bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: eggV2
2,483,276 sequences; 915,453,621 total letters
Query= Eten_1464_orf14
Length=1524
Score E
Sequences producing significant alignments: (Bits) Value
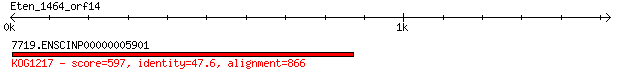
7719.ENSCINP00000005901 597 2e-168
> 7719.ENSCINP00000005901
Length=1187
Score = 597 bits (1538), Expect = 2e-168, Method: Compositional matrix adjust.
Identities = 412/947 (43%), Positives = 513/947 (54%), Gaps = 122/947 (12%)
Query 7 KPGYELVDGKCVKIDFCARGA--CNSLAHCKENPEGTAAICTCIAGYSGDGTAQGHCDDI 64
K G+ C ID CA C++ A+C N G + C C G++G+G +C DI
Sbjct 2 KTGFTGDGVNCTDIDECALNTDNCDTNANCT-NTIG-SFTCACKTGFTGNGV---NCTDI 56
Query 65 DECLAENDCTPADQGGICENTVGSYTCKCAAGYQQDGNSCTDIDECANGTHNCHASATCT 124
DEC D D C NT+GS+TC C G+ DG SCTDIDECA T NCH +A CT
Sbjct 57 DECALNTD--NCDTNANCTNTIGSFTCTCKTGFTGDGVSCTDIDECALNTDNCHTNANCT 114
Query 125 NTQGSFECACNAGFSGNGVECNDVDECSTDADDCGENTLCNNTVGSFECTCMAGFEAADA 184
NT GSF C C GFSG+GV C D+DEC+ + ++C N C NT+GSF CTC GF + +
Sbjct 115 NTVGSFTCTCKTGFSGDGVSCTDIDECALNTNNCDTNANCTNTIGSFTCTCTTGF-SGNG 173
Query 185 KTCKDIDECASGTHTCSTHATCTNTAGSFTCECNPGFDGDGHKCEDVDFCGQGLHDCNVH 244
TC D+DECA T C+ +A CTNT GSFTC C GF GDG C D+D C +C +
Sbjct 174 TTCIDLDECALNTDNCNINANCTNTIGSFTCTCKTGFTGDGVTCTDIDECALNTDNCITN 233
Query 245 AECSESDDNTT---------------------------------------FKCTCGIGYS 265
A C+ + + T F CTC G++
Sbjct 234 ANCTNTIGSFTCTCKTGFTGNGVTCTDIDECALNTDNCNTNANCTNTIGSFTCTCKTGFT 293
Query 266 GEGHGENGCQDIDECAQDA-ICGENTVCTNTPGSFECACVEGFVAVGAKLKGATSLTCID 324
G+G C DIDECA + C N CTNT GSF C C GF G +TC D
Sbjct 294 GDGV---TCTDIDECALNTDNCDTNANCTNTIGSFTCTCKTGFTGDG--------VTCTD 342
Query 325 IDECNDASKNTCATSADGGSCKNTAGSYECSCLPGFQGDGHSCTDIDECATQG-VCGEHA 383
IDEC + N C T+A+ C NT GS+ C+C GF GDG SCTDIDECA C +A
Sbjct 343 IDECALNTDN-CDTNAN---CTNTIGSFTCTCKTGFTGDGVSCTDIDECALNTDNCDTNA 398
Query 384 TCENTAGSYNCTCEAGYTQQDGAVGCIDIDECAASTAVLPANATCVNTEGSYTFECVPGY 443
C NT GS+ CTC+ G+T V C DIDECA +T NA C NT GS+T C G+
Sbjct 399 NCTNTIGSFTCTCKTGFTGN--GVTCTDIDECALNTDNCDTNANCTNTIGSFTCTCKTGF 456
Query 444 RHTENGCTKIDFCSEK--GCNANASCKENDAGTEAICTCHSGYEGNGEGEEGCKNIDECS 501
CT ID C+ C+ NA+C N G CTC +G+ G+G C +IDEC+
Sbjct 457 TGDGVSCTDIDECALNTDNCDINANCT-NTIG-SFTCTCKTGFTGDG---VSCTDIDECA 511
Query 502 VG-EPCKDFGEGGVCVDSPGSFSCSCATGFIKRRSTCQDIDECLDGKMNTCAPVGGICTN 560
+ + CK C ++ GSF+C+C TGF TC DIDEC +TC CTN
Sbjct 512 LNTDNCK---INANCTNTIGSFTCTCKTGFTGNGVTCTDIDECA-LNTDTC-DTNANCTN 566
Query 561 TVGSFTCSCAAGFTGDGLTCEDIDECATAAHTCDPNATCVNTVGSFECGCKEGFSGDGHT 620
T+GSFTC+C GFTG+G+TC DIDECA CD NA C NT+G F C CK GF+G+G T
Sbjct 567 TIGSFTCTCKTGFTGNGVTCTDIDECALNTDNCDTNANCTNTIGRFTCTCKTGFTGNGVT 626
Query 621 CTDIDECADPNLNKCDTH---------------------------------KGICQNGTG 647
CTDIDECA N + CDT+ C N TG
Sbjct 627 CTDIDECA-LNTDNCDTNANCTNTIGSFTCACTTGFTDIDECTLGTHNCHANATCTNTTG 685
Query 648 SYTCGCRPGYSLAADGFTCDNVDECAAGTATCGERSFCVDTQGSYKCECKNGYRQSGEDC 707
S+TC C G+S DG +C ++DEC GT C + C +T GS+ C C G+ G C
Sbjct 686 SFTCICNTGFS--GDGVSCRDIDECTLGTHNCDTNATCTNTTGSFTCTCNTGFTGDGVSC 743
Query 708 VDVDECEADVHTCSEHATCTNTEGSHTCTCNEGYQGDGKKCEKTVGPCDNS--PCGNNAM 765
D+DEC H C +ATCTNT GS TC+CN G+ G+G C + PC N C +NA
Sbjct 744 TDIDECTLGTHNCHANATCTNTTGSFTCSCNTGFLGNGTWC---LYPCANQTHKCHSNAG 800
Query 766 CEATADSYNCTCKAGYEMKDGACVDIDECQSGTHNCDPHADCSNTDGSFTCTCGSGYTGV 825
C T+ ++ C CKAG+ C DIDEC GTHNC+ A C+NT GSFTC+C +GY+G
Sbjct 801 CVNTSKNFFCFCKAGFTGDGLTCTDIDECALGTHNCNTSATCNNTIGSFTCSCDTGYSGN 860
Query 826 GTLCEDVDECAGNHAGCDINAVCTNVPGSFTCECKSGFEGDGHECTE 872
G C D++ECA C NA CTN GSF C C +GF GDG CT+
Sbjct 861 GINCTDINECALRLHNCSYNANCTNTNGSFACSCNTGFTGDGVNCTD 907
Lambda K H
0.314 0.132 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 777828888878
Database: eggV2
Posted date: Dec 15, 2009 4:47 PM
Number of letters in database: 915,453,621
Number of sequences in database: 2,483,276
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40