +
Sbjct 18 LATTLAPRVLLPAIKKTYKQIEKNWKNHMGPFMSILQEHIGAMKKEELTS 67
> At5g53480
Length=870
Score = 29.6 bits (65), Expect = 7.0, Method: Compositional matrix adjust.
Identities = 13/49 (26%), Positives = 23/49 (46%), Gaps = 0/49 (0%)
Query 19 IADRGGDLVSPKEWDRICFDLLDLLRAQKKSIRRATVNTFGYIARTIGP 67
IA G + K+W + LL + +++AT+ T GY+ + P
Sbjct 116 IAKVAGIELPQKQWPELIVSLLSNIHQLPAHVKQATLETLGYLCEEVSP 164
> At2g35630
Length=2021
Score = 29.6 bits (65), Expect = 7.5, Method: Compositional matrix adjust.
Identities = 54/262 (20%), Positives = 105/262 (40%), Gaps = 25/262 (9%)
Query 13 IDLIGRIADRGGDLVSPKEWDRICFDLLDLLRAQKKSIRRATVNTFGYIARTIGP----- 67
I+ + +I + + P + L L K++ T+ T G +A +GP
Sbjct 880 IEAVNKILEEANKRIQPTGTGELFGGLRGRLLDSNKNLVMQTLTTIGGVAAAMGPAVEKA 939
Query 68 -QDVLATLLNNLKMQERQLRLCTTIAI-----AIVAETCLPYSVLPALINEYRVPE--LN 119
+ +L+ +L L ++ +R CT A+ A+ + +PY ++ AL + E +
Sbjct 940 SKGILSDVLKCLGDNKKHMRECTLAALDLWLGAVHLDKMIPYIII-ALTDGKMGAEGRKD 998
Query 120 IQNGVLKTLSFMFEYIGEMAKDYIYTVTPLLEDALMDRDLVHRQTAAWACKHLALGVHGL 179
+ + + K L+ + +++ D I+ + P A+ D+ R+ AA C L V G
Sbjct 999 LFDWLTKQLTGLSDFV-----DAIHLLKP-ASTAMTDKSADVRK-AAEGCISEILRVSGQ 1051
Query 180 S-CEDALTHLFNYVWPNIFE-VSAHMVQAFFDAVDGMRVAIGPGV--VFRYVILGLFHPA 235
E L + + E V VQ F++ M + GV + + G
Sbjct 1052 EMIEKNLKDIQGPALALVLEKVRPGFVQEPFESSKAMAGPVSKGVTKISKSTSNGTLKQG 1111
Query 236 RKVREVYWRVYNNVYIGHQDAM 257
+ R V + + + H A+
Sbjct 1112 NRSRAVPTKGSSQITSVHDIAI 1133
Lambda K H
0.327 0.142 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 6162740004
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40