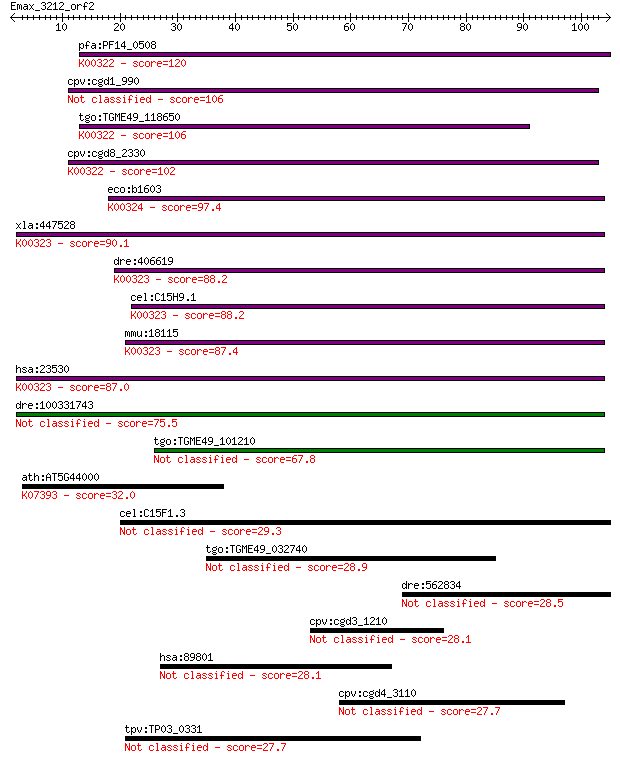
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_865
164,496 sequences; 82,071,388 total letters
Query= Emax_3212_orf2
Length=104
Score E
Sequences producing significant alignments: (Bits) Value
pfa:PF14_0508 pyridine nucleotide transhydrogenase, putative (... 120 1e-27
cpv:cgd1_990 pyridine nucleotide/ NAD(P) transhydrogenase alph... 106 1e-23
tgo:TGME49_118650 transhydrogenase, putative (EC:1.6.1.2); K00... 106 2e-23
cpv:cgd8_2330 pyridine nucleotide/ NAD(P) transhydrogenase alp... 102 4e-22
eco:b1603 pntA, ECK1598, JW1595; pyridine nucleotide transhydr... 97.4 8e-21
xla:447528 nnt, MGC83563; nicotinamide nucleotide transhydroge... 90.1 1e-18
dre:406619 nnt, wu:fa20d10, wu:fc86a04, zgc:76979; nicotinamid... 88.2 5e-18
cel:C15H9.1 nnt-1; Nicotinamide Nucleotide Transhydrogenase fa... 88.2 6e-18
mmu:18115 Nnt, 4930423F13Rik, AI323702, BB168308; nicotinamide... 87.4 1e-17
hsa:23530 NNT, MGC126502, MGC126503; nicotinamide nucleotide t... 87.0 1e-17
dre:100331743 Nicotinamide Nucleotide Transhydrogenase family ... 75.5 3e-14
tgo:TGME49_101210 NAD(P) transhydrogenase, alpha subunit, puta... 67.8 7e-12
ath:AT5G44000 glutathione S-transferase C-terminal domain-cont... 32.0 0.53
cel:C15F1.3 tra-2; TRAnsformer : XX animals transformed into m... 29.3 2.9
tgo:TGME49_032740 hypothetical protein 28.9 4.4
dre:562834 slc12a10.2, si:dkey-114c3.2; solute carrier family ... 28.5 6.0
cpv:cgd3_1210 hypothetical protein 28.1 6.2
hsa:89801 PPP1R3F, HB2E; protein phosphatase 1, regulatory (in... 28.1 6.7
cpv:cgd4_3110 10 transmembrane domain protein, possible transl... 27.7 8.1
tpv:TP03_0331 hypothetical protein 27.7 8.8
> pfa:PF14_0508 pyridine nucleotide transhydrogenase, putative
(EC:1.6.1.2); K00322 NAD(P) transhydrogenase [EC:1.6.1.1]
Length=1176
Score = 120 bits (300), Expect = 1e-27, Method: Composition-based stats.
Identities = 57/92 (61%), Positives = 72/92 (78%), Gaps = 0/92 (0%)
Query 13 LDPMELKHLTLLALSLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIGCMLEYGTALISG 72
L +L+ L L LS IVGYYCVW+VTPALHTPLMS+TNALSGVI+IG M+EYG
Sbjct 1085 LSQSDLQSLFLFTLSTIVGYYCVWSVTPALHTPLMSMTNALSGVIIIGSMIEYGNGYKDL 1144
Query 73 FTLLAIIGTFLASVNIAGGFFVTHRMLKMFQI 104
++L+++ TFL+SVN++GGF+VT RML MF I
Sbjct 1145 SSILSMLATFLSSVNLSGGFYVTKRMLDMFFI 1176
> cpv:cgd1_990 pyridine nucleotide/ NAD(P) transhydrogenase alpha
plus beta subunits, duplicated gene, 12 transmembrane domain
(EC:1.6.1.2)
Length=1147
Score = 106 bits (265), Expect = 1e-23, Method: Composition-based stats.
Identities = 46/92 (50%), Positives = 69/92 (75%), Gaps = 0/92 (0%)
Query 11 FVLDPMELKHLTLLALSLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIGCMLEYGTALI 70
+ ++++++ +S ++GYYCVW V P LHTPLMSVTNALSGVI+IG M++YG +
Sbjct 1054 LTMTTIQIQNIFSFIISTMLGYYCVWDVDPKLHTPLMSVTNALSGVIIIGSMMQYGNQTV 1113
Query 71 SGFTLLAIIGTFLASVNIAGGFFVTHRMLKMF 102
+ TL+++ TFLAS+N GGF+VT++ML +F
Sbjct 1114 TYTTLMSMFSTFLASINTFGGFYVTNKMLTLF 1145
> tgo:TGME49_118650 transhydrogenase, putative (EC:1.6.1.2); K00322
NAD(P) transhydrogenase [EC:1.6.1.1]
Length=1013
Score = 106 bits (265), Expect = 2e-23, Method: Compositional matrix adjust.
Identities = 53/78 (67%), Positives = 62/78 (79%), Gaps = 0/78 (0%)
Query 13 LDPMELKHLTLLALSLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIGCMLEYGTALISG 72
L +EL+ TLLALSLIVGYY VW+VTPALHTPLMSVTNALSGVI+IG MLEYG + S
Sbjct 927 LKALELQQFTLLALSLIVGYYSVWSVTPALHTPLMSVTNALSGVIIIGSMLEYGPSATSA 986
Query 73 FTLLAIIGTFLASVNIAG 90
+ A+ TF +S+NIAG
Sbjct 987 SAICALCATFFSSLNIAG 1004
> cpv:cgd8_2330 pyridine nucleotide/ NAD(P) transhydrogenase alpha
plus beta subunits, duplicated gene, possible signal peptide
plus 12 transmembrane regions ; K00322 NAD(P) transhydrogenase
[EC:1.6.1.1]
Length=1143
Score = 102 bits (253), Expect = 4e-22, Method: Compositional matrix adjust.
Identities = 50/96 (52%), Positives = 70/96 (72%), Gaps = 4/96 (4%)
Query 11 FVLDPMELKHLTLLALSLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIGCMLEYGTALI 70
+++D L ++ + +LS+IVGYYC+W VTP+LHTPLMSVTNALSG+I+IG MLE G ++
Sbjct 1026 YIIDHDTLGNILVFSLSVIVGYYCIWNVTPSLHTPLMSVTNALSGIIIIGAMLECGPVIL 1085
Query 71 ----SGFTLLAIIGTFLASVNIAGGFFVTHRMLKMF 102
++ L + L+S+NI GGF+VT RML MF
Sbjct 1086 FTDFQVYSFLLFLAMLLSSINIIGGFYVTTRMLYMF 1121
> eco:b1603 pntA, ECK1598, JW1595; pyridine nucleotide transhydrogenase,
alpha subunit (EC:1.6.1.2); K00324 NAD(P) transhydrogenase
subunit alpha [EC:1.6.1.2]
Length=510
Score = 97.4 bits (241), Expect = 8e-21, Method: Composition-based stats.
Identities = 48/86 (55%), Positives = 62/86 (72%), Gaps = 2/86 (2%)
Query 18 LKHLTLLALSLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIGCMLEYGTALISGFTLLA 77
L H T+ AL+ +VGYY VW V+ ALHTPLMSVTNA+SG+IV+G +L+ G + L+
Sbjct 425 LGHFTVFALACVVGYYVVWNVSHALHTPLMSVTNAISGIIVVGALLQIGQG--GWVSFLS 482
Query 78 IIGTFLASVNIAGGFFVTHRMLKMFQ 103
I +AS+NI GGF VT RMLKMF+
Sbjct 483 FIAVLIASINIFGGFTVTQRMLKMFR 508
> xla:447528 nnt, MGC83563; nicotinamide nucleotide transhydrogenase
(EC:1.6.1.2); K00323 NAD(P) transhydrogenase [EC:1.6.1.2]
Length=1086
Score = 90.1 bits (222), Expect = 1e-18, Method: Compositional matrix adjust.
Identities = 51/104 (49%), Positives = 66/104 (63%), Gaps = 6/104 (5%)
Query 2 LSYGAPSPKFVLDPMELKHLTLLALSLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIGC 61
LS G SP M +T L+ IVGY+ VW VTPALH+PLMSVTNA+SG+ +G
Sbjct 487 LSLGIASPHSAFTQM----VTTFGLAGIVGYHTVWGVTPALHSPLMSVTNAISGLTAVGG 542
Query 62 MLEYGTALISGFT--LLAIIGTFLASVNIAGGFFVTHRMLKMFQ 103
+ G + T LLA++ F++S+NIAGGF VT RML MF+
Sbjct 543 LALMGGGYLPTNTHELLAVLAAFVSSINIAGGFLVTQRMLDMFK 586
> dre:406619 nnt, wu:fa20d10, wu:fc86a04, zgc:76979; nicotinamide
nucleotide transhydrogenase (EC:1.6.1.2); K00323 NAD(P)
transhydrogenase [EC:1.6.1.2]
Length=1079
Score = 88.2 bits (217), Expect = 5e-18, Method: Compositional matrix adjust.
Identities = 45/87 (51%), Positives = 60/87 (68%), Gaps = 2/87 (2%)
Query 19 KHLTLLALSLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIGCMLEYGTALISGFT--LL 76
+ +T L+ IVGY+ VW VTPALH+PLMSVTNA+SG+ +G + G + T L
Sbjct 496 QMVTTFGLAGIVGYHTVWGVTPALHSPLMSVTNAISGLTAVGGLSLMGGGYLPSSTAETL 555
Query 77 AIIGTFLASVNIAGGFFVTHRMLKMFQ 103
A++ F++SVNIAGGF VT RML MF+
Sbjct 556 AVLAAFISSVNIAGGFLVTQRMLDMFK 582
> cel:C15H9.1 nnt-1; Nicotinamide Nucleotide Transhydrogenase
family member (nnt-1); K00323 NAD(P) transhydrogenase [EC:1.6.1.2]
Length=1041
Score = 88.2 bits (217), Expect = 6e-18, Method: Compositional matrix adjust.
Identities = 44/84 (52%), Positives = 58/84 (69%), Gaps = 2/84 (2%)
Query 22 TLLALSLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIG--CMLEYGTALISGFTLLAII 79
T AL+ +VGY+ VW VTPALH+PLMSVTNA+SG G C++ G + +A++
Sbjct 457 TTFALAGLVGYHTVWGVTPALHSPLMSVTNAISGTTAAGALCLMGGGLMPQNSAQTMALL 516
Query 80 GTFLASVNIAGGFFVTHRMLKMFQ 103
TF++SVNI GGF VT RML MF+
Sbjct 517 ATFISSVNIGGGFLVTKRMLDMFK 540
> mmu:18115 Nnt, 4930423F13Rik, AI323702, BB168308; nicotinamide
nucleotide transhydrogenase (EC:1.6.1.2); K00323 NAD(P) transhydrogenase
[EC:1.6.1.2]
Length=1086
Score = 87.4 bits (215), Expect = 1e-17, Method: Compositional matrix adjust.
Identities = 47/90 (52%), Positives = 60/90 (66%), Gaps = 12/90 (13%)
Query 21 LTLLALSLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIGCMLEYGTALISGF------- 73
+T L+ I+GY+ VW VTPALH+PLMSVTNA+SG+ +G G AL+ G
Sbjct 502 VTTFGLAGIIGYHTVWGVTPALHSPLMSVTNAISGLTAVG-----GLALMGGHFYPSTTS 556
Query 74 TLLAIIGTFLASVNIAGGFFVTHRMLKMFQ 103
LA + TF++SVNIAGGF VT RML MF+
Sbjct 557 QSLAALATFISSVNIAGGFLVTQRMLDMFK 586
> hsa:23530 NNT, MGC126502, MGC126503; nicotinamide nucleotide
transhydrogenase (EC:1.6.1.2); K00323 NAD(P) transhydrogenase
[EC:1.6.1.2]
Length=1086
Score = 87.0 bits (214), Expect = 1e-17, Method: Compositional matrix adjust.
Identities = 50/104 (48%), Positives = 63/104 (60%), Gaps = 6/104 (5%)
Query 2 LSYGAPSPKFVLDPMELKHLTLLALSLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIGC 61
L G +P M +T L+ IVGY+ VW VTPALH+PLMSVTNA+SG+ +G
Sbjct 487 LGLGIAAPNLAFSQM----VTTFGLAGIVGYHTVWGVTPALHSPLMSVTNAISGLTAVGG 542
Query 62 MLEYGTALISGFTL--LAIIGTFLASVNIAGGFFVTHRMLKMFQ 103
+ G L T LA + F++SVNIAGGF VT RML MF+
Sbjct 543 LALMGGHLYPSTTSQGLAALAAFISSVNIAGGFLVTQRMLDMFK 586
> dre:100331743 Nicotinamide Nucleotide Transhydrogenase family
member (nnt-1)-like
Length=518
Score = 75.5 bits (184), Expect = 3e-14, Method: Compositional matrix adjust.
Identities = 47/104 (45%), Positives = 64/104 (61%), Gaps = 6/104 (5%)
Query 2 LSYGAPSPKFVLDPMELKHLTLLALSLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIGC 61
L G SP M +T L+ IVGY+ VW VTPALH+PLMSVTNA+SG+ +G
Sbjct 187 LGLGIASPNAAFTQM----VTTFGLAGIVGYHTVWGVTPALHSPLMSVTNAISGLTAVGG 242
Query 62 MLEYGTALISGF--TLLAIIGTFLASVNIAGGFFVTHRMLKMFQ 103
++ G L LA++ F++S+NIAGGF +T +ML MF+
Sbjct 243 LVLMGGGLTPSSLPESLALLAAFVSSINIAGGFLITQKMLDMFK 286
> tgo:TGME49_101210 NAD(P) transhydrogenase, alpha subunit, putative
(EC:1.6.1.2)
Length=1165
Score = 67.8 bits (164), Expect = 7e-12, Method: Compositional matrix adjust.
Identities = 44/108 (40%), Positives = 54/108 (50%), Gaps = 34/108 (31%)
Query 26 LSLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIGCML---------------------- 63
L+ VGY VW V PALHTPLMSVTNA+SG +++G ML
Sbjct 1057 LACFVGYLLVWNVAPALHTPLMSVTNAISGTVLVGGMLGISARAYQTLDKESICPHNHLG 1116
Query 64 --------EYGTALISGFTLLAIIGTFLASVNIAGGFFVTHRMLKMFQ 103
GTA I+ L I +AS+N+ GGF VT RML MF+
Sbjct 1117 PLEPFYCHSRGTAAIT----LNAIAIAVASMNVFGGFAVTQRMLNMFR 1160
> ath:AT5G44000 glutathione S-transferase C-terminal domain-containing
protein; K07393 putative glutathione S-transferase
Length=399
Score = 32.0 bits (71), Expect = 0.53, Method: Composition-based stats.
Identities = 15/35 (42%), Positives = 17/35 (48%), Gaps = 0/35 (0%)
Query 3 SYGAPSPKFVLDPMELKHLTLLALSLIVGYYCVWA 37
SY P+ KF LDP + L L VG C WA
Sbjct 83 SYTRPTSKFRLDPTQFTSAASSELHLYVGLPCPWA 117
> cel:C15F1.3 tra-2; TRAnsformer : XX animals transformed into
males family member (tra-2)
Length=1475
Score = 29.3 bits (64), Expect = 2.9, Method: Composition-based stats.
Identities = 27/100 (27%), Positives = 42/100 (42%), Gaps = 18/100 (18%)
Query 20 HLTLLALSLIVGYYCVWAVTPAL--------------HTP-LMSVTNALSGVIVIGCMLE 64
H L+L G+ W AL TP L S L ++ + +
Sbjct 977 HQIYTNLALFAGFLAAWDPFCALLRYRRRILYKSETRRTPELASKRRVLLPIVATADIAQ 1036
Query 65 YGTALISGFTLLAIIGTFLASVNIAGGFFVTHRMLKMFQI 104
+ LI+ F++LAII + + +NI FFV +L + QI
Sbjct 1037 FFVLLITAFSILAIICSIVPELNI---FFVPTVILIVIQI 1073
> tgo:TGME49_032740 hypothetical protein
Length=380
Score = 28.9 bits (63), Expect = 4.4, Method: Composition-based stats.
Identities = 15/50 (30%), Positives = 26/50 (52%), Gaps = 0/50 (0%)
Query 35 VWAVTPALHTPLMSVTNALSGVIVIGCMLEYGTALISGFTLLAIIGTFLA 84
+W + P P+++ N L V+ M E+ TA +S + +IG+F A
Sbjct 200 LWILRPLPCLPMLTGANCLGSVLYPITMSEFTTADVSDSVMNGVIGSFPA 249
> dre:562834 slc12a10.2, si:dkey-114c3.2; solute carrier family
12 (sodium/potassium/chloride transporters), member 10.2
Length=998
Score = 28.5 bits (62), Expect = 6.0, Method: Composition-based stats.
Identities = 15/36 (41%), Positives = 25/36 (69%), Gaps = 2/36 (5%)
Query 69 LISGFTLLAIIGTFLASVNIAGGFFVTHRMLKMFQI 104
++SG+ +L IGTF A+++ A GF V+ K+FQ+
Sbjct 419 VVSGWGILITIGTFAATLSSALGFLVSAP--KVFQL 452
> cpv:cgd3_1210 hypothetical protein
Length=464
Score = 28.1 bits (61), Expect = 6.2, Method: Composition-based stats.
Identities = 12/23 (52%), Positives = 17/23 (73%), Gaps = 0/23 (0%)
Query 53 LSGVIVIGCMLEYGTALISGFTL 75
LS +I+I +L +G+AL SG TL
Sbjct 9 LSALIIITLLLSFGSALFSGLTL 31
> hsa:89801 PPP1R3F, HB2E; protein phosphatase 1, regulatory (inhibitor)
subunit 3F
Length=453
Score = 28.1 bits (61), Expect = 6.7, Method: Composition-based stats.
Identities = 14/40 (35%), Positives = 20/40 (50%), Gaps = 0/40 (0%)
Query 27 SLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIGCMLEYG 66
+L V CV A+ P L PL L+G++V+ L G
Sbjct 398 TLAVPAECVCALPPQLRGPLTQTLGVLAGLVVVPVALNSG 437
> cpv:cgd4_3110 10 transmembrane domain protein, possible translocator
Length=736
Score = 27.7 bits (60), Expect = 8.1, Method: Composition-based stats.
Identities = 13/42 (30%), Positives = 22/42 (52%), Gaps = 3/42 (7%)
Query 58 VIGCMLEYGTALISGFT---LLAIIGTFLASVNIAGGFFVTH 96
++ +L Y T L+ G+T ++G F + AGGF + H
Sbjct 410 LLSMLLNYFTFLVVGYTSPVTFNVLGMFKSCAQTAGGFIIFH 451
> tpv:TP03_0331 hypothetical protein
Length=806
Score = 27.7 bits (60), Expect = 8.8, Method: Composition-based stats.
Identities = 13/51 (25%), Positives = 33/51 (64%), Gaps = 1/51 (1%)
Query 21 LTLLALSLIVGYYCVWAVTPALHTPLMSVTNALSGVIVIGCMLEYGTALIS 71
++++ +++++GY ++ V A +T +T+ L +I++G +++G LIS
Sbjct 462 ISMVTIAILMGYIVLFFVPYAFYTITSPMTHILE-IILLGLFVDFGNTLIS 511
Lambda K H
0.329 0.142 0.445
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2022291452
Database: egene_temp_file_orthology_annotation_similarity_blast_database_865
Posted date: Sep 17, 2011 11:19 AM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40