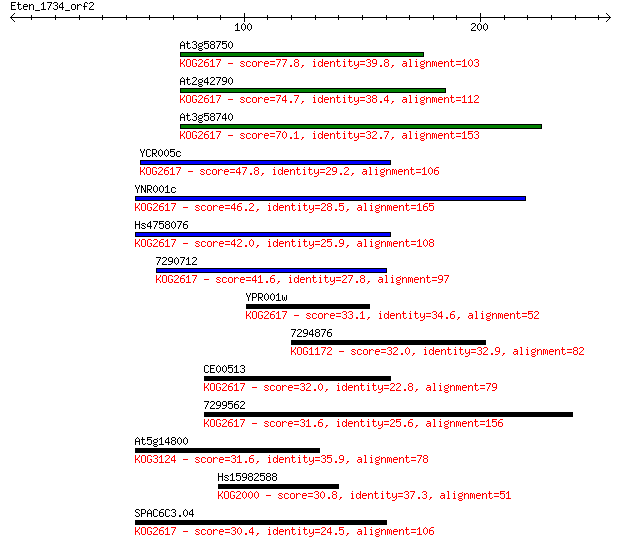
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_1734_orf2
Length=254
Score E
Sequences producing significant alignments: (Bits) Value
At3g58750 77.8 2e-14
At2g42790 74.7 2e-13
At3g58740 70.1 5e-12
YCR005c 47.8 2e-05
YNR001c 46.2 6e-05
Hs4758076 42.0 0.001
7290712 41.6 0.002
YPR001w 33.1 0.60
7294876 32.0 1.4
CE00513 32.0 1.4
7299562 31.6 1.6
At5g14800 31.6 1.6
Hs15982588 30.8 2.6
SPAC6C3.04 30.4 4.0
> At3g58750
Length=514
Score = 77.8 bits (190), Expect = 2e-14, Method: Compositional matrix adjust.
Identities = 41/104 (39%), Positives = 60/104 (57%), Gaps = 1/104 (0%)
Query 73 SSICTVALGSG-LNYRGYNVVDLAREATFEEVVHLLLLKELPTQKQLSDLKQKLYKARQL 131
SSIC + G L YRGY + +LA +TF EV +LL+ LP+Q QL+D + + + +
Sbjct 107 SSICYIDGDEGILRYRGYPIEELAESSTFIEVAYLLMYGNLPSQSQLADWEFTVSQHSAV 166
Query 132 PVQLLTVLKQIPKDAHAMDVLRTACSVLGSLEPERNPTQQQVDI 175
P +L +++ +P DAH M VL +A S L P+ NP DI
Sbjct 167 PQGVLDIIQSMPHDAHPMGVLVSAMSALSIFHPDANPALSGQDI 210
> At2g42790
Length=509
Score = 74.7 bits (182), Expect = 2e-13, Method: Compositional matrix adjust.
Identities = 43/123 (34%), Positives = 66/123 (53%), Gaps = 11/123 (8%)
Query 73 SSICTVALGSG-LNYRGYNVVDLAREATFEEVVHLLLLKELPTQKQLSDLKQKLYKARQL 131
SSI + G L YRGY + ++A +TF EV +LL+ LP++ QLSD + + + +
Sbjct 102 SSISYIDGDEGILRYRGYPIEEMAENSTFLEVAYLLMYGNLPSESQLSDWEFAVSQHSAV 161
Query 132 PVQLLTVLKQIPKDAHAMDVLRTACSVLGSLEPERNPTQQQVDI----------ALRLIG 181
P +L +++ +P DAH M VL +A S L P+ NP + DI +R+IG
Sbjct 162 PQGVLDIIQSMPHDAHPMGVLVSAMSALSIFHPDANPALRGQDIYDSKQVRDKQIIRIIG 221
Query 182 CLP 184
P
Sbjct 222 KAP 224
> At3g58740
Length=480
Score = 70.1 bits (170), Expect = 5e-12, Method: Compositional matrix adjust.
Identities = 50/166 (30%), Positives = 85/166 (51%), Gaps = 15/166 (9%)
Query 73 SSICTVALGSG-LNYRGYNVVDLAREATFEEVVHLLLLKELPTQKQLSDLKQKLYKARQL 131
SSI + G L YRGY V +LA ++T+ EV +LL+ LP+Q+QL+D + + + +
Sbjct 104 SSISYIDGDEGILRYRGYPVEELAEKSTYTEVTYLLIYGNLPSQRQLADWEFAISQNSAV 163
Query 132 PVQLLTVLKQIPKDAHAMDVLRTACSVLGSLEPERNPTQQQVDI----------ALRLIG 181
P +L +++ +P D H + L TA S L P+ NP+ + + +R++G
Sbjct 164 PQGVLDMIQSMPNDVHPVGALVTAMSALSIFYPDANPSLMGLGVYKSKQVRDKQIVRVLG 223
Query 182 CLPGALSYWFHYTNSGKEISFQSDPEDSVATNFLKLLH--GDPSFK 225
P ++ + +GK Q S + NFL ++ GD S+K
Sbjct 224 QAP-TIAAAAYLRKAGKP-PVQPLSNLSYSENFLYMVESMGDRSYK 267
> YCR005c
Length=460
Score = 47.8 bits (112), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 31/115 (26%), Positives = 50/115 (43%), Gaps = 9/115 (7%)
Query 56 VLKSTSSGGLAGVVAGESSICTVALGSGLNYRGYNVVDLAREATF---------EEVVHL 106
VL GG+ G+ + G+ +RG + D+ ++ E + L
Sbjct 56 VLLEQVYGGMRGIPGSVWEGSVLDPEDGIRFRGRTIADIQKDLPKAKGSSQPLPEALFWL 115
Query 107 LLLKELPTQKQLSDLKQKLYKARQLPVQLLTVLKQIPKDAHAMDVLRTACSVLGS 161
LL E+PTQ Q+ +L L +LP ++ +L +PKD H M A + L S
Sbjct 116 LLTGEVPTQAQVENLSADLMSRSELPSHVVQLLDNLPKDLHPMAQFSIAVTALES 170
> YNR001c
Length=479
Score = 46.2 bits (108), Expect = 6e-05, Method: Compositional matrix adjust.
Identities = 47/186 (25%), Positives = 76/186 (40%), Gaps = 24/186 (12%)
Query 54 GKVLKSTSSGGLAGV--VAGESSICTVALGSGLNYRGYNVVDLAREATFEE--------- 102
G+VL + GG+ G+ + E S+ G+ +RG + ++ RE E
Sbjct 73 GEVLLEQAYGGMRGIKGLVWEGSVLDPE--EGIRFRGRTIPEIQRELPKAEGSTEPLPEA 130
Query 103 VVHLLLLKELPTQKQLSDLKQKLYKARQLPVQLLTVLKQIPKDAHAMDVLRTACSVLGSL 162
+ LLL E+PT Q+ L L ++P ++ +L +PKD H M A + L S
Sbjct 131 LFWLLLTGEIPTDAQVKALSADLAARSEIPEHVIQLLDSLPKDLHPMAQFSIAVTALESE 190
Query 163 EPERNPTQQQV----------DIALRLIGCLPGALSYWFHYTNSGKEISFQSDPEDSVAT 212
Q V + +L L+G LP S + +I+ +DP
Sbjct 191 SKFAKAYAQGVSKKEYWSYTFEDSLDLLGKLPVIASKIYRNVFKDGKIT-STDPNADYGK 249
Query 213 NFLKLL 218
N +LL
Sbjct 250 NLAQLL 255
> Hs4758076
Length=466
Score = 42.0 bits (97), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 28/117 (23%), Positives = 53/117 (45%), Gaps = 9/117 (7%)
Query 54 GKVLKSTSSGGLAGVVAGESSICTVALGSGLNYRGYNVVDLAR---------EATFEEVV 104
G++ GG+ G+ + G+ +RG+++ + + E E +
Sbjct 61 GQITVDMMYGGMRGMKGLVYETSVLDPDEGIRFRGFSIPECQKLLPKAKGGEEPLPEGLF 120
Query 105 HLLLLKELPTQKQLSDLKQKLYKARQLPVQLLTVLKQIPKDAHAMDVLRTACSVLGS 161
LL+ +PT++Q+S L ++ K LP ++T+L P + H M L A + L S
Sbjct 121 WLLVTGCIPTEEQVSWLSKEWAKRAALPSHVVTMLDNFPTNLHPMSQLSAAVTALNS 177
> 7290712
Length=464
Score = 41.6 bits (96), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 27/106 (25%), Positives = 47/106 (44%), Gaps = 9/106 (8%)
Query 63 GGLAGVVAGESSICTVALGSGLNYRGYNV---------VDLAREATFEEVVHLLLLKELP 113
GG+ G+ A + + G+ +RG ++ D E E + LLL E+P
Sbjct 71 GGMRGIKALVTETSVLDADEGIRFRGLSIPECQKVLPAADGGTEPLPEGLFWLLLTGEVP 130
Query 114 TQKQLSDLKQKLYKARQLPVQLLTVLKQIPKDAHAMDVLRTACSVL 159
T+ Q+ L ++ + LP ++T+L +P H M A + L
Sbjct 131 TKSQVQQLSREWAERAALPQHVVTMLNNMPTTLHPMSQFAAAVTAL 176
> YPR001w
Length=486
Score = 33.1 bits (74), Expect = 0.60, Method: Compositional matrix adjust.
Identities = 18/53 (33%), Positives = 30/53 (56%), Gaps = 1/53 (1%)
Query 101 EEVVHLLLLKELPTQKQLSDLKQKL-YKARQLPVQLLTVLKQIPKDAHAMDVL 152
E ++ LL+ +PT +Q + +++L + R+LP VL +PKD H M L
Sbjct 114 ESMLWLLMTGGVPTFQQAASFRKELAIRGRKLPHYTEKVLSSLPKDMHPMTQL 166
> 7294876
Length=1239
Score = 32.0 bits (71), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 27/93 (29%), Positives = 41/93 (44%), Gaps = 11/93 (11%)
Query 120 DLKQKLYKARQLPVQLL---TVLKQIPKDAHAMDVLRTACSVLGSL--------EPERNP 168
D+K +Y + Q ++ L T+LK+IP A A VL A L E P
Sbjct 454 DMKDDMYSSSQEDLKKLQNDTILKRIPAGAEATTVLVGAVEFLEQPTIAFVRLSEGVLMP 513
Query 169 TQQQVDIALRLIGCLPGALSYWFHYTNSGKEIS 201
T +V + +R + L G ++ Y G+ IS
Sbjct 514 TLTEVPVPVRFMFVLLGPRNFDLDYHEVGRSIS 546
> CE00513
Length=468
Score = 32.0 bits (71), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 18/88 (20%), Positives = 39/88 (44%), Gaps = 9/88 (10%)
Query 83 GLNYRGYNVVDLAR---------EATFEEVVHLLLLKELPTQKQLSDLKQKLYKARQLPV 133
G+ +RGY++ + + E E + LL ++P++ Q + + ++ LP
Sbjct 92 GIRFRGYSIPECQKLLPKAKGGEEPLPEAIWWLLCTGDVPSEAQTAAITKEWNARADLPT 151
Query 134 QLLTVLKQIPKDAHAMDVLRTACSVLGS 161
++ +L P + H M A + L +
Sbjct 152 HVVRMLDNFPDNLHPMAQFIAAIAALNN 179
> 7299562
Length=407
Score = 31.6 bits (70), Expect = 1.6, Method: Compositional matrix adjust.
Identities = 40/169 (23%), Positives = 67/169 (39%), Gaps = 28/169 (16%)
Query 83 GLNYRGYNVVDLA----------REATFEEVVHLLLLKELPTQKQLSDLKQKLYKARQLP 132
G+ YRG + D+ +E T E LL +PT+K+ ++ + K +P
Sbjct 19 GIYYRGKLLKDVCAKLPRVQEGTQEGTPEGCFFLLTSGSMPTKKEAQEVTNEWLKRGSVP 78
Query 133 VQLLTVLKQIPKDAHAMDVLRTACSVLGSLEPERNPTQQQVDIALRLIGCLPGA--LSYW 190
L ++ + K H M L A + L NP Q V+ + GA YW
Sbjct 79 RYCLRMIDSMDKRVHPMAQLCAASACL-------NPQSQFVEAYTK------GARRADYW 125
Query 191 -FHYTNSGKEISFQSDPEDSVATNFLKLLHGDPSFKVRRGRAKSASPCR 238
+ Y +S I+ ++ +N + G+ S +V S + CR
Sbjct 126 KYSYEDSMNLIAMLPTVAAAIYSNVFR--DGEGSREVNYEEDWSGNFCR 172
> At5g14800
Length=276
Score = 31.6 bits (70), Expect = 1.6, Method: Compositional matrix adjust.
Identities = 28/84 (33%), Positives = 42/84 (50%), Gaps = 7/84 (8%)
Query 54 GKVLKSTSSGGLAGVVAGESSICTVALGSGLNYR------GYNVVDLAREATFEEVVHLL 107
GK+ +S + G +A V + ICT A+ S LN R G NV + E E V +
Sbjct 19 GKMAESIARGVVASGVLPPNRICT-AVHSNLNRRDVFESFGVNVFSTSEEVVKESDVVIF 77
Query 108 LLKELPTQKQLSDLKQKLYKARQL 131
+K +K +++LK KL K + L
Sbjct 78 SVKPQVVKKAVTELKSKLSKNKIL 101
> Hs15982588
Length=1819
Score = 30.8 bits (68), Expect = 2.6, Method: Compositional matrix adjust.
Identities = 19/51 (37%), Positives = 28/51 (54%), Gaps = 0/51 (0%)
Query 89 YNVVDLAREATFEEVVHLLLLKELPTQKQLSDLKQKLYKARQLPVQLLTVL 139
Y V+L EA +E + H LL+++ + LSDL + A Q P +LL L
Sbjct 1506 YFFVELHLEAHYEALRHFLLMEDGEFAQSLSDLLFEKLGAGQTPGELLNPL 1556
> SPAC6C3.04
Length=473
Score = 30.4 bits (67), Expect = 4.0, Method: Compositional matrix adjust.
Identities = 26/115 (22%), Positives = 45/115 (39%), Gaps = 9/115 (7%)
Query 54 GKVLKSTSSGGLAGVVAGESSICTVALGSGLNYRGYNVVDL---------AREATFEEVV 104
G+V + GG GV + + G+ +RGY + + ++ E +
Sbjct 70 GEVTINQMYGGARGVRSLIWEGSVLDPNEGIRFRGYTIPECQKLLPSSPNGKQPLPESLF 129
Query 105 HLLLLKELPTQKQLSDLKQKLYKARQLPVQLLTVLKQIPKDAHAMDVLRTACSVL 159
LL+ E+PT Q+ L QLP + ++ + P H M A + L
Sbjct 130 WLLVTGEIPTLSQVQALSADWAARSQLPKFVEELIDRCPPTLHPMAQFSLAVTAL 184
Lambda K H
0.317 0.131 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 5258582968
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40