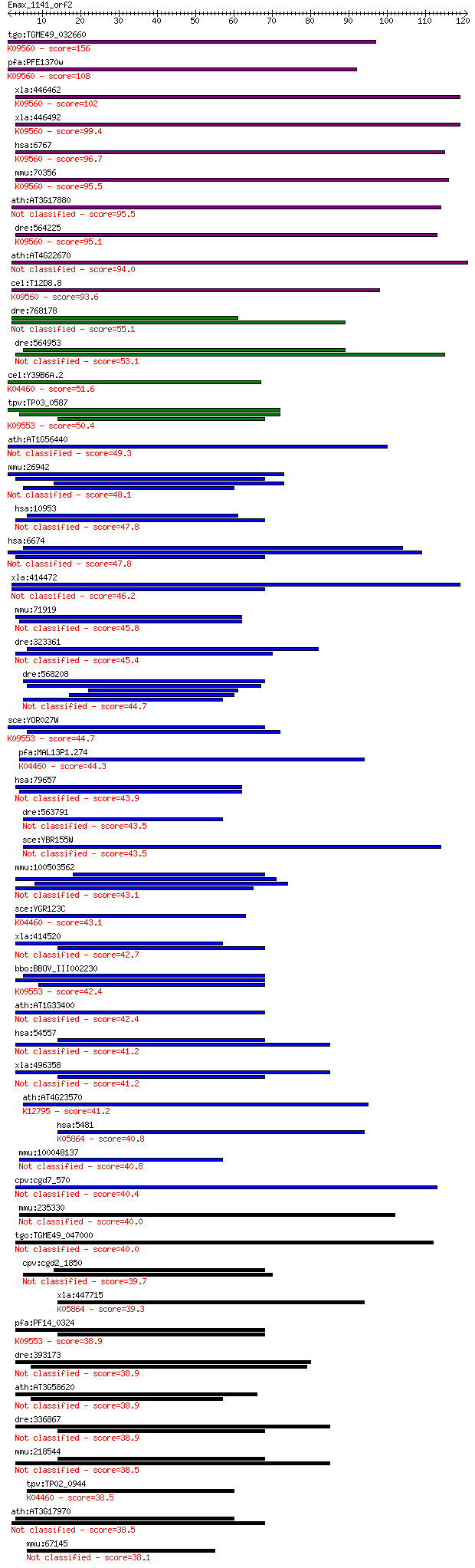
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_865
164,496 sequences; 82,071,388 total letters
Query= Emax_1141_orf2
Length=120
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_032660 58 kDa phosphoprotein, putative ; K09560 sup... 156 2e-38
pfa:PFE1370w hsp70 interacting protein, putative; K09560 suppr... 108 3e-24
xla:446462 st13, MGC78939; suppression of tumorigenicity 13 (c... 102 3e-22
xla:446492 MGC79131 protein; K09560 suppressor of tumorigenici... 99.4 3e-21
hsa:6767 ST13, AAG2, FAM10A1, FAM10A4, FLJ27260, HIP, HOP, HSP... 96.7 2e-20
mmu:70356 St13, 1110007I03Rik, 3110002K08Rik, AW555194, HIP, H... 95.5 3e-20
ath:AT3G17880 ATTDX; ATTDX (TETRATICOPEPTIDE DOMAIN-CONTAINING... 95.5 3e-20
dre:564225 st13, MGC73267, MGC77089, wu:fd15g02, zgc:73267; su... 95.1 5e-20
ath:AT4G22670 AtHip1; AtHip1 (Arabidopsis thaliana Hsp70-inter... 94.0 1e-19
cel:T12D8.8 hypothetical protein; K09560 suppressor of tumorig... 93.6 1e-19
dre:768178 zgc:153288 55.1 5e-08
dre:564953 spag1, MGC162178, cb1089, wu:fj78g10; sperm associa... 53.1 2e-07
cel:Y39B6A.2 pph-5; Protein PHosphatase family member (pph-5);... 51.6 6e-07
tpv:TP03_0587 hypothetical protein; K09553 stress-induced-phos... 50.4 1e-06
ath:AT1G56440 serine/threonine protein phosphatase-related 49.3 3e-06
mmu:26942 Spag1, tpis; sperm associated antigen 1 48.1
hsa:10953 TOMM34, HTOM34P, TOM34, URCC3; translocase of outer ... 47.8 8e-06
hsa:6674 SPAG1, FLJ32920, HSD-3.8, SP75, TPIS; sperm associate... 47.8 8e-06
xla:414472 rpap3, MGC81126; RNA polymerase II associated prote... 46.2 2e-05
mmu:71919 Rpap3, 2310042P20Rik, D15Ertd682e; RNA polymerase II... 45.8 4e-05
dre:323361 tomm34, wu:fb96b08, zgc:56645; translocase of outer... 45.4 5e-05
dre:568208 ttc6; tetratricopeptide repeat domain 6 44.7
sce:YOR027W STI1; Hsp90 cochaperone, interacts with the Ssa gr... 44.7 7e-05
pfa:MAL13P1.274 PfPP5; serine/threonine protein phosphatase (E... 44.3 1e-04
hsa:79657 RPAP3, FLJ21908; RNA polymerase II associated protein 3 43.9
dre:563791 ttc12; tetratricopeptide repeat domain 12 43.5
sce:YBR155W CNS1; Cns1p 43.5 2e-04
mmu:100503562 hypothetical protein LOC100503562 43.1 2e-04
sce:YGR123C PPT1; Ppt1p (EC:3.1.3.16); K04460 protein phosphat... 43.1 2e-04
xla:414520 hypothetical protein MGC81394 42.7 3e-04
bbo:BBOV_III002230 17.m07215; tetratricopeptide repeat domain ... 42.4 4e-04
ath:AT1G33400 tetratricopeptide repeat (TPR)-containing protein 42.4 4e-04
hsa:54557 SGTB, FLJ39002, SGT2; small glutamine-rich tetratric... 41.2 7e-04
xla:496358 sgta; small glutamine-rich tetratricopeptide repeat... 41.2 8e-04
ath:AT4G23570 SGT1A; SGT1A; protein binding; K12795 suppressor... 41.2 8e-04
hsa:5481 PPID, CYP-40, CYPD, MGC33096; peptidylprolyl isomeras... 40.8 0.001
mmu:100048137 tetratricopeptide repeat protein 12-like 40.8 0.001
cpv:cgd7_570 protein with 2 TPR domains 40.4 0.001
mmu:235330 Ttc12, E330017O07Rik; tetratricopeptide repeat doma... 40.0 0.002
tgo:TGME49_047000 TPR domain-containing protein (EC:3.4.21.72) 40.0 0.002
cpv:cgd2_1850 stress-induced protein sti1-like protein 39.7 0.002
xla:447715 ppid, MGC81732, cyp-40, cypd; peptidylprolyl isomer... 39.3 0.003
pfa:PF14_0324 Hsp70/Hsp90 organizing protein, putative; K09553... 38.9 0.004
dre:393173 MGC56178; zgc:56178 38.9 0.004
ath:AT3G58620 TTL4; TTL4 (Tetratricopetide-repeat Thioredoxin-... 38.9 0.004
dre:336867 fk20d10, wu:fa05b08, wu:fc52b05, wu:fk20d10, zgc:77... 38.9 0.004
mmu:218544 Sgtb, C630001O05Rik, MGC27660; small glutamine-rich... 38.5 0.005
tpv:TP02_0944 serine/threonine protein phosphatase; K04460 pro... 38.5 0.005
ath:AT3G17970 atToc64-III (Arabidopsis thaliana translocon at ... 38.5 0.005
mmu:67145 Tomm34, 2610100K07Rik, TOM34, Tomm34a, Tomm34b; tran... 38.1 0.006
> tgo:TGME49_032660 58 kDa phosphoprotein, putative ; K09560 suppressor
of tumorigenicity protein 13
Length=425
Score = 156 bits (394), Expect = 2e-38, Method: Composition-based stats.
Identities = 71/96 (73%), Positives = 85/96 (88%), Gaps = 0/96 (0%)
Query 1 PTALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDI 60
PTALLYTRRAD+LLK KR A IRDCDEALKLNPD+ARAY+IRG A+R LG W++AHSD+
Sbjct 150 PTALLYTRRADVLLKLKRPVACIRDCDEALKLNPDSARAYKIRGKANRLLGKWREAHSDL 209
Query 61 EMGQKIDYDENIWDIQKLVEQKFKIIEEHERSVQRR 96
+MGQKIDYDE +WD+QKLV++KFK IEEHER + R+
Sbjct 210 DMGQKIDYDEGLWDMQKLVDEKFKKIEEHERKIVRK 245
> pfa:PFE1370w hsp70 interacting protein, putative; K09560 suppressor
of tumorigenicity protein 13
Length=458
Score = 108 bits (271), Expect = 3e-24, Method: Composition-based stats.
Identities = 53/91 (58%), Positives = 72/91 (79%), Gaps = 0/91 (0%)
Query 1 PTALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDI 60
P+A++YT+RA +LL KR A IRDC EAL LN D+A AY++R A+R+LG W+ AH+DI
Sbjct 162 PSAMIYTKRASVLLSLKRPKACIRDCTEALNLNIDSANAYKVRAKAYRHLGKWECAHADI 221
Query 61 EMGQKIDYDENIWDIQKLVEQKFKIIEEHER 91
E GQKIDYDE++W++QKL+E+K+K I E R
Sbjct 222 EQGQKIDYDEDLWEMQKLIEEKYKKIYEKRR 252
> xla:446462 st13, MGC78939; suppression of tumorigenicity 13
(colon carcinoma) (Hsp70 interacting protein); K09560 suppressor
of tumorigenicity protein 13
Length=379
Score = 102 bits (254), Expect = 3e-22, Method: Compositional matrix adjust.
Identities = 52/119 (43%), Positives = 80/119 (67%), Gaps = 3/119 (2%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
A+LY +RA + +K ++ AAIRDCD A+ +NPD+A+ Y+ RG AHR LGHW+ + D+ +
Sbjct 146 AILYAKRASVYVKLQKPNAAIRDCDRAIAINPDSAQPYKWRGKAHRLLGHWEDSAHDLAI 205
Query 63 GQKIDYDENIWDIQKLVEQKFKIIEEHERSVQRRKEE---EERKQRERRAREQRANAQR 118
K+DYDE+ + K V+ + I EH R +R++EE ERK+R ++A+E+ AQR
Sbjct 206 ACKLDYDEDASTLLKEVQPRANKIAEHRRKYERKREEREINERKERLKKAKEENERAQR 264
> xla:446492 MGC79131 protein; K09560 suppressor of tumorigenicity
protein 13
Length=376
Score = 99.4 bits (246), Expect = 3e-21, Method: Compositional matrix adjust.
Identities = 50/119 (42%), Positives = 81/119 (68%), Gaps = 3/119 (2%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
A+LY +RA + ++ ++ AAIRDCD A+ +NPD+A+ Y+ RG AHR LGHW+ + D+ +
Sbjct 146 AILYAKRASVYVQLQKPNAAIRDCDRAIAINPDSAQPYKWRGKAHRLLGHWEDSAHDLAI 205
Query 63 GQKIDYDENIWDIQKLVEQKFKIIEEHERSVQRRKEEEE---RKQRERRAREQRANAQR 118
K+DYDE+ + K V+ + I EH R +R++EE+E +K+R ++A+E+ AQR
Sbjct 206 ACKLDYDEDASAMLKEVQPRANKIAEHRRKHERKREEKEINDKKERLKKAKEENERAQR 264
> hsa:6767 ST13, AAG2, FAM10A1, FAM10A4, FLJ27260, HIP, HOP, HSPABP,
HSPABP1, MGC129952, P48, PRO0786, SNC6; suppression of
tumorigenicity 13 (colon carcinoma) (Hsp70 interacting protein);
K09560 suppressor of tumorigenicity protein 13
Length=369
Score = 96.7 bits (239), Expect = 2e-20, Method: Compositional matrix adjust.
Identities = 51/128 (39%), Positives = 76/128 (59%), Gaps = 16/128 (12%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
A+LY +RA + +K ++ AAIRDCD A+++NPD+A+ Y+ RG AHR LGHW++A D+ +
Sbjct 148 AILYAKRASVFVKLQKPNAAIRDCDRAIEINPDSAQPYKWRGKAHRLLGHWEEAAHDLAL 207
Query 63 GQKIDYDENIWDIQKLVEQKFKIIEEH----------------ERSVQRRKEEEERKQRE 106
K+DYDE+ + K V+ + + I EH V++ +EE ER QRE
Sbjct 208 ACKLDYDEDASAMLKEVQPRAQKIAEHRRKYERKREEREIKERIERVKKAREEHERAQRE 267
Query 107 RRAREQRA 114
AR Q
Sbjct 268 EEARRQSG 275
> mmu:70356 St13, 1110007I03Rik, 3110002K08Rik, AW555194, HIP,
HOP, HSPABP, HSPABP1, PRO0786, SNC6, p48; suppression of tumorigenicity
13; K09560 suppressor of tumorigenicity protein
13
Length=371
Score = 95.5 bits (236), Expect = 3e-20, Method: Compositional matrix adjust.
Identities = 50/129 (38%), Positives = 77/129 (59%), Gaps = 16/129 (12%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
A+LY +RA + +K ++ AAIRDCD A+++NPD+A+ Y+ RG AHR LGHW++A D+ +
Sbjct 147 AILYAKRASVFVKLQKPNAAIRDCDRAIEINPDSAQPYKWRGKAHRLLGHWEEAAHDLAL 206
Query 63 GQKIDYDENIWDIQKLVEQKFKIIEEH----------------ERSVQRRKEEEERKQRE 106
K+DYDE+ + + V+ + + I EH V++ +EE ER QRE
Sbjct 207 ACKLDYDEDASAMLREVQPRAQKIAEHRRKYERKREEREIKERIERVKKAREEHERAQRE 266
Query 107 RRAREQRAN 115
AR Q +
Sbjct 267 EEARRQSGS 275
> ath:AT3G17880 ATTDX; ATTDX (TETRATICOPEPTIDE DOMAIN-CONTAINING
THIOREDOXIN); oxidoreductase, acting on sulfur group of donors,
disulfide as acceptor / protein binding
Length=373
Score = 95.5 bits (236), Expect = 3e-20, Method: Compositional matrix adjust.
Identities = 50/112 (44%), Positives = 73/112 (65%), Gaps = 0/112 (0%)
Query 2 TALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIE 61
+A+LY RA + LK K+ AAIRD + AL+ N D+A+ Y+ RG A LG W++A +D+
Sbjct 138 SAILYATRASVFLKVKKPNAAIRDANVALQFNSDSAKGYKSRGMAKAMLGQWEEAAADLH 197
Query 62 MGQKIDYDENIWDIQKLVEQKFKIIEEHERSVQRRKEEEERKQRERRAREQR 113
+ K+DYDE I + K VE K IEEH R QR ++E+E ++ ER R+Q+
Sbjct 198 VASKLDYDEEIGTMLKKVEPNAKRIEEHRRKYQRLRKEKELQRAERERRKQQ 249
> dre:564225 st13, MGC73267, MGC77089, wu:fd15g02, zgc:73267;
suppression of tumorigenicity 13 (colon carcinoma) (Hsp70 interacting
protein); K09560 suppressor of tumorigenicity protein
13
Length=362
Score = 95.1 bits (235), Expect = 5e-20, Method: Compositional matrix adjust.
Identities = 50/126 (39%), Positives = 75/126 (59%), Gaps = 16/126 (12%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
A+LY +RA + +K ++ AAIRDCD A+ +NPD+A+ Y+ RG AH+ LGHW+++ D+ M
Sbjct 151 AILYAKRASVYVKMQKPNAAIRDCDRAISINPDSAQPYKWRGKAHKLLGHWEESARDLAM 210
Query 63 GQKIDYDENIWDIQKLVEQKFKIIEEH----------------ERSVQRRKEEEERKQRE 106
K+DYDE + K V+ K I +H + V++ +EE ER QRE
Sbjct 211 ACKLDYDEEASAMLKEVQPKANKIIDHRRKYERKREEREIRARQERVKKAREEHERAQRE 270
Query 107 RRAREQ 112
AR+Q
Sbjct 271 EEARQQ 276
> ath:AT4G22670 AtHip1; AtHip1 (Arabidopsis thaliana Hsp70-interacting
protein 1); binding
Length=441
Score = 94.0 bits (232), Expect = 1e-19, Method: Compositional matrix adjust.
Identities = 50/119 (42%), Positives = 71/119 (59%), Gaps = 0/119 (0%)
Query 2 TALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIE 61
+A++Y RA + +K K+ AAIRD + AL++NPD+A+ Y+ RG A LG W +A D+
Sbjct 156 SAIMYGNRASVYIKLKKPNAAIRDANAALEINPDSAKGYKSRGMARAMLGEWAEAAKDLH 215
Query 62 MGQKIDYDENIWDIQKLVEQKFKIIEEHERSVQRRKEEEERKQRERRAREQRANAQRTY 120
+ IDYDE I + K VE +EEH R R ++E E K+ ER +RA AQ Y
Sbjct 216 LASTIDYDEEISAVLKKVEPNAHKLEEHRRKYDRLRKEREDKKAERDRLRRRAEAQAAY 274
> cel:T12D8.8 hypothetical protein; K09560 suppressor of tumorigenicity
protein 13
Length=422
Score = 93.6 bits (231), Expect = 1e-19, Method: Compositional matrix adjust.
Identities = 44/96 (45%), Positives = 67/96 (69%), Gaps = 0/96 (0%)
Query 2 TALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIE 61
+A+L+ +RA++LLK KR AAI DCD+A+ +NPD+A+ Y+ RG A+R LG W +A +D+
Sbjct 148 SAMLHAKRANVLLKLKRPVAAIADCDKAISINPDSAQGYKFRGRANRLLGKWVEAKTDLA 207
Query 62 MGQKIDYDENIWDIQKLVEQKFKIIEEHERSVQRRK 97
K+DYDE + K VE I+E+ R+V+R+K
Sbjct 208 TACKLDYDEAANEWLKEVEPNAHKIQEYNRAVERQK 243
> dre:768178 zgc:153288
Length=591
Score = 55.1 bits (131), Expect = 5e-08, Method: Compositional matrix adjust.
Identities = 22/59 (37%), Positives = 39/59 (66%), Gaps = 0/59 (0%)
Query 2 TALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDI 60
+A ++ RA L++ +++ AA+ DCD L+L P N +A R T H++LGH +++H D+
Sbjct 226 SAAVFNNRAQTLIRLQQWPAALSDCDAVLQLEPHNIKALLRRATVHKHLGHQQESHDDL 284
Score = 47.0 bits (110), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 26/87 (29%), Positives = 49/87 (56%), Gaps = 2/87 (2%)
Query 2 TALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIE 61
T YT RA +K +R+ A +DCD AL++ P N +A+ R A++ L + SD++
Sbjct 483 TCAAYTNRALCYIKLERFTEARQDCDSALQIEPTNKKAFYRRALANKGLKDYLSCRSDLQ 542
Query 62 MGQKIDYDENIWDIQKLVEQKFKIIEE 88
Q + D ++ + Q+L+ + ++E+
Sbjct 543 --QVLRLDASVTEAQRLLMELTHLMED 567
> dre:564953 spag1, MGC162178, cb1089, wu:fj78g10; sperm associated
antigen 1
Length=386
Score = 53.1 bits (126), Expect = 2e-07, Method: Compositional matrix adjust.
Identities = 27/84 (32%), Positives = 49/84 (58%), Gaps = 2/84 (2%)
Query 5 LYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQ 64
+YT RA LK +R+A A +DCD AL++ P N +A+ R AH+ L + A +D++ +
Sbjct 297 IYTNRALCFLKLERFAEAKQDCDSALQMEPKNKKAFYRRALAHKGLKDYLSASTDLQ--E 354
Query 65 KIDYDENIWDIQKLVEQKFKIIEE 88
+ D N+ + ++ +E ++ E
Sbjct 355 VLQLDPNVQEAEQELEMVTNLLRE 378
Score = 38.5 bits (88), Expect = 0.005, Method: Compositional matrix adjust.
Identities = 36/129 (27%), Positives = 61/129 (47%), Gaps = 17/129 (13%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
+LY+ RA LK A I+DC AL+L+P + + R A+ L +++A+ D +
Sbjct 126 CVLYSNRAACFLKDGNSADCIQDCTRALELHPFSLKPLLRRAMAYESLERYRKAYVDYKT 185
Query 63 GQKIDYD-----ENIWDIQK-LVEQ-------KFKIIEEHERSVQRRKEEEERKQ----R 105
+ID +++ I K L+EQ K I S Q+ ++EE + R
Sbjct 186 VLQIDISVQAAHDSVHRITKMLIEQDGPDWREKLPEIPAVPLSAQQHRKEEPSAELLQAR 245
Query 106 ERRAREQRA 114
RA +++A
Sbjct 246 AERAEQEKA 254
> cel:Y39B6A.2 pph-5; Protein PHosphatase family member (pph-5);
K04460 protein phosphatase 5 [EC:3.1.3.16]
Length=496
Score = 51.6 bits (122), Expect = 6e-07, Method: Composition-based stats.
Identities = 24/66 (36%), Positives = 40/66 (60%), Gaps = 0/66 (0%)
Query 1 PTALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDI 60
PTA+LY RA LK++ Y +A+ D D A+ ++P + + R TA+ LG +K+A +D
Sbjct 60 PTAVLYGNRAQAYLKKELYGSALEDADNAIAIDPSYVKGFYRRATANMALGRFKKALTDY 119
Query 61 EMGQKI 66
+ K+
Sbjct 120 QAVVKV 125
> tpv:TP03_0587 hypothetical protein; K09553 stress-induced-phosphoprotein
1
Length=540
Score = 50.4 bits (119), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 27/72 (37%), Positives = 42/72 (58%), Gaps = 1/72 (1%)
Query 1 PT-ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSD 59
PT A LY+ RA LLK Y +A+ DC++AL+L+P +A+ +G H L + +A
Sbjct 386 PTDAKLYSNRAAALLKLCEYPSALADCNKALELDPTFVKAWARKGNLHVLLKEYHKAMDS 445
Query 60 IEMGQKIDYDEN 71
+ G K+D + N
Sbjct 446 YDKGLKVDPNNN 457
Score = 33.5 bits (75), Expect = 0.17, Method: Compositional matrix adjust.
Identities = 19/68 (27%), Positives = 32/68 (47%), Gaps = 2/68 (2%)
Query 4 LLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMG 63
+LY+ R+ Y A+ D ++ + L PD + Y +G LG+ ++A MG
Sbjct 36 VLYSNRSGAYASMYMYNEALADANKCIDLKPDWPKGYSRKGLCEYKLGNPEKAKETYNMG 95
Query 64 QKIDYDEN 71
+ YD N
Sbjct 96 --LAYDPN 101
Score = 29.3 bits (64), Expect = 3.6, Method: Compositional matrix adjust.
Identities = 16/54 (29%), Positives = 27/54 (50%), Gaps = 0/54 (0%)
Query 14 LKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKID 67
K ++ A ++ DEA+K NP +A+ Y R A L + A +D ++D
Sbjct 366 FKAFKFPEAKKEYDEAIKRNPTDAKLYSNRAAALLKLCEYPSALADCNKALELD 419
> ath:AT1G56440 serine/threonine protein phosphatase-related
Length=476
Score = 49.3 bits (116), Expect = 3e-06, Method: Composition-based stats.
Identities = 37/104 (35%), Positives = 55/104 (52%), Gaps = 7/104 (6%)
Query 1 PTALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDI 60
P A+ Y RA LK KRY A DC EAL L+ +AY R TA + LG K+A D
Sbjct 115 PNAVTYANRAMAYLKIKRYREAEVDCTEALNLDDRYIKAYSRRATARKELGMIKEAKEDA 174
Query 61 EMGQKIDYD-----ENIWDIQKLVEQKFKIIEEHERSVQRRKEE 99
E +++ + + DI+ L+E+ +IIE+ ++Q +E
Sbjct 175 EFALRLEPESQELKKQYADIKSLLEK--EIIEKATGAMQSTAQE 216
> mmu:26942 Spag1, tpis; sperm associated antigen 1
Length=901
Score = 48.1 bits (113), Expect = 7e-06, Method: Compositional matrix adjust.
Identities = 22/72 (30%), Positives = 45/72 (62%), Gaps = 0/72 (0%)
Query 1 PTALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDI 60
PTA+ Y RA +K +R+++A+ DC++AL+L+P N +A R T +++ ++A D+
Sbjct 244 PTAIAYNNRAQAEIKLQRWSSALEDCEKALELDPGNVKALLRRATTYKHQNKLQEAVDDL 303
Query 61 EMGQKIDYDENI 72
+++ D ++
Sbjct 304 RKVLQVEPDNDL 315
Score = 37.7 bits (86), Expect = 0.008, Method: Compositional matrix adjust.
Identities = 20/65 (30%), Positives = 36/65 (55%), Gaps = 0/65 (0%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
++LY+ RA LK+ I+DC+ AL+L+P + + R A+ L ++ A+ D +
Sbjct 472 SILYSNRAACYLKEGNCRDCIQDCNRALELHPFSVKPLLRRAMAYETLEQYRNAYVDYKT 531
Query 63 GQKID 67
+ID
Sbjct 532 VLQID 536
Score = 34.3 bits (77), Expect = 0.11, Method: Compositional matrix adjust.
Identities = 21/60 (35%), Positives = 30/60 (50%), Gaps = 1/60 (1%)
Query 13 LLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKIDYDENI 72
L+K K Y AI +E LK+N Y R + LG +++A D E +ID EN+
Sbjct 616 LVKDKNYKDAISKYNECLKINSKACAIYTNRALCYLKLGQFEEAKLDCEQALQID-GENV 674
Score = 30.4 bits (67), Expect = 1.5, Method: Compositional matrix adjust.
Identities = 16/55 (29%), Positives = 31/55 (56%), Gaps = 0/55 (0%)
Query 5 LYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSD 59
+YT RA LK ++ A DC++AL+++ +N +A A + L + +++ D
Sbjct 642 IYTNRALCYLKLGQFEEAKLDCEQALQIDGENVKASHRLALAQKGLENCRESGVD 696
> hsa:10953 TOMM34, HTOM34P, TOM34, URCC3; translocase of outer
mitochondrial membrane 34
Length=309
Score = 47.8 bits (112), Expect = 8e-06, Method: Compositional matrix adjust.
Identities = 23/55 (41%), Positives = 34/55 (61%), Gaps = 0/55 (0%)
Query 6 YTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDI 60
Y+ RA L K+Y A++DC EALKL+ N +A+ R AH+ L +K + +DI
Sbjct 230 YSNRALCYLVLKQYTEAVKDCTEALKLDGKNVKAFYRRAQAHKALKDYKSSFADI 284
Score = 34.7 bits (78), Expect = 0.080, Method: Compositional matrix adjust.
Identities = 20/65 (30%), Positives = 32/65 (49%), Gaps = 0/65 (0%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
++LY+ RA LK I+DC AL L P + + R +A+ L + A+ D +
Sbjct 51 SVLYSNRAACHLKDGNCRDCIKDCTSALALVPFSIKPLLRRASAYEALEKYPMAYVDYKT 110
Query 63 GQKID 67
+ID
Sbjct 111 VLQID 115
> hsa:6674 SPAG1, FLJ32920, HSD-3.8, SP75, TPIS; sperm associated
antigen 1
Length=926
Score = 47.8 bits (112), Expect = 8e-06, Method: Compositional matrix adjust.
Identities = 30/99 (30%), Positives = 58/99 (58%), Gaps = 2/99 (2%)
Query 5 LYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQ 64
+YT RA LK ++ A +DCD+AL+L N +A+ R AH+ L ++++ S I++ +
Sbjct 659 IYTNRALCYLKLCQFEEAKQDCDQALQLADGNVKAFYRRALAHKGLKNYQK--SLIDLNK 716
Query 65 KIDYDENIWDIQKLVEQKFKIIEEHERSVQRRKEEEERK 103
I D +I + + +E+ +++ +++ KE+E RK
Sbjct 717 VILLDPSIIEAKMELEEVTRLLNLKDKTAPFNKEKERRK 755
Score = 41.6 bits (96), Expect = 6e-04, Method: Compositional matrix adjust.
Identities = 25/108 (23%), Positives = 52/108 (48%), Gaps = 9/108 (8%)
Query 1 PTALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDI 60
PT + Y RA +K + + +A +DC++ L+L P N +A R T +++ ++A D+
Sbjct 240 PTVVAYNNRAQAEIKLQNWNSAFQDCEKVLELEPGNVKALLRRATTYKHQNKLREATEDL 299
Query 61 EMGQKIDYDENIWDIQKLVEQKFKIIEEHERSVQRRKEEEERKQRERR 108
+ D++ + K + E ER ++ + E + + +R
Sbjct 300 ---------SKVLDVEPDNDLAKKTLSEVERDLKNSEAASETQTKGKR 338
Score = 40.8 bits (94), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 20/65 (30%), Positives = 37/65 (56%), Gaps = 0/65 (0%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
++LY+ RA LK+ + I+DC+ AL+L+P + + R A+ L + +A+ D +
Sbjct 487 SILYSNRAACYLKEGNCSGCIQDCNRALELHPFSMKPLLRRAMAYETLEQYGKAYVDYKT 546
Query 63 GQKID 67
+ID
Sbjct 547 VLQID 551
> xla:414472 rpap3, MGC81126; RNA polymerase II associated protein
3
Length=660
Score = 46.2 bits (108), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 40/133 (30%), Positives = 63/133 (47%), Gaps = 25/133 (18%)
Query 2 TALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIE 61
A+L T RA + K++A A DC+ A+ LN D A+AY RG A L + + A D E
Sbjct 165 NAILPTNRASAFFRLKKFAVAESDCNLAIALNRDYAKAYARRGAARLALKNLQGAKEDYE 224
Query 62 MGQKIDYD----------------ENIWDIQKLVEQKFKIIEEHERSVQRRKEEEERKQR 105
++D + + D+Q+ + + KI E+ EEE+KQ
Sbjct 225 KVLELDANNFEAKNELRKINQELYSSASDVQENMATEAKITVEN---------EEEKKQI 275
Query 106 ERRAREQRANAQR 118
E + R+Q+A Q+
Sbjct 276 EIQQRKQQAIMQK 288
Score = 46.2 bits (108), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 27/66 (40%), Positives = 35/66 (53%), Gaps = 0/66 (0%)
Query 2 TALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIE 61
ALL RA LK ++Y A DC A+ L+ +A+ RGTA LG K+A D E
Sbjct 317 NALLPANRAMAYLKIQKYKEAEADCTLAISLDASYCKAFARRGTASIMLGKQKEAKEDFE 376
Query 62 MGQKID 67
M K+D
Sbjct 377 MVLKLD 382
> mmu:71919 Rpap3, 2310042P20Rik, D15Ertd682e; RNA polymerase
II associated protein 3
Length=660
Score = 45.8 bits (107), Expect = 4e-05, Method: Compositional matrix adjust.
Identities = 25/59 (42%), Positives = 34/59 (57%), Gaps = 0/59 (0%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIE 61
ALL RA LK +RY A RDC +A+ L+ ++A+ RGTA +LG +A D E
Sbjct 318 ALLPANRAMAYLKIQRYEEAERDCTQAIVLDGSYSKAFARRGTARTFLGKINEAKQDFE 376
Score = 32.7 bits (73), Expect = 0.25, Method: Compositional matrix adjust.
Identities = 20/58 (34%), Positives = 29/58 (50%), Gaps = 0/58 (0%)
Query 4 LLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIE 61
+L T RA + K++A A DC+ A+ L+ +AY RG A L + A D E
Sbjct 169 VLPTNRASAYFRLKKFAVAESDCNLAIALSRTYTKAYARRGAARFALQKLEDARKDYE 226
> dre:323361 tomm34, wu:fb96b08, zgc:56645; translocase of outer
mitochondrial membrane 34
Length=305
Score = 45.4 bits (106), Expect = 5e-05, Method: Compositional matrix adjust.
Identities = 30/76 (39%), Positives = 42/76 (55%), Gaps = 2/76 (2%)
Query 6 YTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQK 65
YT RA L K Y AIRDC+EAL+L+ N +A R A++ L + K D+ K
Sbjct 227 YTNRALCYLALKMYKDAIRDCEEALRLDSANIKALYRRAQAYKELKNKKSCIEDLNSVLK 286
Query 66 IDYDENIWDIQKLVEQ 81
I D N +QKL+++
Sbjct 287 I--DPNNTAVQKLLQE 300
Score = 36.6 bits (83), Expect = 0.019, Method: Compositional matrix adjust.
Identities = 21/67 (31%), Positives = 33/67 (49%), Gaps = 0/67 (0%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
+LY+ RA LK I+DC +L L P +A R A L ++QA+ D +
Sbjct 52 GILYSNRAASYLKDGNCNECIKDCTASLDLVPFGFKALLRRAAAFEALERYRQAYVDYKT 111
Query 63 GQKIDYD 69
+ID++
Sbjct 112 VLQIDWN 118
> dre:568208 ttc6; tetratricopeptide repeat domain 6
Length=720
Score = 44.7 bits (104), Expect = 6e-05, Method: Compositional matrix adjust.
Identities = 22/63 (34%), Positives = 37/63 (58%), Gaps = 0/63 (0%)
Query 5 LYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQ 64
++ RA + + R+A AI DC+EA+++ P + RA+ +G YL +K A D+ M
Sbjct 276 VHLSRAALYGAEGRHAKAILDCNEAIRIQPKSLRAHLYKGALKFYLKAYKSAVEDLTMAV 335
Query 65 KID 67
+ID
Sbjct 336 QID 338
Score = 31.6 bits (70), Expect = 0.63, Method: Compositional matrix adjust.
Identities = 18/61 (29%), Positives = 31/61 (50%), Gaps = 0/61 (0%)
Query 6 YTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQK 65
Y RA++ +++ +A RD +AL L P +A Y++R LG ++A D K
Sbjct 657 YFNRANLHCSLRQFQSAERDLTQALVLEPGDALLYKLRADVRGCLGWMEEAMEDYRTALK 716
Query 66 I 66
+
Sbjct 717 L 717
Score = 29.6 bits (65), Expect = 2.3, Method: Compositional matrix adjust.
Identities = 15/39 (38%), Positives = 23/39 (58%), Gaps = 0/39 (0%)
Query 22 AIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDI 60
+++D +EAL LNP +A AY R H L ++ A D+
Sbjct 639 SLQDFNEALCLNPLSAHAYFNRANLHCSLRQFQSAERDL 677
Score = 28.5 bits (62), Expect = 5.5, Method: Compositional matrix adjust.
Identities = 14/43 (32%), Positives = 24/43 (55%), Gaps = 0/43 (0%)
Query 17 KRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSD 59
K Y +A+ D A++++P + AY RG + L H++ A D
Sbjct 322 KAYKSAVEDLTMAVQIDPACSFAYYNRGICFQELQHYEMALRD 364
Score = 27.7 bits (60), Expect = 8.6, Method: Compositional matrix adjust.
Identities = 19/54 (35%), Positives = 29/54 (53%), Gaps = 4/54 (7%)
Query 5 LYTRRADILLKQKRY--AAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQA 56
L+T R L++Q R ++A++D A+ LNP A AY Y G ++QA
Sbjct 554 LFTNRG--LIQQLRGDKSSAMKDYQTAISLNPAYALAYFNAANLFFYNGQFEQA 605
> sce:YOR027W STI1; Hsp90 cochaperone, interacts with the Ssa
group of the cytosolic Hsp70 chaperones; activates the ATPase
activity of Ssa1p; homolog of mammalian Hop protein; K09553
stress-induced-phosphoprotein 1
Length=589
Score = 44.7 bits (104), Expect = 7e-05, Method: Compositional matrix adjust.
Identities = 20/67 (29%), Positives = 37/67 (55%), Gaps = 0/67 (0%)
Query 1 PTALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDI 60
P +LY+ R+ K+++ A+ D +E +K+NP ++ Y G AH LG +A S+
Sbjct 38 PNHVLYSNRSACYTSLKKFSDALNDANECVKINPSWSKGYNRLGAAHLGLGDLDEAESNY 97
Query 61 EMGQKID 67
+ ++D
Sbjct 98 KKALELD 104
Score = 34.3 bits (77), Expect = 0.086, Method: Compositional matrix adjust.
Identities = 19/66 (28%), Positives = 34/66 (51%), Gaps = 0/66 (0%)
Query 6 YTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQK 65
Y+ RA L K + AI DC++A++ +P+ RAY + TA + + A ++ +
Sbjct 433 YSNRAAALAKLMSFPEAIADCNKAIEKDPNFVRAYIRKATAQIAVKEYASALETLDAART 492
Query 66 IDYDEN 71
D + N
Sbjct 493 KDAEVN 498
> pfa:MAL13P1.274 PfPP5; serine/threonine protein phosphatase
(EC:3.1.3.16); K04460 protein phosphatase 5 [EC:3.1.3.16]
Length=658
Score = 44.3 bits (103), Expect = 1e-04, Method: Composition-based stats.
Identities = 31/95 (32%), Positives = 52/95 (54%), Gaps = 5/95 (5%)
Query 4 LLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMG 63
+ YT R+ +K + Y AI D DEA+K+NP A+AY +G ++ L K+A +
Sbjct 226 IYYTNRSFCHIKLENYGTAIEDIDEAIKINPYYAKAYYRKGCSYLLLSDLKRASECFQKV 285
Query 64 QKIDYDEN----IWDIQKLV-EQKFKIIEEHERSV 93
K+ D+N + +KL+ EQ+F+ E E+ +
Sbjct 286 LKLTKDKNSELKLKQCKKLIFEQQFQKAIELEQKM 320
> hsa:79657 RPAP3, FLJ21908; RNA polymerase II associated protein
3
Length=631
Score = 43.9 bits (102), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 23/59 (38%), Positives = 34/59 (57%), Gaps = 0/59 (0%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIE 61
ALL RA LK ++Y A +DC +A+ L+ ++A+ RGTA +LG +A D E
Sbjct 316 ALLPANRAMAYLKIQKYEEAEKDCTQAILLDGSYSKAFARRGTARTFLGKLNEAKQDFE 374
Score = 36.6 bits (83), Expect = 0.021, Method: Compositional matrix adjust.
Identities = 21/58 (36%), Positives = 30/58 (51%), Gaps = 0/58 (0%)
Query 4 LLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIE 61
+L T RA + K++A A DC+ A+ LN +AY RG A L ++A D E
Sbjct 168 VLPTNRASAYFRLKKFAVAESDCNLAVALNRSYTKAYSRRGAARFALQKLEEAKKDYE 225
> dre:563791 ttc12; tetratricopeptide repeat domain 12
Length=700
Score = 43.5 bits (101), Expect = 2e-04, Method: Compositional matrix adjust.
Identities = 22/52 (42%), Positives = 30/52 (57%), Gaps = 0/52 (0%)
Query 5 LYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQA 56
LYT RA +K KRY AI DC+ AL+ N +A+ GT+H L + Q+
Sbjct 149 LYTNRAQAFIKLKRYKEAISDCEWALRCNEKCIKAFIHMGTSHLALKDFTQS 200
> sce:YBR155W CNS1; Cns1p
Length=385
Score = 43.5 bits (101), Expect = 2e-04, Method: Compositional matrix adjust.
Identities = 34/111 (30%), Positives = 54/111 (48%), Gaps = 12/111 (10%)
Query 5 LYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMG- 63
LY RA L+ K Y I DC +AL +NP N + Y A L ++A S
Sbjct 123 LYANRAACELELKNYRRCIEDCSKALTINPKNVKCYYRTSKAFFQLNKLEEAKSAATFAN 182
Query 64 QKIDY-DENIWDIQKLVEQKFKIIEEHERSVQRRKEEEERKQRERRAREQR 113
Q+ID +++I ++ ++ +R Q K +EE++QRE + RE +
Sbjct 183 QRIDPENKSILNMLSVI----------DRKEQELKAKEEKQQREAQERENK 223
> mmu:100503562 hypothetical protein LOC100503562
Length=1010
Score = 43.1 bits (100), Expect = 2e-04, Method: Compositional matrix adjust.
Identities = 20/50 (40%), Positives = 30/50 (60%), Gaps = 0/50 (0%)
Query 18 RYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKID 67
RY+ AI +C+EA+KL P++ RAY RG Y +K +D+ K+D
Sbjct 576 RYSKAILNCNEAIKLYPESVRAYICRGVLKYYNRTYKLGITDLSTAIKMD 625
Score = 33.9 bits (76), Expect = 0.12, Method: Compositional matrix adjust.
Identities = 23/68 (33%), Positives = 30/68 (44%), Gaps = 2/68 (2%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
A LY RA + K+Y A D L L PD A Y +R +G +A SD
Sbjct 940 AALYFNRASFYVYLKKYKLAEEDLGIGLSLKPDEAIMYNLRAQVRGKMGLIAEAMSD--Y 997
Query 63 GQKIDYDE 70
Q +D +E
Sbjct 998 NQALDLEE 1005
Score = 31.6 bits (70), Expect = 0.61, Method: Compositional matrix adjust.
Identities = 20/66 (30%), Positives = 29/66 (43%), Gaps = 2/66 (3%)
Query 8 RRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKID 67
+R + + A+ D LKL+P N++A RG A+ +KQA D I
Sbjct 325 KRGMFYFENGNWIGAVYDFTSLLKLDPYNSKARTYRGRAYFKRNLYKQATQDFSAA--IH 382
Query 68 YDENIW 73
D N W
Sbjct 383 LDPNNW 388
Score = 28.9 bits (63), Expect = 4.0, Method: Compositional matrix adjust.
Identities = 16/62 (25%), Positives = 27/62 (43%), Gaps = 0/62 (0%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
+L Y +I L ++++ A ALK NP+N A R + L +++A D
Sbjct 872 SLAYFNAGNIYLHHRQFSQASDYFSTALKFNPENEYALMNRAVTNSVLKKYEEAEKDFSC 931
Query 63 GQ 64
Sbjct 932 AM 933
> sce:YGR123C PPT1; Ppt1p (EC:3.1.3.16); K04460 protein phosphatase
5 [EC:3.1.3.16]
Length=513
Score = 43.1 bits (100), Expect = 2e-04, Method: Composition-based stats.
Identities = 20/60 (33%), Positives = 35/60 (58%), Gaps = 0/60 (0%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
++ ++ RA K + +A+ DCDEA+KL+P N +AY R + L +K+A D+ +
Sbjct 46 SIYFSNRAFAHFKVDNFQSALNDCDEAIKLDPKNIKAYHRRALSCMALLEFKKARKDLNV 105
> xla:414520 hypothetical protein MGC81394
Length=312
Score = 42.7 bits (99), Expect = 3e-04, Method: Compositional matrix adjust.
Identities = 23/61 (37%), Positives = 32/61 (52%), Gaps = 7/61 (11%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYL-------GHWKQ 55
A+ Y RA K YA A+RDC+EA+ ++P ++AY G A L G +KQ
Sbjct 122 AVYYCNRAAAYSKLGNYAGAVRDCEEAISIDPSYSKAYGRMGLALSSLNKHAESVGFYKQ 181
Query 56 A 56
A
Sbjct 182 A 182
Score = 32.3 bits (72), Expect = 0.38, Method: Compositional matrix adjust.
Identities = 19/54 (35%), Positives = 29/54 (53%), Gaps = 0/54 (0%)
Query 14 LKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKID 67
+K + + +A+ +AL+LNP NA Y R A+ LG++ A D E ID
Sbjct 99 MKVENFESAVTYYTKALELNPRNAVYYCNRAAAYSKLGNYAGAVRDCEEAISID 152
> bbo:BBOV_III002230 17.m07215; tetratricopeptide repeat domain
containing protein; K09553 stress-induced-phosphoprotein 1
Length=546
Score = 42.4 bits (98), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 20/63 (31%), Positives = 35/63 (55%), Gaps = 0/63 (0%)
Query 5 LYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQ 64
LYT RA L K Y +A+ DC++A++++P +A+ +G H L + +A + G
Sbjct 396 LYTNRAAALTKLGEYPSALADCNKAVEMDPTFVKAWARKGNLHVLLKEYSKALEAYDKGL 455
Query 65 KID 67
+D
Sbjct 456 ALD 458
Score = 37.4 bits (85), Expect = 0.012, Method: Compositional matrix adjust.
Identities = 19/65 (29%), Positives = 34/65 (52%), Gaps = 0/65 (0%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
+LY+ R+ +R+ A+ D ++ + L PD + Y +G A LG ++A + +
Sbjct 35 GILYSNRSGAYASLQRFQEALDDANQCVSLKPDWPKGYSRKGLALYKLGRLQEARTAYQE 94
Query 63 GQKID 67
G KID
Sbjct 95 GLKID 99
Score = 30.8 bits (68), Expect = 1.1, Method: Compositional matrix adjust.
Identities = 15/59 (25%), Positives = 30/59 (50%), Gaps = 0/59 (0%)
Query 9 RADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKID 67
+ + K+ ++ A ++ DEA++ NP + + Y R A LG + A +D ++D
Sbjct 366 KGNAFFKKFQFPEAKKEYDEAIRRNPSDIKLYTNRAAALTKLGEYPSALADCNKAVEMD 424
> ath:AT1G33400 tetratricopeptide repeat (TPR)-containing protein
Length=798
Score = 42.4 bits (98), Expect = 4e-04, Method: Composition-based stats.
Identities = 21/65 (32%), Positives = 37/65 (56%), Gaps = 0/65 (0%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
A L+ RA++L ++RDC AL+++P A+A+ RG + LG++K A DI +
Sbjct 106 ASLFLNRANVLHNLGLLKESLRDCHRALRIDPYYAKAWYRRGKLNTLLGNYKDAFRDITV 165
Query 63 GQKID 67
++
Sbjct 166 SMSLE 170
> hsa:54557 SGTB, FLJ39002, SGT2; small glutamine-rich tetratricopeptide
repeat (TPR)-containing, beta
Length=304
Score = 41.2 bits (95), Expect = 7e-04, Method: Compositional matrix adjust.
Identities = 23/55 (41%), Positives = 33/55 (60%), Gaps = 2/55 (3%)
Query 14 LKQKRYAAAIRDC-DEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKID 67
+K++ YAAA+ DC +A++L+P+NA Y R A LGH+ A D E ID
Sbjct 96 MKEENYAAAV-DCYTQAIELDPNNAVYYCNRAAAQSKLGHYTDAIKDCEKAIAID 149
Score = 32.3 bits (72), Expect = 0.37, Method: Compositional matrix adjust.
Identities = 22/83 (26%), Positives = 42/83 (50%), Gaps = 1/83 (1%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
A+ Y RA K Y AI+DC++A+ ++ ++AY G A L +++A + +
Sbjct 119 AVYYCNRAAAQSKLGHYTDAIKDCEKAIAIDSKYSKAYGRMGLALTALNKFEEAVTSYQK 178
Query 63 GQKIDYDENIWDIQ-KLVEQKFK 84
+D + + + K+ EQK +
Sbjct 179 ALDLDPENDSYKSNLKIAEQKLR 201
> xla:496358 sgta; small glutamine-rich tetratricopeptide repeat
(TPR)-containing, alpha
Length=302
Score = 41.2 bits (95), Expect = 8e-04, Method: Compositional matrix adjust.
Identities = 26/83 (31%), Positives = 40/83 (48%), Gaps = 1/83 (1%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
A+ Y RA K YA A+RDC+ A+ ++P+ ++AY G A L +A +
Sbjct 110 AVYYCNRAAAYSKLGNYAGAVRDCEAAITIDPNYSKAYGRMGLALSSLNKHAEAVGFYKQ 169
Query 63 GQKIDYDENIWDIQ-KLVEQKFK 84
+D D + K+ EQK K
Sbjct 170 ALVLDPDNETYKSNLKIAEQKMK 192
Score = 34.7 bits (78), Expect = 0.085, Method: Compositional matrix adjust.
Identities = 20/54 (37%), Positives = 29/54 (53%), Gaps = 0/54 (0%)
Query 14 LKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKID 67
+K + + +AI +AL+LNP NA Y R A+ LG++ A D E ID
Sbjct 87 MKVENFESAISYYTKALELNPANAVYYCNRAAAYSKLGNYAGAVRDCEAAITID 140
> ath:AT4G23570 SGT1A; SGT1A; protein binding; K12795 suppressor
of G2 allele of SKP1
Length=351
Score = 41.2 bits (95), Expect = 8e-04, Method: Compositional matrix adjust.
Identities = 29/92 (31%), Positives = 49/92 (53%), Gaps = 4/92 (4%)
Query 5 LYTRRADILLKQKRYAA-AIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMG 63
+ RA +K + + A A+ D ++A++L+P +AY +GTA L ++ A + +E G
Sbjct 38 FFADRAQAYIKLESFTAEAVADANKAIELDPSLTKAYLRKGTACMKLEEYRTAKTALEKG 97
Query 64 QKIDYDENIWDIQKLV-EQKFKIIEEHERSVQ 94
I E+ +KL+ E F I EE + VQ
Sbjct 98 ASITPSES--KFKKLIDECNFLITEEEKDLVQ 127
> hsa:5481 PPID, CYP-40, CYPD, MGC33096; peptidylprolyl isomerase
D (EC:5.2.1.8); K05864 peptidyl-prolyl isomerase D (cyclophilin
D) [EC:5.2.1.8]
Length=370
Score = 40.8 bits (94), Expect = 0.001, Method: Composition-based stats.
Identities = 26/81 (32%), Positives = 43/81 (53%), Gaps = 1/81 (1%)
Query 14 LKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKIDYDENIW 73
LK + AI C EAL+L+P N +A R + L + QA +D++ Q I ++
Sbjct 284 LKMSNWQGAIDSCLEALELDPSNTKALYRRAQGWQGLKEYDQALADLKKAQGIAPEDKAI 343
Query 74 DIQKL-VEQKFKIIEEHERSV 93
+ L V+QK K ++ E++V
Sbjct 344 QAELLKVKQKIKAQKDKEKAV 364
> mmu:100048137 tetratricopeptide repeat protein 12-like
Length=544
Score = 40.8 bits (94), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 21/53 (39%), Positives = 30/53 (56%), Gaps = 0/53 (0%)
Query 4 LLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQA 56
+LYT RA +K Y A+ DCD ALK + + +AY G AH L ++ +A
Sbjct 140 VLYTNRAQAFIKLGDYQKALVDCDWALKCDENCTKAYFHMGKAHVALKNYSKA 192
> cpv:cgd7_570 protein with 2 TPR domains
Length=390
Score = 40.4 bits (93), Expect = 0.001, Method: Compositional matrix adjust.
Identities = 38/111 (34%), Positives = 57/111 (51%), Gaps = 7/111 (6%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGT-AHRYLGHWKQAHSDIE 61
+LLY+ RA + L KRY + DC +LK NP N +A RG A + ++QA +
Sbjct 130 SLLYSNRAHVYLLLKRYVDCVDDCRASLKENPKNVKA-AYRGCRASMCMQLYRQALNFAL 188
Query 62 MGQKIDYDENIWDIQKLVEQKFKIIEEHERSVQRRKEEEERKQRERRAREQ 112
G K Y+ ++ KL Q + + E E+ RRKE EE ++R+ E
Sbjct 189 HGLK--YEPENPELLKLKSQLEERLSEIEK---RRKEREELEKRDGGKNES 234
> mmu:235330 Ttc12, E330017O07Rik; tetratricopeptide repeat domain
12
Length=704
Score = 40.0 bits (92), Expect = 0.002, Method: Composition-based stats.
Identities = 31/98 (31%), Positives = 50/98 (51%), Gaps = 5/98 (5%)
Query 4 LLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMG 63
+LYT RA +K Y A+ DCD ALK + + +AY G AH L ++ +A E
Sbjct 140 VLYTNRAQAFIKLGDYQKALVDCDWALKCDENCTKAYFHMGKAHVALKNYSKAK---ECY 196
Query 64 QKIDYDENIWDIQKLVEQKFKIIEEHERSVQRRKEEEE 101
QKI +E ++ V++ + E++ + KE +E
Sbjct 197 QKI--EEINPKLKAQVKEHLNQVTLREKADLQEKEAQE 232
> tgo:TGME49_047000 TPR domain-containing protein (EC:3.4.21.72)
Length=888
Score = 40.0 bits (92), Expect = 0.002, Method: Composition-based stats.
Identities = 31/109 (28%), Positives = 49/109 (44%), Gaps = 16/109 (14%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
A+L + R L K + A + DC EA++ +P A+AY R TA+ L W A +DI
Sbjct 744 AVLLSNRGACHLHGKCWEAVVADCTEAIQCDPSYAKAYLRRFTANEALTKWHDAAADINK 803
Query 63 GQKIDYDENIWDIQKLVEQKFKIIEEHERSVQRRKEEEERKQRERRARE 111
++D +E RS Q+R +++ Q E+ E
Sbjct 804 AIELDPS----------------LEARYRSDQQRVKKKSEAQFEKEKEE 836
> cpv:cgd2_1850 stress-induced protein sti1-like protein
Length=326
Score = 39.7 bits (91), Expect = 0.002, Method: Compositional matrix adjust.
Identities = 20/55 (36%), Positives = 30/55 (54%), Gaps = 0/55 (0%)
Query 13 LLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKID 67
L KQK Y AA ++ DEA+K NP ++R Y R + L + A D++ +D
Sbjct 150 LFKQKNYPAAKKEYDEAIKRNPSDSRLYSNRAACYMQLLEYPSALIDVQKALDLD 204
Score = 38.1 bits (87), Expect = 0.007, Method: Compositional matrix adjust.
Identities = 20/65 (30%), Positives = 34/65 (52%), Gaps = 0/65 (0%)
Query 5 LYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQ 64
LY+ RA ++ Y +A+ D +AL L+P +A+ +G H +L + +A + G
Sbjct 176 LYSNRAACYMQLLEYPSALIDVQKALDLDPKFTKAWSRKGNIHYFLKEYHKALHAYQEGL 235
Query 65 KIDYD 69
K D D
Sbjct 236 KCDPD 240
> xla:447715 ppid, MGC81732, cyp-40, cypd; peptidylprolyl isomerase
D (cyclophilin D) (EC:5.2.1.8); K05864 peptidyl-prolyl
isomerase D (cyclophilin D) [EC:5.2.1.8]
Length=370
Score = 39.3 bits (90), Expect = 0.003, Method: Composition-based stats.
Identities = 24/81 (29%), Positives = 45/81 (55%), Gaps = 1/81 (1%)
Query 14 LKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKIDYDENIW 73
LK + AAI C+EAL+++P + +A R + L ++QA D++ ++ D+
Sbjct 284 LKVSDFRAAIDSCNEALEIDPSHTKALYRRAQGWQGLKDYEQALEDLKKAHELSPDDKAV 343
Query 74 DIQKL-VEQKFKIIEEHERSV 93
+ L V+Q+ K +E E++V
Sbjct 344 SSEILRVKQRIKEQKEKEKAV 364
> pfa:PF14_0324 Hsp70/Hsp90 organizing protein, putative; K09553
stress-induced-phosphoprotein 1
Length=564
Score = 38.9 bits (89), Expect = 0.004, Method: Compositional matrix adjust.
Identities = 20/65 (30%), Positives = 35/65 (53%), Gaps = 0/65 (0%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
A LY+ RA L K Y +A+ D +A++L+P +AY +G H ++ + +A
Sbjct 412 AKLYSNRAAALTKLIEYPSALEDVMKAIELDPTFVKAYSRKGNLHFFMKDYYKALQAYNK 471
Query 63 GQKID 67
G ++D
Sbjct 472 GLELD 476
Score = 28.1 bits (61), Expect = 6.3, Method: Compositional matrix adjust.
Identities = 15/54 (27%), Positives = 27/54 (50%), Gaps = 0/54 (0%)
Query 14 LKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKID 67
K + A ++ DEA++ NP++A+ Y R A L + A D+ ++D
Sbjct 389 FKNNDFPNAKKEYDEAIRRNPNDAKLYSNRAAALTKLIEYPSALEDVMKAIELD 442
> dre:393173 MGC56178; zgc:56178
Length=595
Score = 38.9 bits (89), Expect = 0.004, Method: Compositional matrix adjust.
Identities = 30/79 (37%), Positives = 41/79 (51%), Gaps = 2/79 (2%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
ALL RA LK R+A A +DC AL L+P +A+ R TA LG + A D E
Sbjct 313 ALLPANRAMAFLKLNRFAEAEQDCSAALALDPSYTKAFARRATARAALGKCRDARDDFEQ 372
Query 63 GQKIDYD--ENIWDIQKLV 79
K++ + I +I+KL
Sbjct 373 VLKLEPGNKQAISEIEKLT 391
Score = 34.3 bits (77), Expect = 0.095, Method: Compositional matrix adjust.
Identities = 24/74 (32%), Positives = 36/74 (48%), Gaps = 2/74 (2%)
Query 7 TRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKI 66
T RA + K++A A DC+ A+ L+ +AY R L ++A D EM K+
Sbjct 166 TNRATCFYRLKKFAVAESDCNLAIALDSKYVKAYIRRAATRTALQKHREALEDYEMVLKL 225
Query 67 DY--DENIWDIQKL 78
D E ++QKL
Sbjct 226 DPGNSEAQTEVQKL 239
> ath:AT3G58620 TTL4; TTL4 (Tetratricopetide-repeat Thioredoxin-Like
4); binding
Length=682
Score = 38.9 bits (89), Expect = 0.004, Method: Composition-based stats.
Identities = 19/63 (30%), Positives = 34/63 (53%), Gaps = 0/63 (0%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
++LY RA K + ++ DC++AL++ P +A R ++ LG W+ A D E+
Sbjct 483 SVLYCNRAACWFKLGMWEKSVDDCNQALRIQPSYTKALLRRAASYGKLGRWEDAVRDYEV 542
Query 63 GQK 65
+K
Sbjct 543 LRK 545
Score = 28.9 bits (63), Expect = 4.5, Method: Composition-based stats.
Identities = 15/50 (30%), Positives = 27/50 (54%), Gaps = 0/50 (0%)
Query 7 TRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQA 56
+ RA L R A+++C EA++ +P ARA++ + + LG + A
Sbjct 249 SNRAAALAASGRLEEAVKECLEAVRCDPSYARAHQRLASLYLRLGEAENA 298
> dre:336867 fk20d10, wu:fa05b08, wu:fc52b05, wu:fk20d10, zgc:77080;
zgc:55741
Length=320
Score = 38.9 bits (89), Expect = 0.004, Method: Compositional matrix adjust.
Identities = 23/83 (27%), Positives = 44/83 (53%), Gaps = 1/83 (1%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
A+ + RA K YA A++DC+ A+ ++ + ++AY G A L + +A S +
Sbjct 125 AVYFCNRAAAYSKLGNYAGAVQDCERAIGIDANYSKAYGRMGLALASLNKYSEAVSYYKK 184
Query 63 GQKIDYDENIWDIQ-KLVEQKFK 84
++D D + + + ++ EQK K
Sbjct 185 ALELDPDNDTYKVNLQVAEQKVK 207
Score = 35.0 bits (79), Expect = 0.055, Method: Compositional matrix adjust.
Identities = 18/54 (33%), Positives = 30/54 (55%), Gaps = 0/54 (0%)
Query 14 LKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKID 67
+K + ++AA+ +A++LNP NA + R A+ LG++ A D E ID
Sbjct 102 MKVENFSAAVEFYSKAIQLNPQNAVYFCNRAAAYSKLGNYAGAVQDCERAIGID 155
> mmu:218544 Sgtb, C630001O05Rik, MGC27660; small glutamine-rich
tetratricopeptide repeat (TPR)-containing, beta
Length=304
Score = 38.5 bits (88), Expect = 0.005, Method: Compositional matrix adjust.
Identities = 22/55 (40%), Positives = 32/55 (58%), Gaps = 2/55 (3%)
Query 14 LKQKRYAAAIRDC-DEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEMGQKID 67
+K++ YAAA+ DC +A++L+P+NA Y R A L H+ A D E ID
Sbjct 96 MKEENYAAAV-DCYTQAIELDPNNAVYYCNRAAAQSKLSHYTDAIKDCEKAIAID 149
Score = 32.0 bits (71), Expect = 0.47, Method: Compositional matrix adjust.
Identities = 21/83 (25%), Positives = 42/83 (50%), Gaps = 1/83 (1%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIEM 62
A+ Y RA K Y AI+DC++A+ ++ ++AY G A + +++A + +
Sbjct 119 AVYYCNRAAAQSKLSHYTDAIKDCEKAIAIDSKYSKAYGRMGLALTAMNKFEEAVTSYQK 178
Query 63 GQKIDYDENIWDIQ-KLVEQKFK 84
+D + + + K+ EQK +
Sbjct 179 ALDLDPENDSYKSNLKIAEQKLR 201
> tpv:TP02_0944 serine/threonine protein phosphatase; K04460 protein
phosphatase 5 [EC:3.1.3.16]
Length=548
Score = 38.5 bits (88), Expect = 0.005, Method: Composition-based stats.
Identities = 20/54 (37%), Positives = 32/54 (59%), Gaps = 0/54 (0%)
Query 6 YTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSD 59
Y+ RA +K + Y +AI D + A++L PD +AY RG A+ L ++ A +D
Sbjct 117 YSNRAICNIKIENYGSAISDANMAIELRPDFFKAYYRRGCAYLCLLKFQDAETD 170
> ath:AT3G17970 atToc64-III (Arabidopsis thaliana translocon at
the outer membrane of chloroplasts 64-III); binding / carbon-nitrogen
ligase, with glutamine as amido-N-donor
Length=589
Score = 38.5 bits (88), Expect = 0.005, Method: Composition-based stats.
Identities = 22/57 (38%), Positives = 29/57 (50%), Gaps = 0/57 (0%)
Query 3 ALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSD 59
A Y+ RA L+ + A DC +A+ L+ N +AY RGTA LG K A D
Sbjct 508 ATYYSNRAAAYLELGGFLQAEEDCTKAITLDKKNVKAYLRRGTAREMLGDCKGAIED 564
Score = 32.0 bits (71), Expect = 0.53, Method: Composition-based stats.
Identities = 20/66 (30%), Positives = 32/66 (48%), Gaps = 0/66 (0%)
Query 2 TALLYTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWKQAHSDIE 61
+A + + + K+K + AI EA+KL+ +NA Y R A+ LG + QA D
Sbjct 473 SAEIAKEKGNQAFKEKLWQKAIGLYSEAIKLSDNNATYYSNRAAAYLELGGFLQAEEDCT 532
Query 62 MGQKID 67
+D
Sbjct 533 KAITLD 538
> mmu:67145 Tomm34, 2610100K07Rik, TOM34, Tomm34a, Tomm34b; translocase
of outer mitochondrial membrane 34
Length=309
Score = 38.1 bits (87), Expect = 0.006, Method: Compositional matrix adjust.
Identities = 20/49 (40%), Positives = 30/49 (61%), Gaps = 0/49 (0%)
Query 6 YTRRADILLKQKRYAAAIRDCDEALKLNPDNARAYRIRGTAHRYLGHWK 54
Y+ RA L K+Y A++DC EALKL+ N +A+ R A++ L +K
Sbjct 230 YSNRALCHLVLKQYKEAVKDCTEALKLDGKNVKAFYRRAQAYKALKDYK 278
Lambda K H
0.318 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2018002440
Database: egene_temp_file_orthology_annotation_similarity_blast_database_865
Posted date: Sep 17, 2011 11:19 AM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40