bitscore colors: <40, 40-50 , 50-80, 80-200, >200
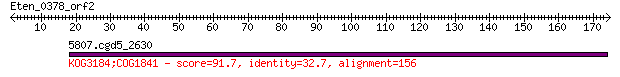
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: eggV2
2,483,276 sequences; 915,453,621 total letters
Query= Eten_0378_orf2
Length=174
Score E
Sequences producing significant alignments: (Bits) Value
5807.cgd5_2630 91.7 8e-18
> 5807.cgd5_2630
Length=268
Score = 91.7 bits (226), Expect = 8e-18, Method: Compositional matrix adjust.
Identities = 51/176 (28%), Positives = 88/176 (50%), Gaps = 20/176 (11%)
Query 18 VYCVVRNHRRGGCRETQEALKALGLPRPYSCTLIANDENATKILRKVSPFVFYGLPSAET 77
+ VVRN R G E+ L+ LGL +S ++++ + ++LR+V P+VFYG P+
Sbjct 92 IVLVVRNSRFCGSLESNRKLRDLGLVNKFSAVILSHTDETIRVLREVKPYVFYGFPNLNL 151
Query 78 IRRLFVTRARLT--------------SEEGVAPVALSDNMMVEEAFGSLGLFCVEDLVH- 122
++ L + T SE+ V L+DN +VE+ G +G+ C ED+V
Sbjct 152 LKTLVFKKGAFTHQNVQSKGQSRAKLSEKKQGKVLLTDNNIVEDRLGEIGVLCTEDIVTG 211
Query 123 -----EVVTCGPHFPQVMQRVGHFQLESLKQVEELEAKRMEYGYIRNKIDMKVAQL 173
E F + + FQL +LK+ E +AK+ E G++ I+ K++++
Sbjct 212 LWNGCETQQDKEIFEAITSNLAPFQLCNLKRAEGFDAKKFESGFLGKSINEKISKI 267
Lambda K H
0.322 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 34096907735
Database: eggV2
Posted date: Dec 15, 2009 4:47 PM
Number of letters in database: 915,453,621
Number of sequences in database: 2,483,276
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40