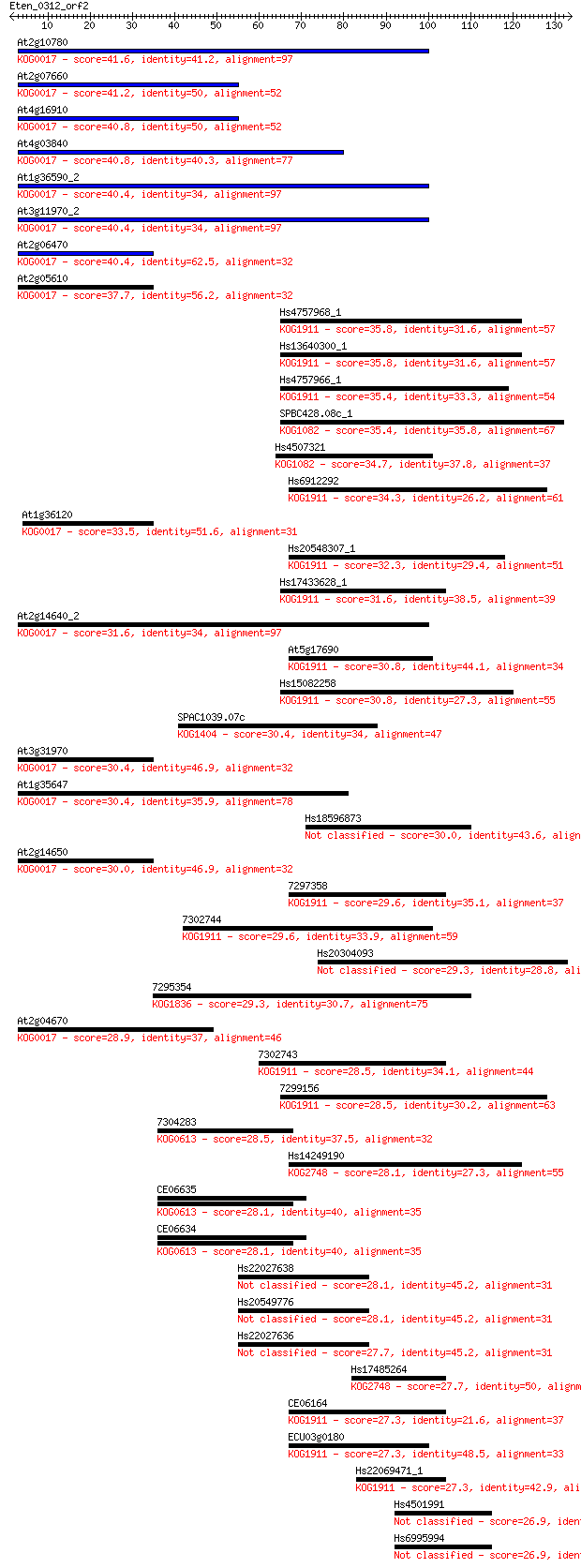
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eten_0312_orf2
Length=133
Score E
Sequences producing significant alignments: (Bits) Value
At2g10780 41.6 4e-04
At2g07660 41.2 6e-04
At4g16910 40.8 6e-04
At4g03840 40.8 7e-04
At1g36590_2 40.4 0.001
At3g11970_2 40.4 0.001
At2g06470 40.4 0.001
At2g05610 37.7 0.006
Hs4757968_1 35.8 0.023
Hs13640300_1 35.8 0.023
Hs4757966_1 35.4 0.027
SPBC428.08c_1 35.4 0.030
Hs4507321 34.7 0.052
Hs6912292 34.3 0.068
At1g36120 33.5 0.10
Hs20548307_1 32.3 0.24
Hs17433628_1 31.6 0.41
At2g14640_2 31.6 0.42
At5g17690 30.8 0.70
Hs15082258 30.8 0.78
SPAC1039.07c 30.4 0.92
At3g31970 30.4 1.0
At1g35647 30.4 1.1
Hs18596873 30.0 1.3
At2g14650 30.0 1.4
7297358 29.6 1.6
7302744 29.6 1.8
Hs20304093 29.3 2.0
7295354 29.3 2.2
At2g04670 28.9 2.7
7302743 28.5 3.7
7299156 28.5 3.8
7304283 28.5 3.9
Hs14249190 28.1 4.2
CE06635 28.1 4.8
CE06634 28.1 4.9
Hs22027638 28.1 5.1
Hs20549776 28.1 5.3
Hs22027636 27.7 6.0
Hs17485264 27.7 6.7
CE06164 27.3 7.7
ECU03g0180 27.3 8.3
Hs22069471_1 27.3 8.7
Hs4501991 26.9 9.6
Hs6995994 26.9 9.9
> At2g10780
Length=1611
Score = 41.6 bits (96), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 40/112 (35%), Positives = 46/112 (41%), Gaps = 32/112 (28%)
Query 3 PSEVIKRIGTVAYCLARPPTYEC-HNVFHVSQL--------VPHRPRPPALALPEADAAW 53
P +VI+R+G VAY L PP HNVFHVSQL PP L AW
Sbjct 1475 PYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQLRKCLSDQEESVEDIPPGLKENMTVEAW 1534
Query 54 PPIRDAGGNPIEEYEADYIMDRLGSG------DAAQYLVKRRGTPEDQATWE 99
P+R IMDR+ G D + L RG E TWE
Sbjct 1535 -PVR--------------IMDRMTKGTRGKARDLLKVLWNCRGREE--YTWE 1569
> At2g07660
Length=949
Score = 41.2 bits (95), Expect = 6e-04, Method: Composition-based stats.
Identities = 26/61 (42%), Positives = 30/61 (49%), Gaps = 9/61 (14%)
Query 3 PSEVIKRIGTVAYCLARPPTYEC-HNVFHVSQLVPHRPR--------PPALALPEADAAW 53
P +VI+R+G VAY L PP HNVFHVSQL + PP L AW
Sbjct 867 PYKVIERVGAVAYKLDLPPKLNVFHNVFHVSQLRKYLSDQEESVEDIPPGLKENMTVEAW 926
Query 54 P 54
P
Sbjct 927 P 927
> At4g16910
Length=687
Score = 40.8 bits (94), Expect = 6e-04, Method: Composition-based stats.
Identities = 26/61 (42%), Positives = 30/61 (49%), Gaps = 9/61 (14%)
Query 3 PSEVIKRIGTVAYCLARPPTYEC-HNVFHVSQL--------VPHRPRPPALALPEADAAW 53
P +VI+R+G VAY L PP + HNVFHVSQL PP L AW
Sbjct 559 PYKVIERVGAVAYKLDLPPKLDAFHNVFHVSQLRKCLSEQEESMEDVPPGLKENMTVEAW 618
Query 54 P 54
P
Sbjct 619 P 619
> At4g03840
Length=973
Score = 40.8 bits (94), Expect = 7e-04, Method: Compositional matrix adjust.
Identities = 31/86 (36%), Positives = 35/86 (40%), Gaps = 24/86 (27%)
Query 3 PSEVIKRIGTVAYCLARPPTYEC-HNVFHVSQLVPHRPR--------PPALALPEADAAW 53
P +VI+R+G VAY L PP HNVFHVSQL PP L AW
Sbjct 837 PYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQLRKCLSNQEESVEDVPPGLKENMTVEAW 896
Query 54 PPIRDAGGNPIEEYEADYIMDRLGSG 79
P IMDR+ G
Sbjct 897 PV---------------QIMDRMTKG 907
> At1g36590_2
Length=958
Score = 40.4 bits (93), Expect = 0.001, Method: Composition-based stats.
Identities = 33/98 (33%), Positives = 46/98 (46%), Gaps = 8/98 (8%)
Query 3 PSEVIKRIGTVAYCLARPPTYECHNVFHVSQL-VPHRPRPPALALPEADAAWPPIRDAGG 61
P ++I R G VAY LA P + H VFHVSQL V + LP ++D
Sbjct 854 PYKIIDRCGEVAYKLALPSYSQVHPVFHVSQLKVLVGNVSTTVHLPSV------MQDVFE 907
Query 62 NPIEEYEADYIMDRLGSGDAAQYLVKRRGTPEDQATWE 99
E+ +++R G + LVK P ++ATWE
Sbjct 908 KVPEKVVERKMVNRQGKA-VTKVLVKWSNEPLEEATWE 944
> At3g11970_2
Length=958
Score = 40.4 bits (93), Expect = 0.001, Method: Composition-based stats.
Identities = 33/98 (33%), Positives = 46/98 (46%), Gaps = 8/98 (8%)
Query 3 PSEVIKRIGTVAYCLARPPTYECHNVFHVSQL-VPHRPRPPALALPEADAAWPPIRDAGG 61
P ++I R G VAY LA P + H VFHVSQL V + LP ++D
Sbjct 854 PYKIIDRCGEVAYKLALPSYSQVHPVFHVSQLKVLVGNVSTTVHLPSV------MQDVFE 907
Query 62 NPIEEYEADYIMDRLGSGDAAQYLVKRRGTPEDQATWE 99
E+ +++R G + LVK P ++ATWE
Sbjct 908 KVPEKVVERKMVNRQGKA-VTKVLVKWSNEPLEEATWE 944
> At2g06470
Length=899
Score = 40.4 bits (93), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 20/33 (60%), Positives = 23/33 (69%), Gaps = 1/33 (3%)
Query 3 PSEVIKRIGTVAYCLARPPTYEC-HNVFHVSQL 34
P +VI+R+G VAY L PP HNVFHVSQL
Sbjct 834 PYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQL 866
> At2g05610
Length=780
Score = 37.7 bits (86), Expect = 0.006, Method: Composition-based stats.
Identities = 18/32 (56%), Positives = 20/32 (62%), Gaps = 0/32 (0%)
Query 3 PSEVIKRIGTVAYCLARPPTYECHNVFHVSQL 34
P +VI R G VAY L P + H VFHVSQL
Sbjct 723 PYKVIDRCGEVAYKLQLPANSQVHPVFHVSQL 754
> Hs4757968_1
Length=227
Score = 35.8 bits (81), Expect = 0.023, Method: Compositional matrix adjust.
Identities = 18/58 (31%), Positives = 28/58 (48%), Gaps = 1/58 (1%)
Query 65 EEYEADYIMD-RLGSGDAAQYLVKRRGTPEDQATWEPAHHPTGGPALLRAWRRRQRRR 121
+E+E + I+D R QYLV+ +G + TWEP H + + RRQ +
Sbjct 4 QEFEVEAIVDKRQDKNGNTQYLVRWKGYDKQDDTWEPEQHLMNCEKCVHDFNRRQTEK 61
> Hs13640300_1
Length=227
Score = 35.8 bits (81), Expect = 0.023, Method: Compositional matrix adjust.
Identities = 18/58 (31%), Positives = 28/58 (48%), Gaps = 1/58 (1%)
Query 65 EEYEADYIMD-RLGSGDAAQYLVKRRGTPEDQATWEPAHHPTGGPALLRAWRRRQRRR 121
+E+E + I+D R QYLV+ +G + TWEP H + + RRQ +
Sbjct 4 QEFEVEAIVDKRQDKNGNTQYLVRWKGYDKQDDTWEPEQHLMNCEKCVHDFNRRQTEK 61
> Hs4757966_1
Length=226
Score = 35.4 bits (80), Expect = 0.027, Method: Compositional matrix adjust.
Identities = 18/55 (32%), Positives = 27/55 (49%), Gaps = 1/55 (1%)
Query 65 EEYEADYIMD-RLGSGDAAQYLVKRRGTPEDQATWEPAHHPTGGPALLRAWRRRQ 118
+E+E + I+D R QYLV+ +G + TWEP H + + RRQ
Sbjct 4 QEFEVEAIVDKRQDKNGNTQYLVRWKGYDKQDDTWEPEQHLMNCEKCVHDFNRRQ 58
> SPBC428.08c_1
Length=126
Score = 35.4 bits (80), Expect = 0.030, Method: Compositional matrix adjust.
Identities = 24/69 (34%), Positives = 37/69 (53%), Gaps = 3/69 (4%)
Query 65 EEYEADYIMD-RLGSGDAAQ-YLVKRRGTPEDQATWEPAHHPTGGPALLRAWRRRQRRRL 122
EEYE + I+D +L A + Y ++ TWEP + +G A+L W+RR +RRL
Sbjct 6 EEYEVERIVDEKLDRNGAVKLYRIRWLNYSSRSDTWEPPENLSGCSAVLAEWKRR-KRRL 64
Query 123 QARNNIGPS 131
+ N+ S
Sbjct 65 KGSNSDSDS 73
> Hs4507321
Length=412
Score = 34.7 bits (78), Expect = 0.052, Method: Compositional matrix adjust.
Identities = 14/37 (37%), Positives = 23/37 (62%), Gaps = 0/37 (0%)
Query 64 IEEYEADYIMDRLGSGDAAQYLVKRRGTPEDQATWEP 100
+ ++E +Y+ D + YLVK RG P+ ++TWEP
Sbjct 40 LYDFEVEYLCDYKKIREQEYYLVKWRGYPDSESTWEP 76
> Hs6912292
Length=191
Score = 34.3 bits (77), Expect = 0.068, Method: Compositional matrix adjust.
Identities = 16/61 (26%), Positives = 31/61 (50%), Gaps = 1/61 (1%)
Query 67 YEADYIMDRLGSGDAAQYLVKRRGTPEDQATWEPAHHPTGGPALLRAWRRRQRRRLQARN 126
Y + ++DR +YL+K +G E+ TWEP + P L+ + ++ ++ + N
Sbjct 20 YVVEKVLDRRVVKGQVEYLLKWKGFSEEHNTWEPEKN-LDCPELISEFMKKYKKMKEGEN 78
Query 127 N 127
N
Sbjct 79 N 79
> At1g36120
Length=1235
Score = 33.5 bits (75), Expect = 0.10, Method: Compositional matrix adjust.
Identities = 16/32 (50%), Positives = 21/32 (65%), Gaps = 1/32 (3%)
Query 4 SEVIKRIGTVAYCLARPPTYEC-HNVFHVSQL 34
+E+++R+G VAY L P HNVFHVS L
Sbjct 1123 TEIVERVGPVAYRLELPDVMRAFHNVFHVSML 1154
> Hs20548307_1
Length=227
Score = 32.3 bits (72), Expect = 0.24, Method: Compositional matrix adjust.
Identities = 15/51 (29%), Positives = 24/51 (47%), Gaps = 0/51 (0%)
Query 67 YEADYIMDRLGSGDAAQYLVKRRGTPEDQATWEPAHHPTGGPALLRAWRRR 117
+E + I+D G Y V+ +G D TWEP H +L +R++
Sbjct 59 FEVEKILDMKTEGGKVLYKVRWKGYTSDDDTWEPEIHLEDCKEVLLEFRKK 109
> Hs17433628_1
Length=179
Score = 31.6 bits (70), Expect = 0.41, Method: Compositional matrix adjust.
Identities = 15/40 (37%), Positives = 22/40 (55%), Gaps = 1/40 (2%)
Query 65 EEYEADYIMD-RLGSGDAAQYLVKRRGTPEDQATWEPAHH 103
+E+E + I+D R G +YLVK + + TWEP H
Sbjct 4 QEFEVETIVDKRQGKNVNIEYLVKWKAYDKQYDTWEPKQH 43
> At2g14640_2
Length=492
Score = 31.6 bits (70), Expect = 0.42, Method: Compositional matrix adjust.
Identities = 33/101 (32%), Positives = 43/101 (42%), Gaps = 12/101 (11%)
Query 3 PSEVIKRIGTVAYCLARPPTYECHNVFHVSQLVPHRPRPPALALPEADAA-WPPIRDAGG 61
P +V + G VAY L P H VFHVS L P + E D PP+R+ G
Sbjct 378 PFQVASKHGEVAYRLTLPEGTRIHPVFHVSLL------KPWVGDGEPDMGQLPPLRNNGE 431
Query 62 NPIEEYEADYIMDRLGSGD---AAQYLVKRRGTPEDQATWE 99
++ + R S D A LV+ G + ATWE
Sbjct 432 LKLQPTAVLEV--RWRSQDKKRVADLLVQWEGLHIEDATWE 470
> At5g17690
Length=445
Score = 30.8 bits (68), Expect = 0.70, Method: Composition-based stats.
Identities = 15/34 (44%), Positives = 18/34 (52%), Gaps = 0/34 (0%)
Query 67 YEADYIMDRLGSGDAAQYLVKRRGTPEDQATWEP 100
YE + I + QYL+K RG PE TWEP
Sbjct 108 YEIEAIRRKRVRKGKVQYLIKWRGWPETANTWEP 141
> Hs15082258
Length=183
Score = 30.8 bits (68), Expect = 0.78, Method: Compositional matrix adjust.
Identities = 15/55 (27%), Positives = 27/55 (49%), Gaps = 1/55 (1%)
Query 65 EEYEADYIMDRLGSGDAAQYLVKRRGTPEDQATWEPAHHPTGGPALLRAWRRRQR 119
EE+ + ++DR +Y +K +G + TWEP + P L+ A+ Q+
Sbjct 28 EEFVVEKVLDRRVVNGKVEYFLKWKGFTDADNTWEPEEN-LDCPELIEAFLNSQK 81
> SPAC1039.07c
Length=448
Score = 30.4 bits (67), Expect = 0.92, Method: Composition-based stats.
Identities = 16/50 (32%), Positives = 26/50 (52%), Gaps = 3/50 (6%)
Query 41 PPALALPEADAAWPPIRDAGGNPIEEYEAD---YIMDRLGSGDAAQYLVK 87
P + +PE + P RDA GN + E D Y++D+ +G A +V+
Sbjct 169 PGSYTIPEPNPKLSPFRDAKGNYDWQKELDYSFYMLDKQSTGSLACMIVE 218
> At3g31970
Length=1329
Score = 30.4 bits (67), Expect = 1.0, Method: Compositional matrix adjust.
Identities = 15/33 (45%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Query 3 PSEVIKRIGTVAYCLARPPTYEC-HNVFHVSQL 34
P +++R+G VAY L P H VFHVS L
Sbjct 1216 PFRIVERVGPVAYRLELPDVMRAFHKVFHVSML 1248
> At1g35647
Length=1495
Score = 30.4 bits (67), Expect = 1.1, Method: Composition-based stats.
Identities = 28/86 (32%), Positives = 36/86 (41%), Gaps = 9/86 (10%)
Query 3 PSEVIKRIGTVAYCLARPPTYECHNVFHVSQLVPHRP-----RPPALALPEADAAWPPIR 57
P +V+KRI AY L Y N F+V+ L P R L E D +
Sbjct 1306 PFKVLKRINNNAYSLDLQGKYNVSNSFNVADLFPFIADNTDLRSNPFQLGEDDVIMTSL- 1364
Query 58 DAGGNPIE---EYEADYIMDRLGSGD 80
D G + I ++ AD IM L GD
Sbjct 1365 DHGADEIMTSLDHGADEIMTSLDHGD 1390
> Hs18596873
Length=234
Score = 30.0 bits (66), Expect = 1.3, Method: Compositional matrix adjust.
Identities = 17/46 (36%), Positives = 20/46 (43%), Gaps = 7/46 (15%)
Query 71 YIMDRLGS-------GDAAQYLVKRRGTPEDQATWEPAHHPTGGPA 109
Y+MD+LG D + L RR ED W HHP PA
Sbjct 20 YLMDQLGLRLGMWYWKDETRTLEFRRFAAEDSVQWLLKHHPHFTPA 65
> At2g14650
Length=1328
Score = 30.0 bits (66), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 15/33 (45%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Query 3 PSEVIKRIGTVAYCLARPPTYEC-HNVFHVSQL 34
P +++R+G VAY L P H VFHVS L
Sbjct 1218 PFRIVERVGPVAYRLELPDVMRAFHKVFHVSML 1250
> 7297358
Length=206
Score = 29.6 bits (65), Expect = 1.6, Method: Compositional matrix adjust.
Identities = 13/37 (35%), Positives = 21/37 (56%), Gaps = 0/37 (0%)
Query 67 YEADYIMDRLGSGDAAQYLVKRRGTPEDQATWEPAHH 103
Y + I+DR +Y +K +G PE + TWEP ++
Sbjct 24 YAVEKIIDRRVRKGKVEYYLKWKGYPETENTWEPENN 60
> 7302744
Length=418
Score = 29.6 bits (65), Expect = 1.8, Method: Composition-based stats.
Identities = 20/59 (33%), Positives = 27/59 (45%), Gaps = 9/59 (15%)
Query 42 PALALPEADAAWPPIRDAGGNPIEEYEADYIMDRLGSGDAAQYLVKRRGTPEDQATWEP 100
P L L +A PP + +EEY + I+ + Q LVK G P + TWEP
Sbjct 8 PNLGLVDA----PP-----NDHVEEYVVEKILGKRFVNGRPQVLVKWSGFPNENNTWEP 57
> Hs20304093
Length=890
Score = 29.3 bits (64), Expect = 2.0, Method: Composition-based stats.
Identities = 17/59 (28%), Positives = 26/59 (44%), Gaps = 0/59 (0%)
Query 74 DRLGSGDAAQYLVKRRGTPEDQATWEPAHHPTGGPALLRAWRRRQRRRLQARNNIGPSE 132
DRLG +++ +K GTP + GG + RR+Q R + R +G E
Sbjct 13 DRLGRRSSSKRALKAEGTPGRRGAQRSQKERAGGSPSPGSPRRKQTGRRRHREELGEQE 71
> 7295354
Length=634
Score = 29.3 bits (64), Expect = 2.2, Method: Composition-based stats.
Identities = 23/85 (27%), Positives = 35/85 (41%), Gaps = 11/85 (12%)
Query 35 VPHRPRPPALALPEADAAW--PPI-RDAGGNPI-------EEYEADYIMDRLGSGDAAQY 84
VP R PP A+ +A W PP+ R N + +E+ Y+ R+G+
Sbjct 83 VPERNHPPENAIDGTEAWWQSPPLSRGMKFNEVNLTINFEQEFHVAYLFIRMGNSPRPGL 142
Query 85 LVKRRGTPEDQATWEPAHHPTGGPA 109
+ T + TW P H + PA
Sbjct 143 WTLEKSTDYGK-TWTPWQHFSDTPA 166
> At2g04670
Length=1411
Score = 28.9 bits (63), Expect = 2.7, Method: Compositional matrix adjust.
Identities = 17/49 (34%), Positives = 24/49 (48%), Gaps = 3/49 (6%)
Query 3 PSEVIKRIGTVAYCLARPPTYEC-HNVFHVSQL--VPHRPRPPALALPE 48
P +++R+G VAY L P H VFHV L H+ + +PE
Sbjct 1298 PFRIVERVGPVAYRLELPDVMRAFHKVFHVLMLRKCLHKDDEVLVKIPE 1346
> 7302743
Length=84
Score = 28.5 bits (62), Expect = 3.7, Method: Compositional matrix adjust.
Identities = 15/48 (31%), Positives = 24/48 (50%), Gaps = 4/48 (8%)
Query 60 GGNPIEEYEADYIMDRLGSGDAA----QYLVKRRGTPEDQATWEPAHH 103
G ++E ++YI+++ QYL K G P +Q TWEP +
Sbjct 11 GVRNVKEKSSEYIVEKFLGKRYLRGRPQYLTKWEGYPIEQCTWEPLEN 58
> 7299156
Length=160
Score = 28.5 bits (62), Expect = 3.8, Method: Compositional matrix adjust.
Identities = 19/63 (30%), Positives = 27/63 (42%), Gaps = 0/63 (0%)
Query 65 EEYEADYIMDRLGSGDAAQYLVKRRGTPEDQATWEPAHHPTGGPALLRAWRRRQRRRLQA 124
EEY + I+DR QYLVK ++ TWE A + ++ +R Q
Sbjct 25 EEYIVERILDRRHYMGQLQYLVKWLDYSDEDNTWESAADLDCHSLIDSIESQKSLKRGQE 84
Query 125 RNN 127
NN
Sbjct 85 LNN 87
> 7304283
Length=7175
Score = 28.5 bits (62), Expect = 3.9, Method: Composition-based stats.
Identities = 12/32 (37%), Positives = 17/32 (53%), Gaps = 0/32 (0%)
Query 36 PHRPRPPALALPEADAAWPPIRDAGGNPIEEY 67
P +P P D AW P ++ GG PI++Y
Sbjct 1846 PGKPEPTNWDKDFVDLAWDPPKNDGGAPIQKY 1877
> Hs14249190
Length=211
Score = 28.1 bits (61), Expect = 4.2, Method: Compositional matrix adjust.
Identities = 15/55 (27%), Positives = 28/55 (50%), Gaps = 1/55 (1%)
Query 67 YEADYIMDRLGSGDAAQYLVKRRGTPEDQATWEPAHHPTGGPALLRAWRRRQRRR 121
+ A+ I+ + +YLVK RG +WEP + P LL A+++++ +
Sbjct 12 FAAECILSKRLRKGKLEYLVKWRGWSSKHNSWEPEENIL-DPRLLLAFQKKEHEK 65
> CE06635
Length=7160
Score = 28.1 bits (61), Expect = 4.8, Method: Compositional matrix adjust.
Identities = 14/35 (40%), Positives = 16/35 (45%), Gaps = 0/35 (0%)
Query 36 PHRPRPPALALPEADAAWPPIRDAGGNPIEEYEAD 70
P RP P D W P GG PIEEY+ +
Sbjct 2877 PGRPEPTDWDSDHVDLKWDPPLSDGGAPIEEYQIE 2911
Score = 27.3 bits (59), Expect = 7.9, Method: Compositional matrix adjust.
Identities = 14/32 (43%), Positives = 14/32 (43%), Gaps = 0/32 (0%)
Query 36 PHRPRPPALALPEADAAWPPIRDAGGNPIEEY 67
P RP D AW P GG PIEEY
Sbjct 1997 PGRPEITDFDADRIDIAWEPPHKDGGAPIEEY 2028
> CE06634
Length=6831
Score = 28.1 bits (61), Expect = 4.9, Method: Compositional matrix adjust.
Identities = 14/35 (40%), Positives = 16/35 (45%), Gaps = 0/35 (0%)
Query 36 PHRPRPPALALPEADAAWPPIRDAGGNPIEEYEAD 70
P RP P D W P GG PIEEY+ +
Sbjct 2548 PGRPEPTDWDSDHVDLKWDPPLSDGGAPIEEYQIE 2582
Score = 27.3 bits (59), Expect = 8.3, Method: Compositional matrix adjust.
Identities = 14/32 (43%), Positives = 14/32 (43%), Gaps = 0/32 (0%)
Query 36 PHRPRPPALALPEADAAWPPIRDAGGNPIEEY 67
P RP D AW P GG PIEEY
Sbjct 1668 PGRPEITDFDADRIDIAWEPPHKDGGAPIEEY 1699
> Hs22027638
Length=1565
Score = 28.1 bits (61), Expect = 5.1, Method: Compositional matrix adjust.
Identities = 14/31 (45%), Positives = 18/31 (58%), Gaps = 0/31 (0%)
Query 55 PIRDAGGNPIEEYEADYIMDRLGSGDAAQYL 85
P++DAGG E EA + RLG+ DA L
Sbjct 599 PVKDAGGGTGREAEARELRFRLGTSDATGSL 629
> Hs20549776
Length=1565
Score = 28.1 bits (61), Expect = 5.3, Method: Compositional matrix adjust.
Identities = 14/31 (45%), Positives = 18/31 (58%), Gaps = 0/31 (0%)
Query 55 PIRDAGGNPIEEYEADYIMDRLGSGDAAQYL 85
P++DAGG E EA + RLG+ DA L
Sbjct 599 PVKDAGGGTGREAEARELRFRLGTSDATGSL 629
> Hs22027636
Length=1253
Score = 27.7 bits (60), Expect = 6.0, Method: Compositional matrix adjust.
Identities = 14/31 (45%), Positives = 18/31 (58%), Gaps = 0/31 (0%)
Query 55 PIRDAGGNPIEEYEADYIMDRLGSGDAAQYL 85
P++DAGG E EA + RLG+ DA L
Sbjct 599 PVKDAGGGTGREAEARELRFRLGTSDATGSL 629
> Hs17485264
Length=259
Score = 27.7 bits (60), Expect = 6.7, Method: Compositional matrix adjust.
Identities = 11/22 (50%), Positives = 14/22 (63%), Gaps = 0/22 (0%)
Query 82 AQYLVKRRGTPEDQATWEPAHH 103
+YLVK +G P +TWEP H
Sbjct 26 VEYLVKWKGWPPKYSTWEPEEH 47
> CE06164
Length=184
Score = 27.3 bits (59), Expect = 7.7, Method: Compositional matrix adjust.
Identities = 8/37 (21%), Positives = 22/37 (59%), Gaps = 0/37 (0%)
Query 67 YEADYIMDRLGSGDAAQYLVKRRGTPEDQATWEPAHH 103
+ + ++++ + ++Y +K +G PE + +WEP +
Sbjct 37 FVVEKVLNKRLTRGGSEYYIKWQGFPESECSWEPIEN 73
> ECU03g0180
Length=156
Score = 27.3 bits (59), Expect = 8.3, Method: Compositional matrix adjust.
Identities = 16/34 (47%), Positives = 19/34 (55%), Gaps = 2/34 (5%)
Query 67 YEADYIM-DRLGSGDAAQYLVKRRGTPEDQATWE 99
Y D I+ DR G QYLVK G P+ + TWE
Sbjct 7 YTVDRIVGDRKKKG-VKQYLVKWEGYPDSENTWE 39
> Hs22069471_1
Length=538
Score = 27.3 bits (59), Expect = 8.7, Method: Compositional matrix adjust.
Identities = 9/21 (42%), Positives = 14/21 (66%), Gaps = 0/21 (0%)
Query 83 QYLVKRRGTPEDQATWEPAHH 103
+YL++ +G + TWEP HH
Sbjct 173 EYLIRWKGYGSTEDTWEPEHH 193
> Hs4501991
Length=2316
Score = 26.9 bits (58), Expect = 9.6, Method: Compositional matrix adjust.
Identities = 12/26 (46%), Positives = 15/26 (57%), Gaps = 3/26 (11%)
Query 92 PEDQATWEPAHHPTGG---PALLRAW 114
PE+Q WEPA+ P G P +L W
Sbjct 735 PENQTEWEPAYTPVGTSPLPGILPTW 760
> Hs6995994
Length=2415
Score = 26.9 bits (58), Expect = 9.9, Method: Compositional matrix adjust.
Identities = 12/26 (46%), Positives = 15/26 (57%), Gaps = 3/26 (11%)
Query 92 PEDQATWEPAHHPTGG---PALLRAW 114
PE+Q WEPA+ P G P +L W
Sbjct 735 PENQTEWEPAYTPVGTSPLPGILPTW 760
Lambda K H
0.319 0.136 0.443
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1356426142
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40