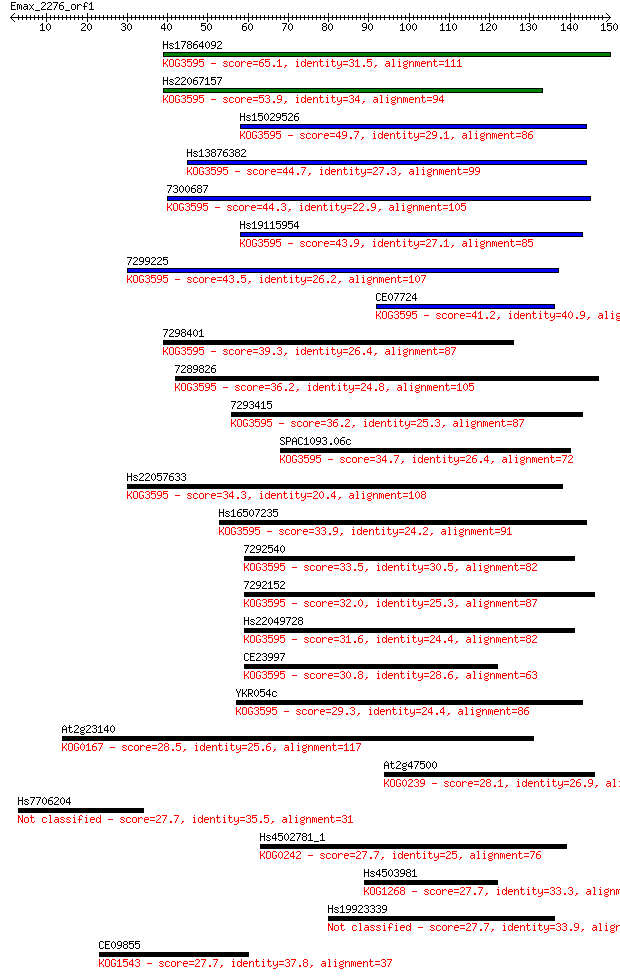
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Emax_2276_orf1
Length=149
Score E
Sequences producing significant alignments: (Bits) Value
Hs17864092 65.1 5e-11
Hs22067157 53.9 1e-07
Hs15029526 49.7 2e-06
Hs13876382 44.7 7e-05
7300687 44.3 1e-04
Hs19115954 43.9 1e-04
7299225 43.5 1e-04
CE07724 41.2 8e-04
7298401 39.3 0.003
7289826 36.2 0.026
7293415 36.2 0.026
SPAC1093.06c 34.7 0.072
Hs22057633 34.3 0.10
Hs16507235 33.9 0.12
7292540 33.5 0.14
7292152 32.0 0.48
Hs22049728 31.6 0.66
CE23997 30.8 0.95
YKR054c 29.3 2.7
At2g23140 28.5 5.2
At2g47500 28.1 6.2
Hs7706204 27.7 8.3
Hs4502781_1 27.7 8.5
Hs4503981 27.7 8.8
Hs19923339 27.7 8.9
CE09855 27.7 9.4
> Hs17864092
Length=4024
Score = 65.1 bits (157), Expect = 5e-11, Method: Compositional matrix adjust.
Identities = 35/111 (31%), Positives = 61/111 (54%), Gaps = 0/111 (0%)
Query 39 CADIFESTKNIADCYRVEQRRFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQ 98
C ST +++ + VE +R+ +VTP S+L + F + + S+ +++YE+GL+
Sbjct 2534 CKSFHTSTIDLSKSFFVELQRYNYVTPTSYLELISTFKLLLEKKRSEVMKMKKRYEVGLE 2593
Query 99 KLHDVSMQVMDMQQQLEKLQPELAKSSMETQQLMQVLTVKQEHAATTMSLV 149
KL S QV MQ +LE L P+L +S E ++M ++ + A T +V
Sbjct 2594 KLDSASSQVATMQMELEALHPQLKVASKEVDEMMIMIEKESVEVAKTEKIV 2644
> Hs22067157
Length=3672
Score = 53.9 bits (128), Expect = 1e-07, Method: Composition-based stats.
Identities = 32/94 (34%), Positives = 51/94 (54%), Gaps = 0/94 (0%)
Query 39 CADIFESTKNIADCYRVEQRRFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQ 98
C ES K ++ Y + RR +VTP S+L + F + ++ + + +Y GLQ
Sbjct 2181 CKYFQESVKKLSLDYYNKLRRHNYVTPTSYLELILTFKTLLNSKRQEVAMMRNRYLTGLQ 2240
Query 99 KLHDVSMQVMDMQQQLEKLQPELAKSSMETQQLM 132
KL + QV MQ++L LQP+L +S ET ++M
Sbjct 2241 KLDFAASQVAVMQRELTALQPQLILTSEETAKMM 2274
> Hs15029526
Length=4490
Score = 49.7 bits (117), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 25/86 (29%), Positives = 49/86 (56%), Gaps = 0/86 (0%)
Query 58 RRFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQKLHDVSMQVMDMQQQLEKL 117
RR HVTP S+L F++G+ ++ + + + ++ +GL KL + S V + Q L
Sbjct 3032 RRRAHVTPKSYLSFINGYKNIYAEKVKFINEQAERMNIGLDKLMEASESVAKLSQDLAVK 3091
Query 118 QPELAKSSMETQQLMQVLTVKQEHAA 143
+ ELA +S++ +++ +TV + +A
Sbjct 3092 EKELAVASIKADEVLAEVTVSAQASA 3117
> Hs13876382
Length=4486
Score = 44.7 bits (104), Expect = 7e-05, Method: Compositional matrix adjust.
Identities = 27/99 (27%), Positives = 51/99 (51%), Gaps = 0/99 (0%)
Query 45 STKNIADCYRVEQRRFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQKLHDVS 104
S + Y ++R+ + TP SFL F+ + + + K ++ E GL KLH S
Sbjct 3021 SVNQTSQSYLSNEQRYNYTTPKSFLEFIRLYQSLLHRHRKELKCKTERLENGLLKLHSTS 3080
Query 105 MQVMDMQQQLEKLQPELAKSSMETQQLMQVLTVKQEHAA 143
QV D++ +L + EL + + + +L+QV+ V+ + +
Sbjct 3081 AQVDDLKAKLAAQEVELKQKNEDADKLIQVVGVETDKVS 3119
> 7300687
Length=4472
Score = 44.3 bits (103), Expect = 1e-04, Method: Composition-based stats.
Identities = 24/105 (22%), Positives = 52/105 (49%), Gaps = 0/105 (0%)
Query 40 ADIFESTKNIADCYRVEQRRFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQK 99
A + S + Y +RR+ + TP S+L ++ + + + K ++ E GL+K
Sbjct 3001 AYVHTSVNTTSKVYLQNERRYNYTTPKSYLEQINLYIKLLNHKNEDLQSKIERLENGLEK 3060
Query 100 LHDVSMQVMDMQQQLEKLQPELAKSSMETQQLMQVLTVKQEHAAT 144
L ++QV D++ +L + EL + + L++++ ++ E T
Sbjct 3061 LRSTALQVADLKVKLAVQEIELKEKNEAADALIEIVGIETEKVQT 3105
> Hs19115954
Length=4624
Score = 43.9 bits (102), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 23/85 (27%), Positives = 47/85 (55%), Gaps = 0/85 (0%)
Query 58 RRFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQKLHDVSMQVMDMQQQLEKL 117
RR HVTP S+L F+ G+ +++ + + + GL+KL + S V + ++LE
Sbjct 3168 RRSTHVTPKSYLSFIQGYKFIYGEKHVEVRTLANRMNTGLEKLKEASESVAALSKELEAK 3227
Query 118 QPELAKSSMETQQLMQVLTVKQEHA 142
+ EL ++ + +++ +T+K + A
Sbjct 3228 EKELQVANDKADMVLKEVTMKAQAA 3252
> 7299225
Length=3508
Score = 43.5 bits (101), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 28/107 (26%), Positives = 54/107 (50%), Gaps = 3/107 (2%)
Query 30 ENLKHLCGACADIFESTKNIADCYRVEQRRFFHVTPASFLRFLDGFCYVFMTRASQHHHK 89
E L + G+ DI T + Y RR HVTP S+L F+ G+ ++ + +
Sbjct 3130 EELVNALGSIQDIVAET---SQEYFQRFRRATHVTPKSYLNFIAGYKNIYQMKQQELRDG 3186
Query 90 QQQYELGLQKLHDVSMQVMDMQQQLEKLQPELAKSSMETQQLMQVLT 136
++ + GL+KL + S V +++ L ++ EL ++S + ++ +T
Sbjct 3187 VEKMDTGLEKLKEASASVEILKKDLVVMEEELVEASKNAESVLVEVT 3233
> CE07724
Length=2632
Score = 41.2 bits (95), Expect = 8e-04, Method: Composition-based stats.
Identities = 18/44 (40%), Positives = 29/44 (65%), Gaps = 0/44 (0%)
Query 92 QYELGLQKLHDVSMQVMDMQQQLEKLQPELAKSSMETQQLMQVL 135
+YE G++K+ QV MQ +L +LQP+L ++S+ET LM +
Sbjct 2069 KYEKGMEKMKRAEEQVAGMQGELLRLQPQLVRTSIETSMLMSTI 2112
> 7298401
Length=4010
Score = 39.3 bits (90), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 23/87 (26%), Positives = 46/87 (52%), Gaps = 0/87 (0%)
Query 39 CADIFESTKNIADCYRVEQRRFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQ 98
C + ST+ ++ + R+ +VTP S+L + F + + + + + +Y G+
Sbjct 2501 CMEFHTSTQELSAKFFSRLHRYNYVTPTSYLELIQTFKALLSQKRNNITNNRNRYLTGIS 2560
Query 99 KLHDVSMQVMDMQQQLEKLQPELAKSS 125
+L + QV MQ+QL L+P+L ++S
Sbjct 2561 QLDIAAQQVAVMQEQLIALEPKLKEAS 2587
> 7289826
Length=507
Score = 36.2 bits (82), Expect = 0.026, Method: Composition-based stats.
Identities = 26/105 (24%), Positives = 48/105 (45%), Gaps = 2/105 (1%)
Query 42 IFESTKNIADCYRVEQRRFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQKLH 101
I ES + Y RR VTP S + FL+ + ++ + ++ GL KL
Sbjct 330 IHESVQETCLSYYDRFRRVTFVTPKSLISFLESYKLLYKDKQDHIVIMSERMSSGLDKLD 389
Query 102 DVSMQVMDMQQQLEKLQPELAKSSMETQQLMQVLTVKQEHAATTM 146
+ V +++ L ++ +A +S E + ++ TV+Q AA +
Sbjct 390 EAGASVAILKKDLIEMNKVIALASEEAEDVLA--TVEQSKAAAEI 432
> 7293415
Length=4081
Score = 36.2 bits (82), Expect = 0.026, Method: Compositional matrix adjust.
Identities = 22/87 (25%), Positives = 46/87 (52%), Gaps = 0/87 (0%)
Query 56 EQRRFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQKLHDVSMQVMDMQQQLE 115
E +R ++ TP+S+L L + + + + K+++ GL KL + + + M ++LE
Sbjct 2590 EMKRHYYTTPSSYLELLKLYQNLLKIKNMEIIAKRKRIANGLNKLLETNEVIAVMGKELE 2649
Query 116 KLQPELAKSSMETQQLMQVLTVKQEHA 142
+ P+L + S + L+ LT + + A
Sbjct 2650 VMVPQLDEKSAMMKSLVDNLTKETKQA 2676
> SPAC1093.06c
Length=1889
Score = 34.7 bits (78), Expect = 0.072, Method: Composition-based stats.
Identities = 19/73 (26%), Positives = 38/73 (52%), Gaps = 1/73 (1%)
Query 68 FLRFLDGFCYVFMTRASQHHHKQQQYELGLQKLHDVSMQVMDMQQQLEKLQPEL-AKSSM 126
F+RFL+ FC +F A++ ++ + E G +K+ + S + ++ L Q L +K+
Sbjct 783 FIRFLNTFCLIFGRDANKLSKEKSRIENGFKKIKETSQGIDKFKEALSDQQNVLFSKTKT 842
Query 127 ETQQLMQVLTVKQ 139
+L ++ KQ
Sbjct 843 ANDRLQCIIQTKQ 855
> Hs22057633
Length=1770
Score = 34.3 bits (77), Expect = 0.10, Method: Compositional matrix adjust.
Identities = 22/108 (20%), Positives = 59/108 (54%), Gaps = 0/108 (0%)
Query 30 ENLKHLCGACADIFESTKNIADCYRVEQRRFFHVTPASFLRFLDGFCYVFMTRASQHHHK 89
EN++++ + +S + + + + RR +VTP ++L F++ + + + + +
Sbjct 1414 ENIENVVKHVVLVHQSVDHYSQQFLQKLRRSNYVTPKNYLDFINTYSKLLDEKTQCNIAQ 1473
Query 90 QQQYELGLQKLHDVSMQVMDMQQQLEKLQPELAKSSMETQQLMQVLTV 137
++ + GL KL + ++Q+ ++ Q+L + + LA+ S + L++ + V
Sbjct 1474 CKRLDGGLDKLKEATIQLDELNQKLAEQKIVLAEKSAACEALLEEIAV 1521
> Hs16507235
Length=4523
Score = 33.9 bits (76), Expect = 0.12, Method: Compositional matrix adjust.
Identities = 22/91 (24%), Positives = 46/91 (50%), Gaps = 0/91 (0%)
Query 53 YRVEQRRFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQKLHDVSMQVMDMQQ 112
Y +RR + TP SFL + F + + ++ K+++ G+QKL + QV D++
Sbjct 3066 YYQNERRHNYTTPKSFLEQISLFKNLLKKKQNEVSEKKERLVNGIQKLKTTASQVGDLKA 3125
Query 113 QLEKLQPELAKSSMETQQLMQVLTVKQEHAA 143
+L + EL + + + L+ + ++ E +
Sbjct 3126 RLASQEAELQLRNHDAEALITKIGLQTEKVS 3156
> 7292540
Length=4680
Score = 33.5 bits (75), Expect = 0.14, Method: Compositional matrix adjust.
Identities = 25/83 (30%), Positives = 41/83 (49%), Gaps = 1/83 (1%)
Query 59 RFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQKLHDVSMQVMDMQQQLEKLQ 118
R VTP +L F+ F ++ + S +Q +GL K+ + QV +MQ+ L +
Sbjct 3136 RTMAVTPRHYLDFIHHFVKLYNEKRSDLEEQQLHLNVGLNKIAETVEQVEEMQKSLAVKK 3195
Query 119 PEL-AKSSMETQQLMQVLTVKQE 140
EL AK+ +L Q+ +QE
Sbjct 3196 QELQAKNEAANAKLKQMFQDQQE 3218
> 7292152
Length=3868
Score = 32.0 bits (71), Expect = 0.48, Method: Compositional matrix adjust.
Identities = 22/87 (25%), Positives = 42/87 (48%), Gaps = 2/87 (2%)
Query 59 RFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQKLHDVSMQVMDMQQQLEKLQ 118
R + T ASF+ + F + + S+ + +Y GL L + + MQ+ L LQ
Sbjct 2493 RHIYQTNASFIELIRSFQTLIERKQSETMLAKMRYIGGLDTLAQAAAAISIMQRDLNALQ 2552
Query 119 PELAKSSMETQQLMQVLTVKQEHAATT 145
P+L + ++++M L + +E A +
Sbjct 2553 PKLVALAESSRKMM--LEINKETLAAS 2577
> Hs22049728
Length=2274
Score = 31.6 bits (70), Expect = 0.66, Method: Compositional matrix adjust.
Identities = 20/83 (24%), Positives = 44/83 (53%), Gaps = 1/83 (1%)
Query 59 RFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQKLHDVSMQVMDMQQQLEKLQ 118
R +TP +L F++ + +F + S+ +Q +GL+K+ + QV ++++ L
Sbjct 795 RTMAITPRHYLDFINHYANLFHEKRSELEEQQMHLNVGLRKIKETVDQVEELRRDLRIKS 854
Query 119 PEL-AKSSMETQQLMQVLTVKQE 140
EL K++ +L +++ +QE
Sbjct 855 QELEVKNAAANDKLKKMVKDQQE 877
> CE23997
Length=4568
Score = 30.8 bits (68), Expect = 0.95, Method: Compositional matrix adjust.
Identities = 18/63 (28%), Positives = 30/63 (47%), Gaps = 0/63 (0%)
Query 59 RFFHVTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQKLHDVSMQVMDMQQQLEKLQ 118
R TP FL F+ F +F + S ++ +GL K+ + QV ++Q+ L+
Sbjct 3110 RVMACTPRHFLDFIKQFMSLFHEKRSDLEEEKIHLNIGLNKISETEEQVKELQKSLKLKS 3169
Query 119 PEL 121
EL
Sbjct 3170 NEL 3172
> YKR054c
Length=4092
Score = 29.3 bits (64), Expect = 2.7, Method: Composition-based stats.
Identities = 21/89 (23%), Positives = 44/89 (49%), Gaps = 3/89 (3%)
Query 57 QRRFFHVTPASFLRFLDGFCYV--FMTRASQHHHKQQQY-ELGLQKLHDVSMQVMDMQQQ 113
Q+ V P S F+DG + +T Q + Q++ +GL+KL++ ++V ++ +
Sbjct 2978 QKMKVGVNPRSPGYFIDGLRALVKLVTAKYQDLQENQRFVNVGLEKLNESVLKVNELNKT 3037
Query 114 LEKLQPELAKSSMETQQLMQVLTVKQEHA 142
L K EL + E + + + ++Q +
Sbjct 3038 LSKKSTELTEKEKEARSTLDKMLMEQNES 3066
> At2g23140
Length=924
Score = 28.5 bits (62), Expect = 5.2, Method: Composition-based stats.
Identities = 30/122 (24%), Positives = 56/122 (45%), Gaps = 11/122 (9%)
Query 14 ADGTKESQLYPSADSNENLKHLCGACADIFESTKNIADCYRVEQRRFFHVTPASFLRFLD 73
A T ++ +P AD+NEN + A +++ I T A+ R L
Sbjct 523 ATSTVSNEEFPRADANENSEESAHATPYSSDASGEI------RSGPLAATTSAATRRDLS 576
Query 74 GFCYVFMTRASQHHHKQQQYE-LGLQKL----HDVSMQVMDMQQQLEKLQPELAKSSMET 128
F FM R ++ ++ E LG + + ++ + +++ Q++KL EL SS++T
Sbjct 577 DFSPKFMDRRTRGQFWRRPSERLGSRIVSAPSNETRRDLSEVETQVKKLVEELKSSSLDT 636
Query 129 QQ 130
Q+
Sbjct 637 QR 638
> At2g47500
Length=861
Score = 28.1 bits (61), Expect = 6.2, Method: Composition-based stats.
Identities = 14/54 (25%), Positives = 32/54 (59%), Gaps = 2/54 (3%)
Query 94 ELGLQKLHDVSMQVMDMQQQLEKLQPELAKSSMETQQ--LMQVLTVKQEHAATT 145
ELG ++++ + V ++++Q+ L+ LA+ E+QQ +++ ++H A T
Sbjct 643 ELGAARVNNDTSDVKELKEQIATLKAALARKEAESQQNNILKTPGGSEKHKAKT 696
> Hs7706204
Length=555
Score = 27.7 bits (60), Expect = 8.3, Method: Composition-based stats.
Identities = 11/31 (35%), Positives = 20/31 (64%), Gaps = 0/31 (0%)
Query 3 DNALPSIQEKMADGTKESQLYPSADSNENLK 33
+N LP+++EK G+K+ + D+N+ LK
Sbjct 314 ENGLPTVREKTRAGSKKCRFPDDLDTNKKLK 344
> Hs4502781_1
Length=731
Score = 27.7 bits (60), Expect = 8.5, Method: Compositional matrix adjust.
Identities = 19/76 (25%), Positives = 34/76 (44%), Gaps = 0/76 (0%)
Query 63 VTPASFLRFLDGFCYVFMTRASQHHHKQQQYELGLQKLHDVSMQVMDMQQQLEKLQPELA 122
+TP SF L + + ++ + L ++MD+++QLE++ E
Sbjct 308 ITPVSFDETLTALQFASTAKYMKNTPYVNEVSTDEALLKRYRKEIMDLKKQLEEVSLETR 367
Query 123 KSSMETQQLMQVLTVK 138
+ME QL Q+L K
Sbjct 368 AQAMEKDQLAQLLEEK 383
> Hs4503981
Length=681
Score = 27.7 bits (60), Expect = 8.8, Method: Compositional matrix adjust.
Identities = 11/33 (33%), Positives = 23/33 (69%), Gaps = 0/33 (0%)
Query 89 KQQQYELGLQKLHDVSMQVMDMQQQLEKLQPEL 121
++++ LGL++L D+ +V+ M +++KL EL
Sbjct 502 RRKEIMLGLKRLPDLIKEVLSMDDEIQKLATEL 534
> Hs19923339
Length=3908
Score = 27.7 bits (60), Expect = 8.9, Method: Compositional matrix adjust.
Identities = 19/61 (31%), Positives = 35/61 (57%), Gaps = 5/61 (8%)
Query 80 MTRASQHHHKQQQYEL-----GLQKLHDVSMQVMDMQQQLEKLQPELAKSSMETQQLMQV 134
M + Q H K Q E G +K+H++ +V D+Q+QLE+ + ++ K +E Q+L +
Sbjct 3381 MEKDRQVHRKTLQTEQEANTEGQKKMHELQSKVEDLQRQLEEKRQQVYKLDLEGQRLQGI 3440
Query 135 L 135
+
Sbjct 3441 M 3441
> CE09855
Length=427
Score = 27.7 bits (60), Expect = 9.4, Method: Compositional matrix adjust.
Identities = 14/37 (37%), Positives = 22/37 (59%), Gaps = 3/37 (8%)
Query 23 YPSADSNENLKHLCGACADIFESTKNIADCYRVEQRR 59
Y SAD N+++ CG+C F +T +AD R+ +R
Sbjct 200 YASADRNQHIPQYCGSCW-AFGATSALAD--RINIKR 233
Lambda K H
0.320 0.129 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1858150626
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40