bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_1378_orf3
Length=259
Score E
Sequences producing significant alignments: (Bits) Value
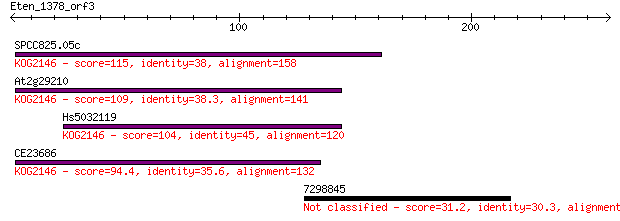
SPCC825.05c 115 1e-25
At2g29210 109 6e-24
Hs5032119 104 2e-22
CE23686 94.4 2e-19
7298845 31.2 2.4
> SPCC825.05c
Length=301
Score = 115 bits (287), Expect = 1e-25, Method: Compositional matrix adjust.
Identities = 60/168 (35%), Positives = 98/168 (58%), Gaps = 12/168 (7%)
Query 3 EQTPFFKDKDSQLIAQRKWPAIFDQEVDMSKVNVEVMKAWINHKITELLGFEDDIVISYC 62
EQ F D +L+ K+PA +D +VDM KVN+EV+K WI ++ EL+GFED++VI++
Sbjct 11 EQETLFTTADKKLMRSTKFPASYDTKVDMKKVNIEVLKPWIATRLNELIGFEDEVVINFV 70
Query 63 LSQLVPEEALAADV---DERKNYLCPKKLAISLTGFVGKQATHFVRELWKLLLSAQNTPT 119
L EEA+ A + ++ L P+K+ ++LTGF+ AT F ELW L++SA
Sbjct 71 YGML--EEAVEASKTSDSQNESTLDPRKVQLNLTGFLESNATAFTEELWSLIISASQNQY 128
Query 120 GIPPEFLENRRLEM-------ERKREEAARINEELVRRHEQMMATEFN 160
GIP +F+ ++ E+ E +EE+ + + RR + +T ++
Sbjct 129 GIPEKFILEKKEEISKLKDRTEASKEESKTVTDHSNRRESRRESTYYD 176
> At2g29210
Length=891
Score = 109 bits (273), Expect = 6e-24, Method: Compositional matrix adjust.
Identities = 54/141 (38%), Positives = 86/141 (60%), Gaps = 12/141 (8%)
Query 3 EQTPFFKDKDSQLIAQRKWPAIFDQEVDMSKVNVEVMKAWINHKITELLGFEDDIVISYC 62
EQ F +K ++L+ +K+ + VD++KV ++VMK WI ++TELLG ED+++I++
Sbjct 32 EQDTRFSNKQAKLMKSQKFAPELENLVDITKVKMDVMKPWIATRVTELLGIEDEVLINFI 91
Query 63 LSQLVPEEALAADVDERKNYLCPKKLAISLTGFVGKQATHFVRELWKLLLSAQNTPTGIP 122
L + K++ I+LTGF+ K F++ELW LLLSAQN P+G+P
Sbjct 92 YGLL------------DGKVVNGKEIQITLTGFMEKNTGKFMKELWTLLLSAQNNPSGVP 139
Query 123 PEFLENRRLEMERKREEAARI 143
+FL+ R E ++K+EEA I
Sbjct 140 QQFLDARAAETKKKQEEANEI 160
> Hs5032119
Length=820
Score = 104 bits (260), Expect = 2e-22, Method: Compositional matrix adjust.
Identities = 54/121 (44%), Positives = 78/121 (64%), Gaps = 13/121 (10%)
Query 24 IFDQEVDMSKVNVEVMKAWINHKITELLGFEDDIVISYCLSQLVPEEALAADVDERKNYL 83
+++VDMSKVN+EV+K WI ++TE+LGFEDD+VI + +QL E KN
Sbjct 33 CLEKKVDMSKVNLEVIKPWITKRVTEILGFEDDVVIEFIFNQL-----------EVKNPD 81
Query 84 CPKKLAISLTGFV-GKQATHFVRELWKLLLSAQNTPTGIPPEFLENRRLEMERKREEAAR 142
K + I+LTGF+ GK A F+ ELW LLLSAQ GIP FLE ++ E+++++ E +
Sbjct 82 S-KMMQINLTGFLNGKNAREFMGELWPLLLSAQENIAGIPSAFLELKKEEIKQRQIEQEK 140
Query 143 I 143
+
Sbjct 141 L 141
> CE23686
Length=335
Score = 94.4 bits (233), Expect = 2e-19, Method: Compositional matrix adjust.
Identities = 47/133 (35%), Positives = 79/133 (59%), Gaps = 13/133 (9%)
Query 3 EQTPFFKDKDSQLIAQRKWPAIFDQEVDMSKVNVEVMKAWINHKITELLGFEDDIVISYC 62
EQ F DK+ +L+ K+ +Q++D+++ N++V+K WI ++ ++LG EDD+V+ Y
Sbjct 12 EQDGRFSDKEKKLLKTMKFEPQLEQKIDLNRCNMDVIKPWITARVNDILGMEDDVVVEYI 71
Query 63 LSQLVPEEALAADVDERKNYLCPKKLAISLTGFV-GKQATHFVRELWKLLLSAQNTPTGI 121
LSQ+ + KN L PK L I++TGF+ ++A FV +LW LL+ A + GI
Sbjct 72 LSQI-----------DDKN-LNPKLLQINVTGFLNARRAREFVGDLWNLLIEANASEDGI 119
Query 122 PPEFLENRRLEME 134
P + + EM+
Sbjct 120 PASLVNQKMAEMK 132
> 7298845
Length=891
Score = 31.2 bits (69), Expect = 2.4, Method: Compositional matrix adjust.
Identities = 27/92 (29%), Positives = 40/92 (43%), Gaps = 7/92 (7%)
Query 128 NRRLEMERKREEAARINEELV---RRHEQMMATEFNKQIKQFATGTAPPKPPTLVAYDTI 184
N R E+E R+E R+ +ELV R + M E N+ I + AP P+ D +
Sbjct 560 NYRDEIELLRKENKRLEQELVKIGRENNSKMLQEINRNIARL----APKVLPSATMSDIL 615
Query 185 PDTGGGMRGWDNDDAPAARGRAASHERSRSRS 216
+ G + D A+ R A + R S S
Sbjct 616 DENGRRTQSLDGTAGKPAKQREAQNRRQSSGS 647
Lambda K H
0.316 0.131 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 5436237798
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40