bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: eggV2
2,483,276 sequences; 915,453,621 total letters
Query= Eten_0355_orf1
Length=472
Score E
Sequences producing significant alignments: (Bits) Value
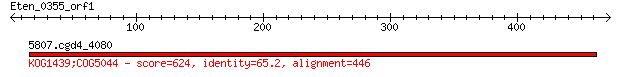
5807.cgd4_4080 624 2e-177
> 5807.cgd4_4080
Length=473
Score = 624 bits (1610), Expect = 2e-177, Method: Compositional matrix adjust.
Identities = 291/449 (64%), Positives = 365/449 (81%), Gaps = 4/449 (0%)
Query 16 MDEEYDVVVCGTGLKECILSGLLSSAGKKVLHLDRNNYYGGECASLSLTNLYNKFKPGTS 75
MDE+YDV++CGTGL ECI+SGLLS++GKKVLH+DRN+YYGGE ASL+LT LY KF+PGTS
Sbjct 14 MDEKYDVLICGTGLTECIISGLLSTSGKKVLHIDRNSYYGGEAASLNLTTLYQKFRPGTS 73
Query 76 PPPEYGSNREWNVDLIPKFVMACGKLVKILLKTKVTRYLEWQVVEGTSVYQYQKAGYFSG 135
PP YG+NR+WNVDLIPKFVMA G LVKILLKTKVTRYLEWQV+EGT VYQ+QK G
Sbjct 74 PPANYGANRDWNVDLIPKFVMASGDLVKILLKTKVTRYLEWQVIEGTYVYQFQKGGLLFN 133
Query 136 EKYIHKVPATESEALMSPLMPLLEKNRCRSFFAYCAQRQADKPETWKGFDPKRHSMEQVY 195
K+IHKVPATE EAL SPL+ ++EKNRCRSFF++ A D GF+ +++M+ +Y
Sbjct 134 PKFIHKVPATEMEALKSPLLGIMEKNRCRSFFSFVANWSDDDVSKQMGFNRDKNTMKDIY 193
Query 196 KYFGLETNTIDFVGHAVALYTCDDYLKQPMQETMERIKLYMYSVSRYGRSPFIYPVYGLG 255
+FGL + TIDFVGHA+ALYT DDY+ +P ET+++I+LYM S+SRYG+SPFIYPVYGLG
Sbjct 194 DHFGLSSTTIDFVGHALALYTNDDYINKPCGETLDKIRLYMMSLSRYGKSPFIYPVYGLG 253
Query 256 GLPEGFSRLCAVNGGTYMLNAPVDSFVLDEKGKVCGVKTKDGQVARCKMVVCDPSYIAEQ 315
GLPEGFSRLCA++GGT+MLN ++ F+ DE+GKV GV T G+ A CKMV+CDPSY+ +
Sbjct 254 GLPEGFSRLCAIHGGTFMLNTNIEKFLYDEQGKVSGVVTSQGK-AECKMVICDPSYVLNE 312
Query 316 FPS--RLKCTGRVIRCICILGAPVPSTNGSTSCQIIIPQRQLNRHNDIYIMVVSSPHGIC 373
S ++KC G+V+RCICIL +P+ TN +SCQIIIPQ +L R NDIY+M+VS HG+
Sbjct 313 NDSKPKVKCIGKVLRCICILNSPINDTNDVSSCQIIIPQNELGRKNDIYVMMVSCTHGVA 372
Query 374 LKGKTIAIVSTTVETDDPEKELAPALKLLGN-IEEKFVAVSDLLERTDNGRDSNIFVSNS 432
LKGK IAI+STTVET++P E+ PA+KLLGN IEE+F SDL E D+G+D N+FVS S
Sbjct 373 LKGKYIAIISTTVETENPLLEINPAIKLLGNSIEEQFFYTSDLYEPIDSGKDDNVFVSKS 432
Query 433 FDATSHFESATQDVLRMWEDMTGEPLDLS 461
DA+SHFES TQDVLR+W+++TGE LDL+
Sbjct 433 CDASSHFESLTQDVLRLWKNITGEDLDLT 461
Lambda K H
0.318 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 199804804260
Database: eggV2
Posted date: Dec 15, 2009 4:47 PM
Number of letters in database: 915,453,621
Number of sequences in database: 2,483,276
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40