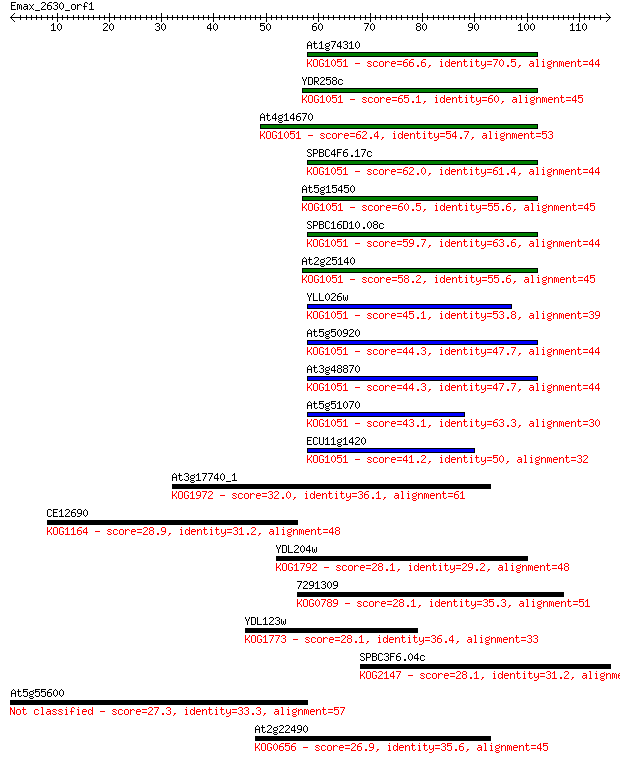
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Emax_2630_orf1
Length=115
Score E
Sequences producing significant alignments: (Bits) Value
At1g74310 66.6 9e-12
YDR258c 65.1 3e-11
At4g14670 62.4 2e-10
SPBC4F6.17c 62.0 3e-10
At5g15450 60.5 8e-10
SPBC16D10.08c 59.7 1e-09
At2g25140 58.2 4e-09
YLL026w 45.1 3e-05
At5g50920 44.3 5e-05
At3g48870 44.3 5e-05
At5g51070 43.1 1e-04
ECU11g1420 41.2 5e-04
At3g17740_1 32.0 0.32
CE12690 28.9 2.7
YDL204w 28.1 4.1
7291309 28.1 4.3
YDL123w 28.1 4.6
SPBC3F6.04c 28.1 4.6
At5g55600 27.3 6.8
At2g22490 26.9 8.4
> At1g74310
Length=911
Score = 66.6 bits (161), Expect = 9e-12, Method: Composition-based stats.
Identities = 31/44 (70%), Positives = 35/44 (79%), Gaps = 0/44 (0%)
Query 58 RYITSRFLPDKAIDLMDEACAIARVQVDSKPEVMDRLERFIFQL 101
RYIT R LPDKAIDL+DEACA RVQ+DS+PE +D LER QL
Sbjct 380 RYITGRHLPDKAIDLVDEACANVRVQLDSQPEEIDNLERKRMQL 423
> YDR258c
Length=811
Score = 65.1 bits (157), Expect = 3e-11, Method: Composition-based stats.
Identities = 27/45 (60%), Positives = 38/45 (84%), Gaps = 0/45 (0%)
Query 57 DRYITSRFLPDKAIDLMDEACAIARVQVDSKPEVMDRLERFIFQL 101
+RYIT RFLPDKAIDL+DEACA+ R+Q +SKP+ + +L+R I ++
Sbjct 313 NRYITDRFLPDKAIDLVDEACAVLRLQHESKPDEIQKLDRAIMKI 357
> At4g14670
Length=623
Score = 62.4 bits (150), Expect = 2e-10, Method: Composition-based stats.
Identities = 29/53 (54%), Positives = 38/53 (71%), Gaps = 0/53 (0%)
Query 49 LCLTKLKFDRYITSRFLPDKAIDLMDEACAIARVQVDSKPEVMDRLERFIFQL 101
L L+ +RYIT R LPDKAIDL+DE+CA + Q+D +PE +D LER + QL
Sbjct 336 LVLSAQLSERYITGRRLPDKAIDLVDESCAHVKAQLDIQPEEIDSLERKVMQL 388
> SPBC4F6.17c
Length=803
Score = 62.0 bits (149), Expect = 3e-10, Method: Composition-based stats.
Identities = 27/44 (61%), Positives = 35/44 (79%), Gaps = 0/44 (0%)
Query 58 RYITSRFLPDKAIDLMDEACAIARVQVDSKPEVMDRLERFIFQL 101
RYIT RFLPDKAIDL+DEAC+ R+Q +SKP+ + RL+R I +
Sbjct 312 RYITDRFLPDKAIDLVDEACSSLRLQQESKPDELRRLDRQIMTI 355
> At5g15450
Length=968
Score = 60.5 bits (145), Expect = 8e-10, Method: Composition-based stats.
Identities = 25/45 (55%), Positives = 35/45 (77%), Gaps = 0/45 (0%)
Query 57 DRYITSRFLPDKAIDLMDEACAIARVQVDSKPEVMDRLERFIFQL 101
DRYI+ RFLPDKAIDL+DEA A ++++ SKP +D L+R + +L
Sbjct 454 DRYISGRFLPDKAIDLVDEAAAKLKMEITSKPTALDELDRSVIKL 498
> SPBC16D10.08c
Length=905
Score = 59.7 bits (143), Expect = 1e-09, Method: Composition-based stats.
Identities = 28/44 (63%), Positives = 34/44 (77%), Gaps = 0/44 (0%)
Query 58 RYITSRFLPDKAIDLMDEACAIARVQVDSKPEVMDRLERFIFQL 101
RY+TSR LPD AIDL+DEA A RV +S+PEV+D LER + QL
Sbjct 383 RYLTSRRLPDSAIDLVDEAAAAVRVTRESQPEVLDNLERKLRQL 426
> At2g25140
Length=874
Score = 58.2 bits (139), Expect = 4e-09, Method: Composition-based stats.
Identities = 25/45 (55%), Positives = 35/45 (77%), Gaps = 0/45 (0%)
Query 57 DRYITSRFLPDKAIDLMDEACAIARVQVDSKPEVMDRLERFIFQL 101
DRYIT RFLPDKAIDL+DEA A ++++ SKP +D ++R + +L
Sbjct 369 DRYITERFLPDKAIDLVDEAGAKLKMEITSKPTELDGIDRAVIKL 413
> YLL026w
Length=908
Score = 45.1 bits (105), Expect = 3e-05, Method: Composition-based stats.
Identities = 21/39 (53%), Positives = 26/39 (66%), Gaps = 0/39 (0%)
Query 58 RYITSRFLPDKAIDLMDEACAIARVQVDSKPEVMDRLER 96
RY+ R LPD A+DL+D +CA V DSKPE +D ER
Sbjct 381 RYLPYRRLPDSALDLVDISCAGVAVARDSKPEELDSKER 419
> At5g50920
Length=929
Score = 44.3 bits (103), Expect = 5e-05, Method: Composition-based stats.
Identities = 21/44 (47%), Positives = 30/44 (68%), Gaps = 0/44 (0%)
Query 58 RYITSRFLPDKAIDLMDEACAIARVQVDSKPEVMDRLERFIFQL 101
+YI+ RFLPDKAIDL+DEA + R++ PE LE+ + Q+
Sbjct 474 QYISDRFLPDKAIDLIDEAGSRVRLRHAQVPEEARELEKELRQI 517
> At3g48870
Length=952
Score = 44.3 bits (103), Expect = 5e-05, Method: Composition-based stats.
Identities = 21/44 (47%), Positives = 30/44 (68%), Gaps = 0/44 (0%)
Query 58 RYITSRFLPDKAIDLMDEACAIARVQVDSKPEVMDRLERFIFQL 101
+YI+ RFLPDKAIDL+DEA + R++ PE LE+ + Q+
Sbjct 495 QYISDRFLPDKAIDLIDEAGSRVRLRHAQLPEEARELEKQLRQI 538
> At5g51070
Length=945
Score = 43.1 bits (100), Expect = 1e-04, Method: Composition-based stats.
Identities = 19/30 (63%), Positives = 23/30 (76%), Gaps = 0/30 (0%)
Query 58 RYITSRFLPDKAIDLMDEACAIARVQVDSK 87
RYI RFLPDKAIDL+DEA + AR++ K
Sbjct 493 RYIADRFLPDKAIDLIDEAGSRARIEAFRK 522
> ECU11g1420
Length=851
Score = 41.2 bits (95), Expect = 5e-04, Method: Composition-based stats.
Identities = 16/32 (50%), Positives = 25/32 (78%), Gaps = 0/32 (0%)
Query 58 RYITSRFLPDKAIDLMDEACAIARVQVDSKPE 89
+YI++R LPD AIDL+D ACA + ++S+P+
Sbjct 366 KYISNRRLPDVAIDLLDTACASVMIALESEPQ 397
> At3g17740_1
Length=640
Score = 32.0 bits (71), Expect = 0.32, Method: Composition-based stats.
Identities = 22/63 (34%), Positives = 32/63 (50%), Gaps = 3/63 (4%)
Query 32 KALHQDMGSSDNCWCWCLCLTKLKFDRYITS--RFLPDKAIDLMDEACAIARVQVDSKPE 89
KAL Q+ +S W LC+ + +F R+ S R L AI + AC+ QVD+ E
Sbjct 362 KALMQN-SASYKLWREFLCVVQGEFSRFKVSEVRRLYSYAIQALSSACSKRHRQVDTTSE 420
Query 90 VMD 92
+D
Sbjct 421 PLD 423
> CE12690
Length=425
Score = 28.9 bits (63), Expect = 2.7, Method: Composition-based stats.
Identities = 15/48 (31%), Positives = 21/48 (43%), Gaps = 5/48 (10%)
Query 8 RSPRSSNIGQCPCRSGSTGACLPLKALHQDMGSSDNCWCWCLCLTKLK 55
R PR + P R GS C P H++ G D+ W W +L+
Sbjct 185 RKPRD----KAPFR-GSDQYCSPRMHNHEEQGRVDDLWSWLYVFVELR 227
> YDL204w
Length=393
Score = 28.1 bits (61), Expect = 4.1, Method: Composition-based stats.
Identities = 14/48 (29%), Positives = 26/48 (54%), Gaps = 0/48 (0%)
Query 52 TKLKFDRYITSRFLPDKAIDLMDEACAIARVQVDSKPEVMDRLERFIF 99
TK+ FD+ + SRF ++ DL+ ++D P + DR+ + +F
Sbjct 83 TKVLFDKGVVSRFGMQESPDLVGVLKPHIDRELDRLPALEDRIRKLVF 130
> 7291309
Length=1377
Score = 28.1 bits (61), Expect = 4.3, Method: Composition-based stats.
Identities = 18/56 (32%), Positives = 29/56 (51%), Gaps = 5/56 (8%)
Query 56 FDRYITSRFLPDKAI---DLMDEACAIARVQVDSKPEVM--DRLERFIFQLAFFVL 106
FDRY+T F+PD + D +D + + V+S + D LE Q +FF++
Sbjct 1076 FDRYLTPTFVPDTVVASSDAIDYEISGRHIGVESVKVTLEGDLLEALPAQGSFFLI 1131
> YDL123w
Length=140
Score = 28.1 bits (61), Expect = 4.6, Method: Compositional matrix adjust.
Identities = 12/33 (36%), Positives = 17/33 (51%), Gaps = 0/33 (0%)
Query 46 CWCLCLTKLKFDRYITSRFLPDKAIDLMDEACA 78
C+C+C T F YI + F P A+ L C+
Sbjct 3 CYCVCCTVSDFILYIVAFFFPPAAVLLRSGPCS 35
> SPBC3F6.04c
Length=827
Score = 28.1 bits (61), Expect = 4.6, Method: Composition-based stats.
Identities = 15/52 (28%), Positives = 26/52 (50%), Gaps = 4/52 (7%)
Query 68 KAIDLMDEACAIARVQ----VDSKPEVMDRLERFIFQLAFFVLHISPMPLIT 115
K +D D ++R++ V P+ RLE F L +LH++ P+I+
Sbjct 424 KGLDYKDYPTVVSRIRTLHHVKLHPDNKSRLENFSVILLQHILHLTRQPMIS 475
> At5g55600
Length=581
Score = 27.3 bits (59), Expect = 6.8, Method: Composition-based stats.
Identities = 19/63 (30%), Positives = 28/63 (44%), Gaps = 7/63 (11%)
Query 1 RLKRSICRSPRSSNIGQCPCRSGSTGACLPLKALHQDMGSSDNCWCWCLCL------TKL 54
+L S R P +N+ + PC +G ++ L QD G CW C L KL
Sbjct 275 KLNASGKRFPSPTNVQKHPCYNGVVKTDAKIEFLCQDSGIR-GCWFRCTVLDVSRKQVKL 333
Query 55 KFD 57
++D
Sbjct 334 QYD 336
> At2g22490
Length=361
Score = 26.9 bits (58), Expect = 8.4, Method: Composition-based stats.
Identities = 16/45 (35%), Positives = 25/45 (55%), Gaps = 1/45 (2%)
Query 48 CLCLTKLKFDRYITSRFLPDKAIDLMDEACAIARVQVDSKPEVMD 92
C+CL+ DR++TS LP K D + A++ + + SK E D
Sbjct 117 CICLSMNYLDRFLTSYELP-KDKDWAAQLLAVSCLSLASKMEETD 160
Lambda K H
0.329 0.140 0.466
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1178471386
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40