bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Emax_2858_orf1
Length=93
Score E
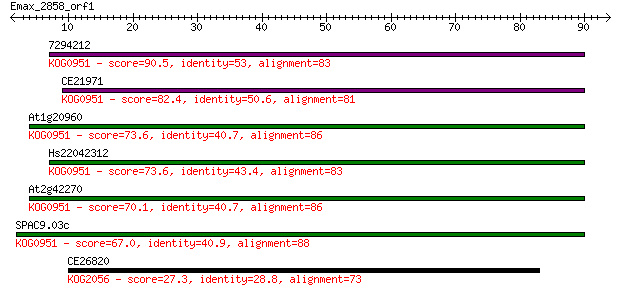
Sequences producing significant alignments: (Bits) Value
7294212 90.5 7e-19
CE21971 82.4 2e-16
At1g20960 73.6 8e-14
Hs22042312 73.6 8e-14
At2g42270 70.1 8e-13
SPAC9.03c 67.0 8e-12
CE26820 27.3 7.7
> 7294212
Length=678
Score = 90.5 bits (223), Expect = 7e-19, Method: Compositional matrix adjust.
Identities = 44/83 (53%), Positives = 57/83 (68%), Gaps = 0/83 (0%)
Query 7 FFYRPTTRVTNQVYEQLLVVLQQQLGDRPDEVLRGAADEVLQVLKEDGLKEGERKRGVEG 66
YRP T+ T Q YE LL +Q+ LGD+P ++L GAADE+L VLK D LK+ ERK+ ++
Sbjct 100 IVYRPKTQETRQTYEVLLSFIQEALGDQPRDILCGAADEILAVLKNDRLKDRERKKDIDS 159
Query 67 VLGPITQERFTKLFQLSKSITDF 89
+LG +T ERF L L K ITDF
Sbjct 160 LLGAVTDERFALLVNLGKKITDF 182
> CE21971
Length=2145
Score = 82.4 bits (202), Expect = 2e-16, Method: Compositional matrix adjust.
Identities = 41/81 (50%), Positives = 53/81 (65%), Gaps = 0/81 (0%)
Query 9 YRPTTRVTNQVYEQLLVVLQQQLGDRPDEVLRGAADEVLQVLKEDGLKEGERKRGVEGVL 68
Y+P T+ T Q YE +L + LGD P EVL GAADEVL LK D ++ E+K+ VE +L
Sbjct 97 YKPRTQETKQTYEVILSFILDALGDVPREVLCGAADEVLLTLKNDKFRDKEKKKEVEALL 156
Query 69 GPITQERFTKLFQLSKSITDF 89
GP+T +R L LSK I+DF
Sbjct 157 GPLTDDRIAVLINLSKKISDF 177
> At1g20960
Length=2171
Score = 73.6 bits (179), Expect = 8e-14, Method: Compositional matrix adjust.
Identities = 35/86 (40%), Positives = 53/86 (61%), Gaps = 0/86 (0%)
Query 4 TENFFYRPTTRVTNQVYEQLLVVLQQQLGDRPDEVLRGAADEVLQVLKEDGLKEGERKRG 63
T++ Y+P T+ T YE +L ++Q+QLG +P ++ GAADE+L VLK D + E+K
Sbjct 105 TDDAVYQPKTKETRAAYEAMLGLIQKQLGGQPPSIVSGAADEILAVLKNDAFRNPEKKME 164
Query 64 VEGVLGPITQERFTKLFQLSKSITDF 89
+E +L I F +L + K ITDF
Sbjct 165 IEKLLNKIENHEFDQLVSIGKLITDF 190
> Hs22042312
Length=2136
Score = 73.6 bits (179), Expect = 8e-14, Method: Compositional matrix adjust.
Identities = 36/83 (43%), Positives = 53/83 (63%), Gaps = 0/83 (0%)
Query 7 FFYRPTTRVTNQVYEQLLVVLQQQLGDRPDEVLRGAADEVLQVLKEDGLKEGERKRGVEG 66
Y+P T+ T + YE LL +Q LGD+P ++L GAADEVL VLK + L++ ER++ ++
Sbjct 100 IIYKPKTKETRETYEVLLSFIQAALGDQPRDILCGAADEVLAVLKNEKLRDKERRKEIDL 159
Query 67 VLGPITQERFTKLFQLSKSITDF 89
+LG R+ L L K ITD+
Sbjct 160 LLGQTDDTRYHVLVNLGKKITDY 182
> At2g42270
Length=2172
Score = 70.1 bits (170), Expect = 8e-13, Method: Compositional matrix adjust.
Identities = 35/86 (40%), Positives = 57/86 (66%), Gaps = 0/86 (0%)
Query 4 TENFFYRPTTRVTNQVYEQLLVVLQQQLGDRPDEVLRGAADEVLQVLKEDGLKEGERKRG 63
TE+ Y+P T+ T +E +L ++QQQLG +P +++ GAADE+L VLK + +K E+K
Sbjct 105 TEDGVYQPKTKETRVAFEIMLGLIQQQLGGQPLDIVCGAADEILAVLKNESVKNHEKKVE 164
Query 64 VEGVLGPITQERFTKLFQLSKSITDF 89
+E +L IT + F++ + K ITD+
Sbjct 165 IEKLLNVITDQVFSQFVSIGKLITDY 190
> SPAC9.03c
Length=2176
Score = 67.0 bits (162), Expect = 8e-12, Method: Compositional matrix adjust.
Identities = 36/89 (40%), Positives = 55/89 (61%), Gaps = 1/89 (1%)
Query 2 DVTENFFYRPTTRVTNQVYEQLLVVLQQQLGDRPDEVLRGAADEVLQVLKEDGLKEGERK 61
D E Y P T T +VY+ +L +QQ LGD+ E+LR AAD ++++LK+ L E RK
Sbjct 124 DSFEILKYNPLTDETREVYDYILSFIQQYLGDQSPEILRSAADLIIELLKDSSLDEQGRK 183
Query 62 RGVEGVLGP-ITQERFTKLFQLSKSITDF 89
+ +E VL + Q+RF++L L +TD+
Sbjct 184 KQIEEVLSTELPQDRFSQLVNLGNRLTDY 212
> CE26820
Length=467
Score = 27.3 bits (59), Expect = 7.7, Method: Composition-based stats.
Identities = 21/77 (27%), Positives = 38/77 (49%), Gaps = 6/77 (7%)
Query 10 RPTTRVTNQVYEQLLVVLQQQLGDRPDEVLRGAADEVLQVLKEDGLKEGER----KRGVE 65
RP+T ++ Q+ + L ++ ++ DE +R + D LQ+ E+ K+ +R K+G
Sbjct 186 RPSTATEEEMQLQIALALSREECEKADE-MRKSDDARLQMALEESQKDADRLATTKQGTV 244
Query 66 GVLGPITQERFTKLFQL 82
G +TQ L L
Sbjct 245 SS-GQLTQSALDDLLSL 260
Lambda K H
0.317 0.137 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1167934574
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40