bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: eggV2
2,483,276 sequences; 915,453,621 total letters
Query= Eten_0516_orf6
Length=623
Score E
Sequences producing significant alignments: (Bits) Value
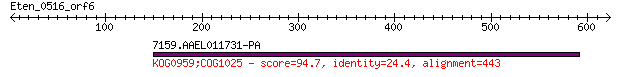
7159.AAEL011731-PA 94.7 8e-18
> 7159.AAEL011731-PA
Length=1003
Score = 94.7 bits (234), Expect = 8e-18, Method: Compositional matrix adjust.
Identities = 108/457 (23%), Positives = 200/457 (43%), Gaps = 38/457 (8%)
Query 149 VWWQGQGFSALPRVAVQLSGSIAKDKADLLSRTQGSVALAAIAEHLQEETADFQYCGITH 208
VW++ P+ + L +D L+ + + +HL E G+
Sbjct 538 VWFKQDVEFLKPKTLMNLDFCSPIVYSDPLNCNLTHLFVQLFKDHLNEYLYAAGLAGLRL 597
Query 209 SLAFKGTGFHMTFEGYTKTQ---LDKLMSHVASVLRDPSVVESERFERVKQKQIKLLADP 265
+A G ++ GY+ Q L+K++ + + ++ +RFE +K++ I+ L +
Sbjct 598 GVANTTYGVSVSIGGYSHKQHILLEKVLDDMFNF-----KIDEKRFEILKEQYIRNLKNY 652
Query 266 ATNMAFEHALQAAAILTRNDAFSRMDLLNALEQTTYDDSIAKLSEL-KNVHVDAFVMGNI 324
++HA+ A+L A+S+ +L++A E T D + EL +HV+ F+ GN+
Sbjct 653 QAEQPYQHAVYYLALLLTEQAWSKQELIDATELVTVDRLRTFIDELLSRMHVECFIYGNV 712
Query 325 DRDQSLLLTESFLEQAGFTPIEHDDAVESLAMEQKQTIEATLAN--------PIKGDKDH 376
+++++L ++ ++ T + V LA + E L N + K
Sbjct 713 NKEKALEMSSKVEDKLKKTDA---NVVPLLARQLMLKREYKLNNGENCLFEMTNEFHKSS 769
Query 377 ASLVQFQLGVPSIEERVNLAILSQFLNRRIYDSLRTEAQLGYIAGAKESQAASTALLQCF 436
+ + Q G+ + + V + +++Q L+ Y+ LRT+ QLGYI ++ ++
Sbjct 770 CAELYLQCGMQNDQANVYVDLVTQILSEPCYNQLRTKEQLGYIVFCGSRKSNGVQGIRVI 829
Query 437 VEGSKSHPDEVVKMIDAELAKAKDYISNMPEAELSRWKEAAHAKLTKVEANFSEDFKKSA 496
V+ S +HP V + I+ L DY+ NM E E R KEA A + S F K
Sbjct 830 VQ-SANHPAFVEERIEHFLNGMVDYLENMTEEEFKRHKEALAAMKLEKPKRLSSQFTKFL 888
Query 497 DEIFSHANCFSKRDLEVKYLDNDFSRKHLLRTFTK--LSDPSRRMVVKLIADLEPAKEVT 554
+EI F++ +EV + L+T TK + D + +VK A L + +
Sbjct 889 NEIALQQYHFNRAQVEVAF----------LQTLTKQQIVDYYKEYIVK-DASLRRSLSIH 937
Query 555 LIGEAKGQDAALLRRGDTAEEVVKAPTKLLSLEDKFV 591
++ A+G D + +V K T S + FV
Sbjct 938 VVSTAEGGAG----HKDASADVAKQSTDDASTQKDFV 970
Lambda K H
0.316 0.132 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 284937677493
Database: eggV2
Posted date: Dec 15, 2009 4:47 PM
Number of letters in database: 915,453,621
Number of sequences in database: 2,483,276
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40