bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: eggV2
2,483,276 sequences; 915,453,621 total letters
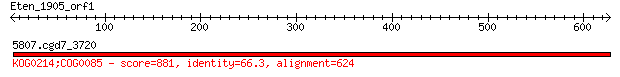
Query= Eten_1905_orf1
Length=627
Score E
Sequences producing significant alignments: (Bits) Value
5807.cgd7_3720 881 0.0
> 5807.cgd7_3720
Length=1177
Score = 881 bits (2276), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 414/633 (65%), Positives = 506/633 (79%), Gaps = 17/633 (2%)
Query 4 PESIKDRCKVFINGDWAGCFDEAETLCNTLKEIRRKGHIRNDTSIVRDIINREVKIFTDA 63
PE IK + KVF+NG+W GCFD++E + L+ IRR G I +TSIV D++NREVK+FTD+
Sbjct 553 PEGIKRKIKVFLNGNWVGCFDDSEDSISNLRMIRRSGEIPYETSIVLDVVNREVKLFTDS 612
Query 64 GRAMRPLYVVDSDTGQLRIRKADVDKVRS-------MVGTSEKGAGWNHLMSCGVIEYID 116
GR+MRPLY+V D G L+I+K+ ++ + + + + W+ L++ G+IEYID
Sbjct 613 GRSMRPLYIVGED-GDLKIKKSHINSLLNENPSFDEKMEDEKHDYTWDDLVNTGIIEYID 671
Query 117 CEEEETSMVAMFHKDLS-EKGKSIPYTHCEIHPSLILGVCASIIPFPDHNQSPRNVYQSA 175
CEEEETSM+AMF DL ++G YTHCEIHPSLILGVCASIIPFPDHNQSPRNVYQSA
Sbjct 672 CEEEETSMIAMFINDLRIDRGYCSTYTHCEIHPSLILGVCASIIPFPDHNQSPRNVYQSA 731
Query 176 MGKQAMGVYTTNFNLRLDTLAHLLYYPQKPLVCTRAMEHLKFRELPAGINTIVAILCYSG 235
MGKQAMGVYT+N+N+R+DTL ++LYYPQKPLV TRAM +++FRELPAGIN IV I+CYSG
Sbjct 732 MGKQAMGVYTSNYNVRMDTLGYVLYYPQKPLVTTRAMTYMRFRELPAGINCIVGIMCYSG 791
Query 236 YNQEDSLIMNRNSIDRGLFRSLFCRTYCAEEKQQGSMIVEGFEQPNPETTSGMKRGDYSK 295
YNQEDSLIMN++SIDRGLFRS+F RTY +EEKQ GS ++E FE PNPE SG++ G+Y
Sbjct 792 YNQEDSLIMNQSSIDRGLFRSVFFRTYVSEEKQVGSNLIEAFELPNPEEVSGLRFGNYGN 851
Query 296 LDTDGLVEPGSRVLGDDVIVGKTSPNFDDNDPLAPPGAEANLSK-TKKDCSLCLRSSESG 354
LD DGL+EPG+RVLGDD+I+GK P + P + + K TK+DCS+ +R+SE G
Sbjct 852 LDLDGLIEPGNRVLGDDIIIGKVGP-------INPEDRDTRIQKLTKRDCSVGIRTSEHG 904
Query 355 VVDTVMLSVNSKGSRFVKVKVRSVRIPQTGDKFASRHGQKGTIGITYRQEDMPFTEDGVV 414
VVD V+LS NSKG +F KVKVR++RIP GDKFASRHGQKGTIGITYR EDMPFT+DG++
Sbjct 905 VVDEVLLSTNSKGVKFAKVKVRTIRIPHVGDKFASRHGQKGTIGITYRTEDMPFTQDGII 964
Query 415 PDLLMNPHAVPSRMTIGHLVECLLGKTAAIIGGEGDATPFNDYTVSWIAGELHRLGFERH 474
PD++MNPHA+PSRMTIGHL+ECLLGK AI G EGDATPF TV I+ LH GF+++
Sbjct 965 PDIIMNPHAIPSRMTIGHLIECLLGKACAIQGLEGDATPFTKITVEEISSRLHSSGFQKN 1024
Query 475 GNERLYHGHTGLHMPSLIFIGPTYYQRLKHMVDDKIHARARGPVAALTRQPMEGKSREGG 534
GNE +Y+GHTG + S IFIGPTYYQRLKHMVDDKIHARARG V LTRQP EG++REGG
Sbjct 1025 GNEVMYNGHTGKRLESRIFIGPTYYQRLKHMVDDKIHARARGAVTMLTRQPREGRAREGG 1084
Query 535 LRFGEMERDCMISHGAARMLKERFFDQSDAYRIHCCEICGTMCVADLQRLAFECKLCGNS 594
LRFGEMERDCMISHGAA+MLKER FDQ DAYR+H CE+CG +C+ADL + FEC C N
Sbjct 1085 LRFGEMERDCMISHGAAKMLKERLFDQCDAYRVHVCEMCGLICIADLGKQHFECNACNNK 1144
Query 595 NKISQVYIPYACKLLLQELTSMCIYPKLVLQVA 627
++ISQ+ +PYACKLLLQEL +M IYP+L LQ A
Sbjct 1145 SRISQICLPYACKLLLQELMAMSIYPRLHLQTA 1177
Lambda K H
0.321 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 287268456041
Database: eggV2
Posted date: Dec 15, 2009 4:47 PM
Number of letters in database: 915,453,621
Number of sequences in database: 2,483,276
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40