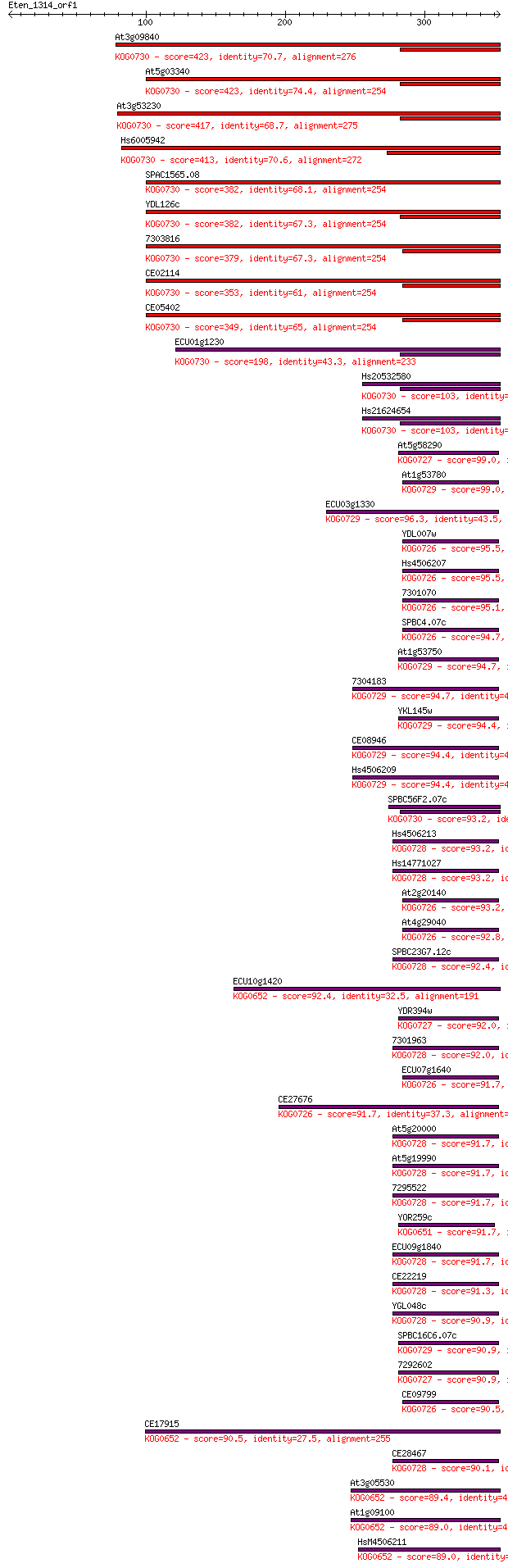
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_1314_orf1
Length=353
Score E
Sequences producing significant alignments: (Bits) Value
At3g09840 423 3e-118
At5g03340 423 4e-118
At3g53230 417 1e-116
Hs6005942 413 4e-115
SPAC1565.08 382 7e-106
YDL126c 382 8e-106
7303816 379 7e-105
CE02114 353 3e-97
CE05402 349 4e-96
ECU01g1230 198 2e-50
Hs20532580 103 6e-22
Hs21624654 103 7e-22
At5g58290 99.0 1e-20
At1g53780 99.0 1e-20
ECU03g1330 96.3 8e-20
YDL007w 95.5 2e-19
Hs4506207 95.5 2e-19
7301070 95.1 2e-19
SPBC4.07c 94.7 2e-19
At1g53750 94.7 3e-19
7304183 94.7 3e-19
YKL145w 94.4 3e-19
CE08946 94.4 4e-19
Hs4506209 94.4 4e-19
SPBC56F2.07c 93.2 7e-19
Hs4506213 93.2 8e-19
Hs14771027 93.2 8e-19
At2g20140 93.2 8e-19
At4g29040 92.8 9e-19
SPBC23G7.12c 92.4 1e-18
ECU10g1420 92.4 1e-18
YDR394w 92.0 2e-18
7301963 92.0 2e-18
ECU07g1640 91.7 2e-18
CE27676 91.7 2e-18
At5g20000 91.7 2e-18
At5g19990 91.7 2e-18
7295522 91.7 2e-18
YOR259c 91.7 2e-18
ECU09g1840 91.7 2e-18
CE22219 91.3 3e-18
YGL048c 90.9 3e-18
SPBC16C6.07c 90.9 4e-18
7292602 90.9 4e-18
CE09799 90.5 5e-18
CE17915 90.5 5e-18
CE28467 90.1 6e-18
At3g05530 89.4 1e-17
At1g09100 89.0 1e-17
HsM4506211 89.0 2e-17
> At3g09840
Length=809
Score = 423 bits (1087), Expect = 3e-118, Method: Compositional matrix adjust.
Identities = 195/277 (70%), Positives = 237/277 (85%), Gaps = 4/277 (1%)
Query 78 GAPAASAGNK-KADANAAANQQPKKKRSPNRLVVEEAINDDNSVVALNPSRMETLQIFRG 136
PA S+ +K K D + A + +K+SPNRLVV+EAINDDNSVV+L+P+ ME LQ+FRG
Sbjct 2 STPAESSDSKSKKDFSTAILE---RKKSPNRLVVDEAINDDNSVVSLHPATMEKLQLFRG 58
Query 137 DTVLLRGKMRHDTVCVVLADQDLDEGKIRLNKVVRKNLRVRLGDIVHVSACADCPYGKRI 196
DT+L++GK R DTVC+ LAD+ +E KIR+NKVVR NLRVRLGD++ V C D YGKR+
Sbjct 59 DTILIKGKKRKDTVCIALADETCEEPKIRMNKVVRSNLRVRLGDVISVHQCPDVKYGKRV 118
Query 197 HVLPLDDTIEGITGNLFEIYLKPYFMEAYRPVRKNDLFLVRGGFRPVEFKVVGVDPGEFC 256
H+LP+DDT+EG+TGNLF+ YLKPYF+EAYRPVRK DLFLVRGG R VEFKV+ DP E+C
Sbjct 119 HILPVDDTVEGVTGNLFDAYLKPYFLEAYRPVRKGDLFLVRGGMRSVEFKVIETDPAEYC 178
Query 257 IVAPDTVIHCEGEPVKREEEERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGV 316
+VAPDT I CEGEPVKRE+EERLD++GY+D+GG RKQMAQIRE++ELPLRHP LFK++GV
Sbjct 179 VVAPDTEIFCEGEPVKREDEERLDDVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGV 238
Query 317 KPPRGVLLYGPPGSGKTLIAKAVANETGAFFFLINGP 353
KPP+G+LLYGPPGSGKTLIA+AVANETGAFFF INGP
Sbjct 239 KPPKGILLYGPPGSGKTLIARAVANETGAFFFCINGP 275
Score = 80.1 bits (196), Expect = 6e-15, Method: Compositional matrix adjust.
Identities = 33/72 (45%), Positives = 47/72 (65%), Gaps = 0/72 (0%)
Query 282 IGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVAN 341
+ + DIGG +++E ++ P+ HP F+ G+ P +GVL YGPPG GKTL+AKA+AN
Sbjct 477 VSWNDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGKTLLAKAIAN 536
Query 342 ETGAFFFLINGP 353
E A F + GP
Sbjct 537 ECQANFISVKGP 548
> At5g03340
Length=843
Score = 423 bits (1087), Expect = 4e-118, Method: Compositional matrix adjust.
Identities = 189/254 (74%), Positives = 227/254 (89%), Gaps = 0/254 (0%)
Query 100 KKKRSPNRLVVEEAINDDNSVVALNPSRMETLQIFRGDTVLLRGKMRHDTVCVVLADQDL 159
++K+SPNRLVV+EAINDDNSVV+L+P+ ME LQ+FRGDT+L++GK R DTVC+ LAD+
Sbjct 55 ERKKSPNRLVVDEAINDDNSVVSLHPTTMEKLQLFRGDTILIKGKKRKDTVCIALADETC 114
Query 160 DEGKIRLNKVVRKNLRVRLGDIVHVSACADCPYGKRIHVLPLDDTIEGITGNLFEIYLKP 219
+E KIR+NKVVR NLRVRLGD++ V C D YGKR+H+LP+DDT+EG+TGNLF+ YLKP
Sbjct 115 EEPKIRMNKVVRSNLRVRLGDVISVHQCPDVKYGKRVHILPVDDTVEGVTGNLFDAYLKP 174
Query 220 YFMEAYRPVRKNDLFLVRGGFRPVEFKVVGVDPGEFCIVAPDTVIHCEGEPVKREEEERL 279
YF+EAYRPVRK DLFLVRGG R VEFKV+ DP E+C+VAPDT I CEGEPVKRE+EERL
Sbjct 175 YFLEAYRPVRKGDLFLVRGGMRSVEFKVIETDPAEYCVVAPDTEIFCEGEPVKREDEERL 234
Query 280 DEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAV 339
DE+GY+D+GG RKQMAQIRE++ELPLRHP LFK++GVKPP+G+LLYGPPGSGKTLIA+AV
Sbjct 235 DEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPPGSGKTLIARAV 294
Query 340 ANETGAFFFLINGP 353
ANETGAFFF INGP
Sbjct 295 ANETGAFFFCINGP 308
Score = 82.0 bits (201), Expect = 2e-15, Method: Compositional matrix adjust.
Identities = 34/72 (47%), Positives = 48/72 (66%), Gaps = 0/72 (0%)
Query 282 IGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVAN 341
+ +EDIGG +++E ++ P+ HP F+ G+ P +GVL YGPPG GKTL+AKA+AN
Sbjct 510 VSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGKTLLAKAIAN 569
Query 342 ETGAFFFLINGP 353
E A F + GP
Sbjct 570 ECQANFISVKGP 581
> At3g53230
Length=815
Score = 417 bits (1073), Expect = 1e-116, Method: Compositional matrix adjust.
Identities = 189/275 (68%), Positives = 230/275 (83%), Gaps = 0/275 (0%)
Query 79 APAASAGNKKADANAAANQQPKKKRSPNRLVVEEAINDDNSVVALNPSRMETLQIFRGDT 138
A A + + K + +KK++ NRLVV+EAINDDNSVV+L+P ME LQ+FRGDT
Sbjct 2 ANQAESSDSKGTKKDFSTAILEKKKAANRLVVDEAINDDNSVVSLHPDTMEKLQLFRGDT 61
Query 139 VLLRGKMRHDTVCVVLADQDLDEGKIRLNKVVRKNLRVRLGDIVHVSACADCPYGKRIHV 198
+L++GK R DTVC+ LAD+ DE KIR+NKVVR NLRVRLGD++ V C D YG R+H+
Sbjct 62 ILIKGKKRKDTVCIALADETCDEPKIRMNKVVRSNLRVRLGDVISVHQCPDVKYGNRVHI 121
Query 199 LPLDDTIEGITGNLFEIYLKPYFMEAYRPVRKNDLFLVRGGFRPVEFKVVGVDPGEFCIV 258
LPLDDTIEG++GN+F+ YLKPYF+EAYRPVRK DLFLVRGG R +EFKV+ DP E+C+V
Sbjct 122 LPLDDTIEGVSGNIFDAYLKPYFLEAYRPVRKGDLFLVRGGMRSIEFKVIETDPAEYCVV 181
Query 259 APDTVIHCEGEPVKREEEERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKP 318
APDT I CEGEP+KRE+EERLDE+GY+D+GG RKQMAQIRE++ELPLRHP LFK++GVKP
Sbjct 182 APDTEIFCEGEPIKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKP 241
Query 319 PRGVLLYGPPGSGKTLIAKAVANETGAFFFLINGP 353
P+G+LLYGPPGSGKTLIA+AVANETGAFFF INGP
Sbjct 242 PKGILLYGPPGSGKTLIARAVANETGAFFFCINGP 276
Score = 82.4 bits (202), Expect = 1e-15, Method: Compositional matrix adjust.
Identities = 35/72 (48%), Positives = 48/72 (66%), Gaps = 0/72 (0%)
Query 282 IGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVAN 341
+ +EDIGG +++E ++ P+ HP F+ G+ P +GVL YGPPG GKTL+AKA+AN
Sbjct 478 VSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGKTLLAKAIAN 537
Query 342 ETGAFFFLINGP 353
E A F I GP
Sbjct 538 ECQANFISIKGP 549
> Hs6005942
Length=806
Score = 413 bits (1061), Expect = 4e-115, Method: Compositional matrix adjust.
Identities = 192/273 (70%), Positives = 231/273 (84%), Gaps = 3/273 (1%)
Query 82 ASAGNKKADANAAANQQPKKKRSPNRLVVEEAINDDNSVVALNPSRMETLQIFRGDTVLL 141
AS + K D + A K+K PNRL+V+EAIN+DNSVV+L+ +M+ LQ+FRGDTVLL
Sbjct 2 ASGADSKGDDLSTAIL--KQKNRPNRLIVDEAINEDNSVVSLSQPKMDELQLFRGDTVLL 59
Query 142 RGKMRHDTVCVVLADQDLDEGKIRLNKVVRKNLRVRLGDIVHVSACADCPYGKRIHVLPL 201
+GK R + VC+VL+D + KIR+N+VVR NLRVRLGD++ + C D YGKRIHVLP+
Sbjct 60 KGKKRREAVCIVLSDDTCSDEKIRMNRVVRNNLRVRLGDVISIQPCPDVKYGKRIHVLPI 119
Query 202 DDTIEGITGNLFEIYLKPYFMEAYRPVRKNDLFLVRGGFRPVEFKVVGVDPGEFCIVAPD 261
DDT+EGITGNLFE+YLKPYF+EAYRP+RK D+FLVRGG R VEFKVV DP +CIVAPD
Sbjct 120 DDTVEGITGNLFEVYLKPYFLEAYRPIRKGDIFLVRGGMRAVEFKVVETDPSPYCIVAPD 179
Query 262 TVIHCEGEPVKRE-EEERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPR 320
TVIHCEGEP+KRE EEE L+E+GY+DIGGCRKQ+AQI+EM+ELPLRHP LFK +GVKPPR
Sbjct 180 TVIHCEGEPIKREDEEESLNEVGYDDIGGCRKQLAQIKEMVELPLRHPALFKAIGVKPPR 239
Query 321 GVLLYGPPGSGKTLIAKAVANETGAFFFLINGP 353
G+LLYGPPG+GKTLIA+AVANETGAFFFLINGP
Sbjct 240 GILLYGPPGTGKTLIARAVANETGAFFFLINGP 272
Score = 82.0 bits (201), Expect = 2e-15, Method: Compositional matrix adjust.
Identities = 37/81 (45%), Positives = 52/81 (64%), Gaps = 0/81 (0%)
Query 273 REEEERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGK 332
RE + ++ +EDIGG +++E+++ P+ HP F G+ P +GVL YGPPG GK
Sbjct 465 RETVVEVPQVTWEDIGGLEDVKRELQELVQYPVEHPDKFLKFGMTPSKGVLFYGPPGCGK 524
Query 333 TLIAKAVANETGAFFFLINGP 353
TL+AKA+ANE A F I GP
Sbjct 525 TLLAKAIANECQANFISIKGP 545
> SPAC1565.08
Length=418
Score = 382 bits (980), Expect = 7e-106, Method: Compositional matrix adjust.
Identities = 173/255 (67%), Positives = 219/255 (85%), Gaps = 1/255 (0%)
Query 100 KKKRSPNRLVVEEAINDDNSVVALNPSRMETLQIFRGDTVLLRGKMRHDTVCVVLADQDL 159
+KKR PN LVV++A NDDNSV+ L+ + METLQ+FRGDTV+++GK R DTV +VL D+++
Sbjct 38 RKKRKPNSLVVDDATNDDNSVITLSSNTMETLQLFRGDTVVVKGKRRKDTVLIVLTDEEM 97
Query 160 DEGKIRLNKVVRKNLRVRLGDIVHVSACADCPYGKRIHVLPLDDTIEGITGNLFEIYLKP 219
++G R+N+VVR NLRVRLGDIV ++ C D Y +RI VLPL DT+EG+TG+LF++YLKP
Sbjct 98 EDGVARINRVVRNNLRVRLGDIVTINPCPDIKYAERISVLPLADTVEGLTGSLFDVYLKP 157
Query 220 YFMEAYRPVRKNDLFLVRGGFRPVEFKVVGVDPGEFCIVAPDTVIHCEGEPVKREEEE-R 278
YF+EAYRP+RK DLF+VRG R VEFKVV V P EF IV+ DT+IH EGEP+ RE+EE
Sbjct 158 YFVEAYRPIRKGDLFVVRGSMRQVEFKVVDVAPDEFGIVSQDTIIHWEGEPINREDEESS 217
Query 279 LDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKA 338
L E+GY+DIGGCR+QMAQIRE++ELPLRHP LFK++G+KPPRG+L+YGPPG+GKTL+A+A
Sbjct 218 LAEVGYDDIGGCRRQMAQIRELVELPLRHPQLFKSIGIKPPRGILMYGPPGTGKTLMARA 277
Query 339 VANETGAFFFLINGP 353
VANETGAFFFLINGP
Sbjct 278 VANETGAFFFLINGP 292
> YDL126c
Length=835
Score = 382 bits (980), Expect = 8e-106, Method: Compositional matrix adjust.
Identities = 171/255 (67%), Positives = 217/255 (85%), Gaps = 1/255 (0%)
Query 100 KKKRSPNRLVVEEAINDDNSVVALNPSRMETLQIFRGDTVLLRGKMRHDTVCVVLADQDL 159
++K+ N L+V++AINDDNSV+A+N + M+ L++FRGDTVL++GK R DTV +VL D +L
Sbjct 28 RRKKKDNMLLVDDAINDDNSVIAINSNTMDKLELFRGDTVLVKGKKRKDTVLIVLIDDEL 87
Query 160 DEGKIRLNKVVRKNLRVRLGDIVHVSACADCPYGKRIHVLPLDDTIEGITGNLFEIYLKP 219
++G R+N+VVR NLR+RLGD+V + C D Y RI VLP+ DTIEGITGNLF+++LKP
Sbjct 88 EDGACRINRVVRNNLRIRLGDLVTIHPCPDIKYATRISVLPIADTIEGITGNLFDVFLKP 147
Query 220 YFMEAYRPVRKNDLFLVRGGFRPVEFKVVGVDPGEFCIVAPDTVIHCEGEPVKREEEE-R 278
YF+EAYRPVRK D F+VRGG R VEFKVV V+P E+ +VA DT+IH EGEP+ RE+EE
Sbjct 148 YFVEAYRPVRKGDHFVVRGGMRQVEFKVVDVEPEEYAVVAQDTIIHWEGEPINREDEENN 207
Query 279 LDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKA 338
++E+GY+DIGGCRKQMAQIREM+ELPLRHP LFK +G+KPPRGVL+YGPPG+GKTL+A+A
Sbjct 208 MNEVGYDDIGGCRKQMAQIREMVELPLRHPQLFKAIGIKPPRGVLMYGPPGTGKTLMARA 267
Query 339 VANETGAFFFLINGP 353
VANETGAFFFLINGP
Sbjct 268 VANETGAFFFLINGP 282
Score = 79.7 bits (195), Expect = 1e-14, Method: Compositional matrix adjust.
Identities = 32/72 (44%), Positives = 48/72 (66%), Gaps = 0/72 (0%)
Query 282 IGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVAN 341
+ ++D+GG + +++E +E P+ HP + G+ P +GVL YGPPG+GKTL+AKAVA
Sbjct 484 VTWDDVGGLDEIKEELKETVEYPVLHPDQYTKFGLSPSKGVLFYGPPGTGKTLLAKAVAT 543
Query 342 ETGAFFFLINGP 353
E A F + GP
Sbjct 544 EVSANFISVKGP 555
> 7303816
Length=801
Score = 379 bits (972), Expect = 7e-105, Method: Compositional matrix adjust.
Identities = 171/255 (67%), Positives = 216/255 (84%), Gaps = 1/255 (0%)
Query 100 KKKRSPNRLVVEEAINDDNSVVALNPSRMETLQIFRGDTVLLRGKMRHDTVCVVLADQDL 159
K+K PNRL+VEEA NDDNSVV+L+ ++M+ LQ+FRGDTV+L+GK R +TVC+VL+D
Sbjct 15 KRKDRPNRLIVEEAQNDDNSVVSLSQAKMDELQLFRGDTVILKGKRRKETVCIVLSDDTC 74
Query 160 DEGKIRLNKVVRKNLRVRLGDIVHVSACADCPYGKRIHVLPLDDTIEGITGNLFEIYLKP 219
+ KIR+N+VVR NL V L D+V V +C D YGKR+ +LP+D++ EG+TGNLFEIYLKP
Sbjct 75 PDEKIRMNRVVRNNLCVHLSDVVSVQSCPDVKYGKRVRILPIDESTEGVTGNLFEIYLKP 134
Query 220 YFMEAYRPVRKNDLFLVRGGFRPVEFKVVGVDPGEFCIVAPDTVIHCEGEPVKREEEER- 278
YF+EAYRP+ D F+VR RP+EFKVV DP +CIVAP+TVI C+G+P+KREEEE
Sbjct 135 YFLEAYRPIHMGDNFIVRAAMRPIEFKVVLTDPEPYCIVAPETVIFCDGDPIKREEEEES 194
Query 279 LDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKA 338
L+ +GY+DIGGCRKQ+AQI+EM+ELPLRHP+LFK +GVKPPRG+L+YGPPG+GKTLIA+A
Sbjct 195 LNAVGYDDIGGCRKQLAQIKEMVELPLRHPSLFKAIGVKPPRGILMYGPPGTGKTLIARA 254
Query 339 VANETGAFFFLINGP 353
VANETGAFFFLINGP
Sbjct 255 VANETGAFFFLINGP 269
Score = 80.5 bits (197), Expect = 5e-15, Method: Compositional matrix adjust.
Identities = 34/70 (48%), Positives = 47/70 (67%), Gaps = 0/70 (0%)
Query 284 YEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANET 343
+ DIGG +++E+++ P+ HP F G++P RGVL YGPPG GKTL+AKA+ANE
Sbjct 473 WTDIGGLESVKKELQELVQYPVEHPDKFLKFGMQPSRGVLFYGPPGCGKTLLAKAIANEC 532
Query 344 GAFFFLINGP 353
A F + GP
Sbjct 533 QANFISVKGP 542
> CE02114
Length=809
Score = 353 bits (906), Expect = 3e-97, Method: Compositional matrix adjust.
Identities = 155/255 (60%), Positives = 212/255 (83%), Gaps = 1/255 (0%)
Query 100 KKKRSPNRLVVEEAINDDNSVVALNPSRMETLQIFRGDTVLLRGKMRHDTVCVVLADQDL 159
K K PNRL+V+++ DDNSV+A++ ++M+ L +FRGD V+L+GK R ++V ++++D+
Sbjct 24 KDKVKPNRLIVDQSEQDDNSVIAVSQAKMDELGLFRGDAVILKGKKRKESVAIIVSDESC 83
Query 160 DEGKIRLNKVVRKNLRVRLGDIVHVSACADCPYGKRIHVLPLDDTIEGITGNLFEIYLKP 219
K+R+N+VVR NLR+RLGD+V ++ + YG RIHVLP+DDTIEG+TGNLF+++LKP
Sbjct 84 PNEKVRMNRVVRNNLRIRLGDVVSITPAPNLSYGTRIHVLPIDDTIEGLTGNLFDVFLKP 143
Query 220 YFMEAYRPVRKNDLFLVRGGFRPVEFKVVGVDPGEFCIVAPDTVIHCEGEPVKREEEER- 278
YF+EAYRP+ K D+F V+ R VEFKVV +P CIV+PDT+IH EG+P+KREEEE
Sbjct 144 YFLEAYRPLHKGDIFTVQAAMRTVEFKVVETEPAPACIVSPDTMIHYEGDPIKREEEEES 203
Query 279 LDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKA 338
+++IGY+D+GG RKQ+AQI+EM+ELPLRHP LFK +G+KPPRG+LL+GPPG+GKTLIA+A
Sbjct 204 MNDIGYDDLGGVRKQLAQIKEMVELPLRHPQLFKAIGIKPPRGILLFGPPGTGKTLIARA 263
Query 339 VANETGAFFFLINGP 353
VANETG+FFFLINGP
Sbjct 264 VANETGSFFFLINGP 278
Score = 80.5 bits (197), Expect = 6e-15, Method: Compositional matrix adjust.
Identities = 34/70 (48%), Positives = 48/70 (68%), Gaps = 0/70 (0%)
Query 284 YEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANET 343
+ DIGG + +++E+++ P+ HP + G++P RGVL YGPPG GKTL+AKA+ANE
Sbjct 482 WSDIGGLQNVKRELQELVQYPVEHPEKYLKFGMQPSRGVLFYGPPGCGKTLLAKAIANEC 541
Query 344 GAFFFLINGP 353
A F I GP
Sbjct 542 QANFISIKGP 551
> CE05402
Length=810
Score = 349 bits (896), Expect = 4e-96, Method: Compositional matrix adjust.
Identities = 165/255 (64%), Positives = 212/255 (83%), Gaps = 2/255 (0%)
Query 100 KKKRSPNRLVVEEAINDDNSVVALNPSRMETLQIFRGDTVLLRGKMRHDTVCVVLADQDL 159
K K+ PNRL+++++ NDDNS+V L+ ++M+ L +FRGD+V+L+GK R +TV +VL +
Sbjct 24 KDKKRPNRLIIDQSDNDDNSMVMLSQAKMDELGLFRGDSVILKGKKRRETVSIVLNADNC 83
Query 160 DEGKIRLNKVVRKNLRVRLGDIVHVSACADCPYGKRIHVLPLDDTIEGITGNLFEIYLKP 219
KI++NKVVR NLR RLGD+V +S+ A YGKR+HVLP+DDTIEG+TGNLF+++L+P
Sbjct 84 PNDKIKMNKVVRNNLRSRLGDVVSISS-AQLEYGKRVHVLPIDDTIEGLTGNLFDVFLRP 142
Query 220 YFMEAYRPVRKNDLFLVRGGFRPVEFKVVGVDPGEFCIVAPDTVIHCEGEPVKREEEER- 278
YF +AYRPV K D+F V+ R VEFKVV DP CIVAPDTVIH EG+P+KREEEE
Sbjct 143 YFTDAYRPVHKGDIFTVQAAMRTVEFKVVETDPAPACIVAPDTVIHYEGDPIKREEEEEA 202
Query 279 LDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKA 338
L+E+GY+D+GG RKQ+AQI+EM+ELPLRHP LFK +GVKPPRG+LL+GPPG+GKTLIA+A
Sbjct 203 LNEVGYDDLGGVRKQLAQIKEMVELPLRHPQLFKAIGVKPPRGILLFGPPGTGKTLIARA 262
Query 339 VANETGAFFFLINGP 353
VANETGAFFFLINGP
Sbjct 263 VANETGAFFFLINGP 277
Score = 80.1 bits (196), Expect = 7e-15, Method: Compositional matrix adjust.
Identities = 34/70 (48%), Positives = 48/70 (68%), Gaps = 0/70 (0%)
Query 284 YEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANET 343
+ DIGG + +++E+++ P+ HP + G++P RGVL YGPPG GKTL+AKA+ANE
Sbjct 481 WSDIGGLQNVKRELQELVQYPVEHPEKYLKFGMQPSRGVLFYGPPGCGKTLLAKAIANEC 540
Query 344 GAFFFLINGP 353
A F I GP
Sbjct 541 QANFISIKGP 550
> ECU01g1230
Length=780
Score = 198 bits (503), Expect = 2e-50, Method: Compositional matrix adjust.
Identities = 101/240 (42%), Positives = 154/240 (64%), Gaps = 8/240 (3%)
Query 121 VALNPSRMETLQIFRGDTVLLRGKMRHDTVCVVLADQDLDEGKIRLNKVVRKNLRVRLGD 180
V L+P+ + L++F D V + GK + + + +A + + I + + R NLR+R+ D
Sbjct 38 VGLHPTTLNELELFESDYVRILGKKKAELIFSTVALESVPPRHIAIVRDGRFNLRIRITD 97
Query 181 IVHVSAC-ADCPYGKRIHVLPLDDTIEGITGNLFEIYLKPYFMEAYRPVRKNDLFLVRGG 239
V + D P +++ LP+ DT+E I GN+F+ +++P+ + P+ ++ V G
Sbjct 98 TVKLYRVDKDIPVVSKLNFLPIKDTVENIRGNIFDEFVRPFLDFNFMPLTTGSIYGVTSG 157
Query 240 FRPVEFKVVG-VDPGEFCI----VAPDTVIHCEGEPVKREE-EERLDEIGYEDIGGCRKQ 293
VEFKV +D + I V T ++C+ E + REE E+ + +GY+D+GGCR Q
Sbjct 158 LGRVEFKVTKMIDAQDMEIKHGSVTSTTSVYCD-ETISREEVEKEFNMVGYDDVGGCRAQ 216
Query 294 MAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANETGAFFFLINGP 353
MA+IRE++ELPLRH L+ +GVKPP+G+LLYGPPG+GKTLIA+A+ANETGAF FLINGP
Sbjct 217 MAKIRELVELPLRHSQLYSKIGVKPPKGILLYGPPGTGKTLIARAIANETGAFLFLINGP 276
Score = 77.0 bits (188), Expect = 6e-14, Method: Compositional matrix adjust.
Identities = 34/72 (47%), Positives = 46/72 (63%), Gaps = 0/72 (0%)
Query 282 IGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVAN 341
+ + DIGG + ++RE ++ P+ +P F G+ P +GVL YGPPG GKTL+AKAVA
Sbjct 478 VKWSDIGGLEQVKQELRETVQYPVEYPEKFIKFGMTPAKGVLFYGPPGCGKTLLAKAVAT 537
Query 342 ETGAFFFLINGP 353
E A F I GP
Sbjct 538 ECKANFISIKGP 549
> Hs20532580
Length=893
Score = 103 bits (257), Expect = 6e-22, Method: Compositional matrix adjust.
Identities = 48/99 (48%), Positives = 68/99 (68%), Gaps = 0/99 (0%)
Query 255 FCIVAPDTVIHCEGEPVKREEEERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTL 314
F ++ T ++ +E++ ++ Y+ IGG Q+ IRE+IELPL+ P LFK+
Sbjct 323 FYFISSTTRVNFTEIDKNSKEQDNQFKVTYDMIGGLSSQLKAIREIIELPLKQPELFKSY 382
Query 315 GVKPPRGVLLYGPPGSGKTLIAKAVANETGAFFFLINGP 353
G+ PRGVLLYGPPG+GKT+IA+AVANE GA+ +INGP
Sbjct 383 GIPAPRGVLLYGPPGTGKTMIARAVANEVGAYVSVINGP 421
Score = 85.9 bits (211), Expect = 1e-16, Method: Compositional matrix adjust.
Identities = 36/72 (50%), Positives = 50/72 (69%), Gaps = 0/72 (0%)
Query 282 IGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVAN 341
+ + DIGG ++ + +E PL+HP F +G++PP+GVLLYGPPG KT+IAKA+AN
Sbjct 624 VSWSDIGGLESIKLKLEQAVEWPLKHPESFIRMGIQPPKGVLLYGPPGCSKTMIAKALAN 683
Query 342 ETGAFFFLINGP 353
E+G F I GP
Sbjct 684 ESGLNFLAIKGP 695
> Hs21624654
Length=893
Score = 103 bits (256), Expect = 7e-22, Method: Compositional matrix adjust.
Identities = 48/99 (48%), Positives = 68/99 (68%), Gaps = 0/99 (0%)
Query 255 FCIVAPDTVIHCEGEPVKREEEERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTL 314
F ++ T ++ +E++ ++ Y+ IGG Q+ IRE+IELPL+ P LFK+
Sbjct 323 FYFISSTTRVNFTEIDKNSKEQDNQFKVTYDMIGGLSSQLKAIREIIELPLKQPELFKSY 382
Query 315 GVKPPRGVLLYGPPGSGKTLIAKAVANETGAFFFLINGP 353
G+ PRGVLLYGPPG+GKT+IA+AVANE GA+ +INGP
Sbjct 383 GIPAPRGVLLYGPPGTGKTMIARAVANEVGAYVSVINGP 421
Score = 85.9 bits (211), Expect = 1e-16, Method: Compositional matrix adjust.
Identities = 36/72 (50%), Positives = 50/72 (69%), Gaps = 0/72 (0%)
Query 282 IGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVAN 341
+ + DIGG ++ + +E PL+HP F +G++PP+GVLLYGPPG KT+IAKA+AN
Sbjct 624 VSWSDIGGLESIKLKLEQAVEWPLKHPESFIRMGIQPPKGVLLYGPPGCSKTMIAKALAN 683
Query 342 ETGAFFFLINGP 353
E+G F I GP
Sbjct 684 ESGLNFLAIKGP 695
> At5g58290
Length=408
Score = 99.0 bits (245), Expect = 1e-20, Method: Compositional matrix adjust.
Identities = 43/72 (59%), Positives = 54/72 (75%), Gaps = 0/72 (0%)
Query 281 EIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVA 340
++ Y DIGGC Q +IRE +ELPL H L+K +G+ PPRGVLLYGPPG+GKT++AKAVA
Sbjct 151 DVSYNDIGGCDIQKQEIREAVELPLTHHELYKQIGIDPPRGVLLYGPPGTGKTMLAKAVA 210
Query 341 NETGAFFFLING 352
N T A F + G
Sbjct 211 NHTTAAFIRVVG 222
> At1g53780
Length=464
Score = 99.0 bits (245), Expect = 1e-20, Method: Compositional matrix adjust.
Identities = 43/69 (62%), Positives = 55/69 (79%), Gaps = 0/69 (0%)
Query 284 YEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANET 343
Y DIGGC++Q+ +IRE++ELP+ HP F LG+ PP+GVL YGPPGSGKTL+A+AVAN T
Sbjct 204 YSDIGGCKEQIEKIREVVELPMLHPEKFVRLGIDPPKGVLCYGPPGSGKTLVARAVANRT 263
Query 344 GAFFFLING 352
GA F + G
Sbjct 264 GACFIRVVG 272
> ECU03g1330
Length=415
Score = 96.3 bits (238), Expect = 8e-20, Method: Compositional matrix adjust.
Identities = 54/124 (43%), Positives = 78/124 (62%), Gaps = 8/124 (6%)
Query 229 RKNDLFLVRGGFRPVEFKVVGVDPGEFCIVAPDTVIHCEGEPVKREEEERLDEIGYEDIG 288
++ D L++ G R VGVD ++ I+ P + + EER D + Y DIG
Sbjct 111 KRVDSSLIQEGMR------VGVDRNKYQIILP-LPRKIDASVTLMQVEERPD-VTYHDIG 162
Query 289 GCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANETGAFFF 348
GC++++ +IRE++E PL +P F LG+ PP+GVLLYGPPG+GKTL+A+AVAN T A F
Sbjct 163 GCKEEIEKIREVVEAPLLNPERFVALGIDPPKGVLLYGPPGTGKTLLARAVANRTNACFI 222
Query 349 LING 352
+ G
Sbjct 223 RVIG 226
> YDL007w
Length=437
Score = 95.5 bits (236), Expect = 2e-19, Method: Compositional matrix adjust.
Identities = 41/69 (59%), Positives = 54/69 (78%), Gaps = 0/69 (0%)
Query 284 YEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANET 343
Y DIGG Q+ +I+E +ELPL HP L++ +G+KPP+GV+LYG PG+GKTL+AKAVAN+T
Sbjct 181 YSDIGGLESQIQEIKESVELPLTHPELYEEMGIKPPKGVILYGAPGTGKTLLAKAVANQT 240
Query 344 GAFFFLING 352
A F I G
Sbjct 241 SATFLRIVG 249
> Hs4506207
Length=440
Score = 95.5 bits (236), Expect = 2e-19, Method: Compositional matrix adjust.
Identities = 40/69 (57%), Positives = 54/69 (78%), Gaps = 0/69 (0%)
Query 284 YEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANET 343
Y DIGG Q+ +I+E +ELPL HP ++ +G+KPP+GV+LYGPPG+GKTL+AKAVAN+T
Sbjct 184 YADIGGLDNQIQEIKESVELPLTHPEYYEEMGIKPPKGVILYGPPGTGKTLLAKAVANQT 243
Query 344 GAFFFLING 352
A F + G
Sbjct 244 SATFLRVVG 252
> 7301070
Length=439
Score = 95.1 bits (235), Expect = 2e-19, Method: Compositional matrix adjust.
Identities = 40/69 (57%), Positives = 54/69 (78%), Gaps = 0/69 (0%)
Query 284 YEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANET 343
Y DIGG Q+ +I+E +ELPL HP ++ +G+KPP+GV+LYGPPG+GKTL+AKAVAN+T
Sbjct 183 YADIGGLDTQIQEIKESVELPLTHPEYYEEMGIKPPKGVILYGPPGTGKTLLAKAVANQT 242
Query 344 GAFFFLING 352
A F + G
Sbjct 243 SATFLRVVG 251
> SPBC4.07c
Length=448
Score = 94.7 bits (234), Expect = 2e-19, Method: Compositional matrix adjust.
Identities = 40/69 (57%), Positives = 54/69 (78%), Gaps = 0/69 (0%)
Query 284 YEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANET 343
Y DIGG Q+ +I+E +ELPL HP L++ +G+KPP+GV+LYG PG+GKTL+AKAVAN+T
Sbjct 190 YADIGGLESQIQEIKEAVELPLTHPELYEEMGIKPPKGVILYGAPGTGKTLLAKAVANQT 249
Query 344 GAFFFLING 352
A F + G
Sbjct 250 SATFLRVVG 258
> At1g53750
Length=426
Score = 94.7 bits (234), Expect = 3e-19, Method: Compositional matrix adjust.
Identities = 39/72 (54%), Positives = 56/72 (77%), Gaps = 0/72 (0%)
Query 281 EIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVA 340
++ Y D+GGC++Q+ ++RE++ELP+ HP F LG+ PP+GVL YGPPG+GKTL+A+AVA
Sbjct 164 DVTYNDVGGCKEQIEKMREVVELPMLHPEKFVKLGIDPPKGVLCYGPPGTGKTLLARAVA 223
Query 341 NETGAFFFLING 352
N T A F + G
Sbjct 224 NRTDACFIRVIG 235
> 7304183
Length=433
Score = 94.7 bits (234), Expect = 3e-19, Method: Compositional matrix adjust.
Identities = 48/113 (42%), Positives = 70/113 (61%), Gaps = 18/113 (15%)
Query 248 VGVDPGEFCIVAP--------DTVIHCEGEPVKREEEERLDEIGYEDIGGCRKQMAQIRE 299
VGVD ++ I P T++ E +P ++ Y D+GGC++Q+ ++RE
Sbjct 140 VGVDRNKYQIHIPLPPKIDPTVTMMQVEDKP----------DVTYSDVGGCKEQIEKLRE 189
Query 300 MIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANETGAFFFLING 352
++E PL HP F LG++PP+GVLL+GPPG+GKTL A+AVAN T A F + G
Sbjct 190 VVETPLLHPEKFVNLGIEPPKGVLLFGPPGTGKTLCARAVANRTDACFIRVIG 242
> YKL145w
Length=467
Score = 94.4 bits (233), Expect = 3e-19, Method: Compositional matrix adjust.
Identities = 40/72 (55%), Positives = 55/72 (76%), Gaps = 0/72 (0%)
Query 281 EIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVA 340
++ Y D+GGC+ Q+ ++RE++ELPL P F TLG+ PP+G+LLYGPPG+GKTL A+AVA
Sbjct 205 DVTYSDVGGCKDQIEKLREVVELPLLSPERFATLGIDPPKGILLYGPPGTGKTLCARAVA 264
Query 341 NETGAFFFLING 352
N T A F + G
Sbjct 265 NRTDATFIRVIG 276
> CE08946
Length=435
Score = 94.4 bits (233), Expect = 4e-19, Method: Compositional matrix adjust.
Identities = 48/106 (45%), Positives = 70/106 (66%), Gaps = 4/106 (3%)
Query 248 VGVDPGEFCIVAPDTVIHCEGEP-VKREEEERLDEIGYEDIGGCRKQMAQIREMIELPLR 306
VGVD ++ I P + + +P V + E ++ Y D+GGC+ Q+ ++RE++E PL
Sbjct 142 VGVDRNKYQIHLP---LPAKIDPTVTMMQVEEKPDVTYSDVGGCKDQIEKLREVVETPLL 198
Query 307 HPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANETGAFFFLING 352
HP + LG++PP+GVLLYGPPG+GKTL A+AVAN T A F + G
Sbjct 199 HPERYVNLGIEPPKGVLLYGPPGTGKTLCARAVANRTDACFIRVIG 244
> Hs4506209
Length=433
Score = 94.4 bits (233), Expect = 4e-19, Method: Compositional matrix adjust.
Identities = 48/113 (42%), Positives = 70/113 (61%), Gaps = 18/113 (15%)
Query 248 VGVDPGEFCIVAP--------DTVIHCEGEPVKREEEERLDEIGYEDIGGCRKQMAQIRE 299
VGVD ++ I P T++ E +P ++ Y D+GGC++Q+ ++RE
Sbjct 140 VGVDRNKYQIHIPLPPKIDPTVTMMQVEEKP----------DVTYSDVGGCKEQIEKLRE 189
Query 300 MIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANETGAFFFLING 352
++E PL HP F LG++PP+GVLL+GPPG+GKTL A+AVAN T A F + G
Sbjct 190 VVETPLLHPERFVNLGIEPPKGVLLFGPPGTGKTLCARAVANRTDACFIRVIG 242
> SPBC56F2.07c
Length=809
Score = 93.2 bits (230), Expect = 7e-19, Method: Compositional matrix adjust.
Identities = 42/84 (50%), Positives = 58/84 (69%), Gaps = 4/84 (4%)
Query 274 EEEERLD----EIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPG 329
EE + D + + IGG + Q+AQIR+++ELP ++P LFK + PPRGVLLYGPPG
Sbjct 264 EETQNFDGPPSAVTFSSIGGLQAQIAQIRDIVELPFQNPELFKFFNIMPPRGVLLYGPPG 323
Query 330 SGKTLIAKAVANETGAFFFLINGP 353
+GKT++ +AVA E A F I+GP
Sbjct 324 TGKTMVMRAVAAEANAQVFTIDGP 347
Score = 82.4 bits (202), Expect = 1e-15, Method: Compositional matrix adjust.
Identities = 36/72 (50%), Positives = 48/72 (66%), Gaps = 0/72 (0%)
Query 282 IGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVAN 341
+ + DIGG + +++E +E PL H F LGV+PP+GVLLYGPPG KT+ AKA+A
Sbjct 545 VHWSDIGGQEEVKQKLKESVEWPLTHGETFSRLGVRPPKGVLLYGPPGCSKTITAKAIAT 604
Query 342 ETGAFFFLINGP 353
ETG F + GP
Sbjct 605 ETGLNFIAVKGP 616
> Hs4506213
Length=406
Score = 93.2 bits (230), Expect = 8e-19, Method: Compositional matrix adjust.
Identities = 41/76 (53%), Positives = 59/76 (77%), Gaps = 0/76 (0%)
Query 277 ERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIA 336
E++ + YE IGG KQ+ +I+E+IELP++HP LF+ LG+ P+GVLLYGPPG+GKTL+A
Sbjct 141 EKVPDSTYEMIGGLDKQIKEIKEVIELPVKHPELFEALGIAQPKGVLLYGPPGTGKTLLA 200
Query 337 KAVANETGAFFFLING 352
+AVA+ T F ++G
Sbjct 201 RAVAHHTDCTFIRVSG 216
> Hs14771027
Length=406
Score = 93.2 bits (230), Expect = 8e-19, Method: Compositional matrix adjust.
Identities = 41/76 (53%), Positives = 59/76 (77%), Gaps = 0/76 (0%)
Query 277 ERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIA 336
E++ + YE IGG KQ+ +I+E+IELP++HP LF+ LG+ P+GVLLYGPPG+GKTL+A
Sbjct 141 EKVPDSTYEMIGGLDKQIKEIKEVIELPVKHPELFEALGIAQPKGVLLYGPPGTGKTLLA 200
Query 337 KAVANETGAFFFLING 352
+AVA+ T F ++G
Sbjct 201 RAVAHHTDCTFIRVSG 216
> At2g20140
Length=443
Score = 93.2 bits (230), Expect = 8e-19, Method: Compositional matrix adjust.
Identities = 40/69 (57%), Positives = 53/69 (76%), Gaps = 0/69 (0%)
Query 284 YEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANET 343
Y DIGG Q+ +I+E +ELPL HP L++ +G+KPP+GV+LYG PG+GKTL+AKAVAN T
Sbjct 187 YADIGGLEAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLLAKAVANST 246
Query 344 GAFFFLING 352
A F + G
Sbjct 247 SATFLRVVG 255
> At4g29040
Length=443
Score = 92.8 bits (229), Expect = 9e-19, Method: Compositional matrix adjust.
Identities = 40/69 (57%), Positives = 53/69 (76%), Gaps = 0/69 (0%)
Query 284 YEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANET 343
Y DIGG Q+ +I+E +ELPL HP L++ +G+KPP+GV+LYG PG+GKTL+AKAVAN T
Sbjct 187 YADIGGLEAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLLAKAVANST 246
Query 344 GAFFFLING 352
A F + G
Sbjct 247 SATFLRVVG 255
> SPBC23G7.12c
Length=403
Score = 92.4 bits (228), Expect = 1e-18, Method: Compositional matrix adjust.
Identities = 39/76 (51%), Positives = 60/76 (78%), Gaps = 0/76 (0%)
Query 277 ERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIA 336
E++ + YE +GG KQ+ +I+E+IELP++HP LF++LG+ P+G+LLYGPPG+GKTL+A
Sbjct 137 EKIPDSTYEMVGGLEKQIKEIKEVIELPVKHPELFESLGIPQPKGILLYGPPGTGKTLLA 196
Query 337 KAVANETGAFFFLING 352
+AVA+ T F ++G
Sbjct 197 RAVAHHTDCKFIRVSG 212
> ECU10g1420
Length=401
Score = 92.4 bits (228), Expect = 1e-18, Method: Compositional matrix adjust.
Identities = 62/200 (31%), Positives = 100/200 (50%), Gaps = 16/200 (8%)
Query 163 KIRLNKVVRKNLRVRLGDIVHVSACADCPYGKRIHVLPLDDTIEGITGNLFEIYLKPYFM 222
KIR+ + L+ R+ I H + + + + + L+ + + GN+ E+ L +
Sbjct 24 KIRMLNSETRILQSRVNTIKHDISTKNASIAENMERIRLNKQLPYLVGNVVEV-LDEHSG 82
Query 223 EAYRPVRKNDLFLVRGGFRPVEFKVVGVDPGEFCIVAPDTVIHCEGEP---------VKR 273
R + + G E + PG+ + DT I E P +
Sbjct 83 VVNASTRMSSYLPITGLIPNSELR-----PGDLVALHKDTNIVFEKLPPDYDMKVGGMVL 137
Query 274 EEEERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKT 333
+ + + DE YEDIGG +Q+ ++ E I L L HP F+ L +KPP+GVL+YGPPG+GKT
Sbjct 138 KSDAKPDET-YEDIGGLERQIEELNEAIVLSLTHPERFEKLNIKPPKGVLMYGPPGTGKT 196
Query 334 LIAKAVANETGAFFFLINGP 353
L+A+A A++T A F + GP
Sbjct 197 LMARACASKTNATFLKLAGP 216
> YDR394w
Length=428
Score = 92.0 bits (227), Expect = 2e-18, Method: Compositional matrix adjust.
Identities = 39/72 (54%), Positives = 52/72 (72%), Gaps = 0/72 (0%)
Query 281 EIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVA 340
++ Y D+GG Q +IRE +ELPL L++ +G+ PPRGVLLYGPPG+GKT++ KAVA
Sbjct 168 DVTYADVGGLDMQKQEIREAVELPLVQADLYEQIGIDPPRGVLLYGPPGTGKTMLVKAVA 227
Query 341 NETGAFFFLING 352
N T A F +NG
Sbjct 228 NSTKAAFIRVNG 239
> 7301963
Length=399
Score = 92.0 bits (227), Expect = 2e-18, Method: Compositional matrix adjust.
Identities = 40/76 (52%), Positives = 58/76 (76%), Gaps = 0/76 (0%)
Query 277 ERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIA 336
E++ + YE +GG KQ+ +I+E+IELP++HP LF LG+ P+GVLLYGPPG+GKTL+A
Sbjct 135 EKVPDSTYEMVGGLDKQIQEIKEVIELPVKHPELFDALGITQPKGVLLYGPPGTGKTLLA 194
Query 337 KAVANETGAFFFLING 352
+AVA+ T F ++G
Sbjct 195 RAVAHHTECTFIRVSG 210
> ECU07g1640
Length=424
Score = 91.7 bits (226), Expect = 2e-18, Method: Compositional matrix adjust.
Identities = 39/69 (56%), Positives = 55/69 (79%), Gaps = 0/69 (0%)
Query 284 YEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANET 343
Y DIGG +Q+ +I+E +ELPL +P L++ +G+KPP+GV+LYG PG+GKTL+AKAVAN+T
Sbjct 168 YADIGGLEEQIQEIKESVELPLTNPELYQEMGIKPPKGVILYGLPGTGKTLLAKAVANQT 227
Query 344 GAFFFLING 352
A F + G
Sbjct 228 SATFLRVVG 236
> CE27676
Length=438
Score = 91.7 bits (226), Expect = 2e-18, Method: Compositional matrix adjust.
Identities = 59/163 (36%), Positives = 88/163 (53%), Gaps = 18/163 (11%)
Query 195 RIHVLPLDDTIEGIT---GNLFEIYLKPYFMEAYRPVRKNDLFLVRGG--FRPVEFKVVG 249
R+H L DD + G+ E Y+ + +R N L +V+ G F+ V +VG
Sbjct 101 RVHEL-FDDEKHAVVVVEGSRREWYVPILSIVDKDLLRLNALVMVKAGGMFKTVPSAIVG 159
Query 250 VDPGEFCIVAPDTVIHCEGEPVKREEEERLDEIGYEDIGGCRKQMAQIREMIELPLRHPT 309
V + D+ + G V++ +E D DIGGC Q+ +++E +ELPL HP
Sbjct 160 VLDDKI-----DS--NAMGHKVEKTPKETFD-----DIGGCESQIQELKESVELPLTHPE 207
Query 310 LFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANETGAFFFLING 352
++ +G+ P+GV+LYG PG+GKTL+AKAVAN T A F G
Sbjct 208 YYEEMGITAPKGVILYGEPGTGKTLLAKAVANSTSATFIRATG 250
> At5g20000
Length=419
Score = 91.7 bits (226), Expect = 2e-18, Method: Compositional matrix adjust.
Identities = 39/76 (51%), Positives = 60/76 (78%), Gaps = 0/76 (0%)
Query 277 ERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIA 336
E++ + Y+ IGG +Q+ +I+E+IELP++HP LF++LG+ P+GVLLYGPPG+GKTL+A
Sbjct 153 EKVPDSTYDMIGGLDQQIKEIKEVIELPIKHPELFESLGIAQPKGVLLYGPPGTGKTLLA 212
Query 337 KAVANETGAFFFLING 352
+AVA+ T F ++G
Sbjct 213 RAVAHHTDCTFIRVSG 228
> At5g19990
Length=405
Score = 91.7 bits (226), Expect = 2e-18, Method: Compositional matrix adjust.
Identities = 39/76 (51%), Positives = 60/76 (78%), Gaps = 0/76 (0%)
Query 277 ERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIA 336
E++ + Y+ IGG +Q+ +I+E+IELP++HP LF++LG+ P+GVLLYGPPG+GKTL+A
Sbjct 139 EKVPDSTYDMIGGLDQQIKEIKEVIELPIKHPELFESLGIAQPKGVLLYGPPGTGKTLLA 198
Query 337 KAVANETGAFFFLING 352
+AVA+ T F ++G
Sbjct 199 RAVAHHTDCTFIRVSG 214
> 7295522
Length=405
Score = 91.7 bits (226), Expect = 2e-18, Method: Compositional matrix adjust.
Identities = 40/76 (52%), Positives = 58/76 (76%), Gaps = 0/76 (0%)
Query 277 ERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIA 336
E++ + YE +GG KQ+ +I+E+IELP++HP LF LG+ P+GVLLYGPPG+GKTL+A
Sbjct 140 EKVPDSTYEMVGGLDKQIKEIKEVIELPVKHPELFDALGIAQPKGVLLYGPPGTGKTLLA 199
Query 337 KAVANETGAFFFLING 352
+AVA+ T F ++G
Sbjct 200 RAVAHHTECTFIRVSG 215
> YOR259c
Length=437
Score = 91.7 bits (226), Expect = 2e-18, Method: Compositional matrix adjust.
Identities = 40/69 (57%), Positives = 56/69 (81%), Gaps = 0/69 (0%)
Query 281 EIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVA 340
EI ++ IGG +Q+ ++RE+IELPL++P +F+ +G+KPP+GVLLYGPPG+GKTL+AKAVA
Sbjct 177 EITFDGIGGLTEQIRELREVIELPLKNPEIFQRVGIKPPKGVLLYGPPGTGKTLLAKAVA 236
Query 341 NETGAFFFL 349
GA F
Sbjct 237 ATIGANFIF 245
> ECU09g1840
Length=453
Score = 91.7 bits (226), Expect = 2e-18, Method: Compositional matrix adjust.
Identities = 40/76 (52%), Positives = 59/76 (77%), Gaps = 0/76 (0%)
Query 277 ERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIA 336
E++ + Y+ IGG +Q+ +IRE+IELP++HP LF+ LG+ P+GVLLYGPPG+GKTL+A
Sbjct 188 EKVPDSTYQMIGGLDEQIKEIREVIELPIKHPELFENLGIAQPKGVLLYGPPGTGKTLLA 247
Query 337 KAVANETGAFFFLING 352
+AVA+ T F ++G
Sbjct 248 RAVAHHTQCKFIRVSG 263
> CE22219
Length=416
Score = 91.3 bits (225), Expect = 3e-18, Method: Compositional matrix adjust.
Identities = 39/76 (51%), Positives = 58/76 (76%), Gaps = 0/76 (0%)
Query 277 ERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIA 336
E++ + YE +GG KQ+ +I+E+IELP++HP LF LG+ P+GVLL+GPPG+GKTL+A
Sbjct 151 EKVPDSTYEMVGGLDKQIKEIKEVIELPVKHPELFDALGIAQPKGVLLFGPPGTGKTLLA 210
Query 337 KAVANETGAFFFLING 352
+AVA+ T F ++G
Sbjct 211 RAVAHHTECTFIRVSG 226
> YGL048c
Length=405
Score = 90.9 bits (224), Expect = 3e-18, Method: Compositional matrix adjust.
Identities = 38/76 (50%), Positives = 60/76 (78%), Gaps = 0/76 (0%)
Query 277 ERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIA 336
E++ + Y+ +GG KQ+ +I+E+IELP++HP LF++LG+ P+GV+LYGPPG+GKTL+A
Sbjct 140 EKVPDSTYDMVGGLTKQIKEIKEVIELPVKHPELFESLGIAQPKGVILYGPPGTGKTLLA 199
Query 337 KAVANETGAFFFLING 352
+AVA+ T F ++G
Sbjct 200 RAVAHHTDCKFIRVSG 215
> SPBC16C6.07c
Length=438
Score = 90.9 bits (224), Expect = 4e-18, Method: Compositional matrix adjust.
Identities = 38/72 (52%), Positives = 55/72 (76%), Gaps = 0/72 (0%)
Query 281 EIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVA 340
++ Y D+GGC++Q+ ++RE++ELPL P F LG+ PP+G++LYGPPG+GKTL A+AVA
Sbjct 175 DVTYGDVGGCKEQIERLREVVELPLLSPERFVKLGIDPPKGIMLYGPPGTGKTLCARAVA 234
Query 341 NETGAFFFLING 352
N T A F + G
Sbjct 235 NRTDATFIRVIG 246
> 7292602
Length=413
Score = 90.9 bits (224), Expect = 4e-18, Method: Compositional matrix adjust.
Identities = 40/72 (55%), Positives = 52/72 (72%), Gaps = 0/72 (0%)
Query 281 EIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVA 340
++ Y DIGG Q +IRE +ELPL H L+K +G+ PPRGVL+YGPPG GKT++AKAVA
Sbjct 156 DVSYADIGGMDMQKQEIREAVELPLTHFELYKQIGIDPPRGVLMYGPPGCGKTMLAKAVA 215
Query 341 NETGAFFFLING 352
+ T A F + G
Sbjct 216 HHTTASFIRVVG 227
> CE09799
Length=443
Score = 90.5 bits (223), Expect = 5e-18, Method: Compositional matrix adjust.
Identities = 38/69 (55%), Positives = 54/69 (78%), Gaps = 0/69 (0%)
Query 284 YEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANET 343
Y D+GG +Q+ +I+E +ELPL HP ++ +G++PP+GV+LYG PG+GKTL+AKAVAN+T
Sbjct 187 YADVGGLDQQIQEIKEAVELPLTHPEYYEEMGIRPPKGVILYGCPGTGKTLLAKAVANQT 246
Query 344 GAFFFLING 352
A F I G
Sbjct 247 SATFLRIVG 255
> CE17915
Length=430
Score = 90.5 bits (223), Expect = 5e-18, Method: Compositional matrix adjust.
Identities = 70/265 (26%), Positives = 123/265 (46%), Gaps = 36/265 (13%)
Query 99 PKKKRSPNRLVVEEAINDDNSVVALNPSRMET-LQIFRGDTVLLRGKMRHDTVCVVLADQ 157
PK ++P + VE+AI D ++ ++ +++ + + ++R ++ Q
Sbjct 7 PKDGKNPEAMEVEDAI--DEEILKMSTEDLKSRTHLLDNEIRIMRSEV-----------Q 53
Query 158 DLDEGKIRLNKVVRKNL-RVRLGDIVHVSACADCPY--GKRIHVLPLDDTIEGITGNLFE 214
++ L + +++N R+++ + PY + +L L+D E N+
Sbjct 54 RINHSATTLKERIKENTERIKVNKTL--------PYLVSNVVELLDLEDNTEEEGANVDL 105
Query 215 IYLKPYFMEAYRPVRKNDLFLVRGGFRPVEFKVVGVDPGEFCIVAPDTVIHCEGEPVKRE 274
K R V G P E K PG+ V D+ + E P + +
Sbjct 106 DAQKTKCAVIKTSTRATYFLPVVGLVDPDELK-----PGDLVGVNKDSYLILEKLPAEYD 160
Query 275 EEERLDEIG------YEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPP 328
+ E+ Y DIGGC KQ+ ++ E + LP+ H F LG+ PP+GVL+YGPP
Sbjct 161 SRVKAMEVDERPTEQYSDIGGCDKQIQELIEAVVLPMTHKDRFVNLGIHPPKGVLMYGPP 220
Query 329 GSGKTLIAKAVANETGAFFFLINGP 353
G+GKT++A+AVA +T + F + GP
Sbjct 221 GTGKTMMARAVAAQTKSTFLKLAGP 245
> CE28467
Length=411
Score = 90.1 bits (222), Expect = 6e-18, Method: Compositional matrix adjust.
Identities = 39/76 (51%), Positives = 57/76 (75%), Gaps = 0/76 (0%)
Query 277 ERLDEIGYEDIGGCRKQMAQIREMIELPLRHPTLFKTLGVKPPRGVLLYGPPGSGKTLIA 336
E++ + YE +GG Q+ +I+E+IELP++HP LF LG+ P+GVLLYGPPG+GKTL+A
Sbjct 146 EKVPDSTYEMVGGLDTQIKEIKEVIELPVKHPELFDALGIAQPKGVLLYGPPGTGKTLLA 205
Query 337 KAVANETGAFFFLING 352
+AVA+ T F ++G
Sbjct 206 RAVAHHTECTFIRVSG 221
> At3g05530
Length=424
Score = 89.4 bits (220), Expect = 1e-17, Method: Compositional matrix adjust.
Identities = 48/107 (44%), Positives = 66/107 (61%), Gaps = 2/107 (1%)
Query 247 VVGVDPGEFCIVAPDTVIHCEGEPVKREEEERLDEIGYEDIGGCRKQMAQIREMIELPLR 306
+VGV+ + I+ DT+ VK E + Y DIGG KQ+ ++ E I LP+
Sbjct 135 LVGVNKDSYLIL--DTLPSEYDSRVKAMEVDEKPTEDYNDIGGLEKQIQELVEAIVLPMT 192
Query 307 HPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANETGAFFFLINGP 353
H F+ LGV+PP+GVLLYGPPG+GKTL+A+A A +T A F + GP
Sbjct 193 HKERFEKLGVRPPKGVLLYGPPGTGKTLMARACAAQTNATFLKLAGP 239
> At1g09100
Length=423
Score = 89.0 bits (219), Expect = 1e-17, Method: Compositional matrix adjust.
Identities = 47/107 (43%), Positives = 66/107 (61%), Gaps = 2/107 (1%)
Query 247 VVGVDPGEFCIVAPDTVIHCEGEPVKREEEERLDEIGYEDIGGCRKQMAQIREMIELPLR 306
+VGV+ + I+ DT+ VK E + Y DIGG KQ+ ++ E I LP+
Sbjct 134 LVGVNKDSYLIL--DTLPSEYDSRVKAMEVDEKPTEDYNDIGGLEKQIQELVEAIVLPMT 191
Query 307 HPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANETGAFFFLINGP 353
H F+ LG++PP+GVLLYGPPG+GKTL+A+A A +T A F + GP
Sbjct 192 HKEQFEKLGIRPPKGVLLYGPPGTGKTLMARACAAQTNATFLKLAGP 238
> HsM4506211
Length=404
Score = 89.0 bits (219), Expect = 2e-17, Method: Compositional matrix adjust.
Identities = 44/108 (40%), Positives = 65/108 (60%), Gaps = 6/108 (5%)
Query 252 PGEFCIVAPDTVIHCEGEPVKREEEERLDEIG------YEDIGGCRKQMAQIREMIELPL 305
PG+ V D+ + E P + + + E+ Y DIGG KQ+ ++ E I LP+
Sbjct 112 PGDLVGVNKDSYLILETLPTEYDSRVKAMEVDERPTEQYSDIGGLDKQIQELVEAIVLPM 171
Query 306 RHPTLFKTLGVKPPRGVLLYGPPGSGKTLIAKAVANETGAFFFLINGP 353
H F+ LG++PP+GVL+YGPPG+GKTL+A+A A +T A F + GP
Sbjct 172 NHKEKFENLGIQPPKGVLMYGPPGTGKTLLARACAAQTKATFLKLAGP 219
Lambda K H
0.319 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 8586898264
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40