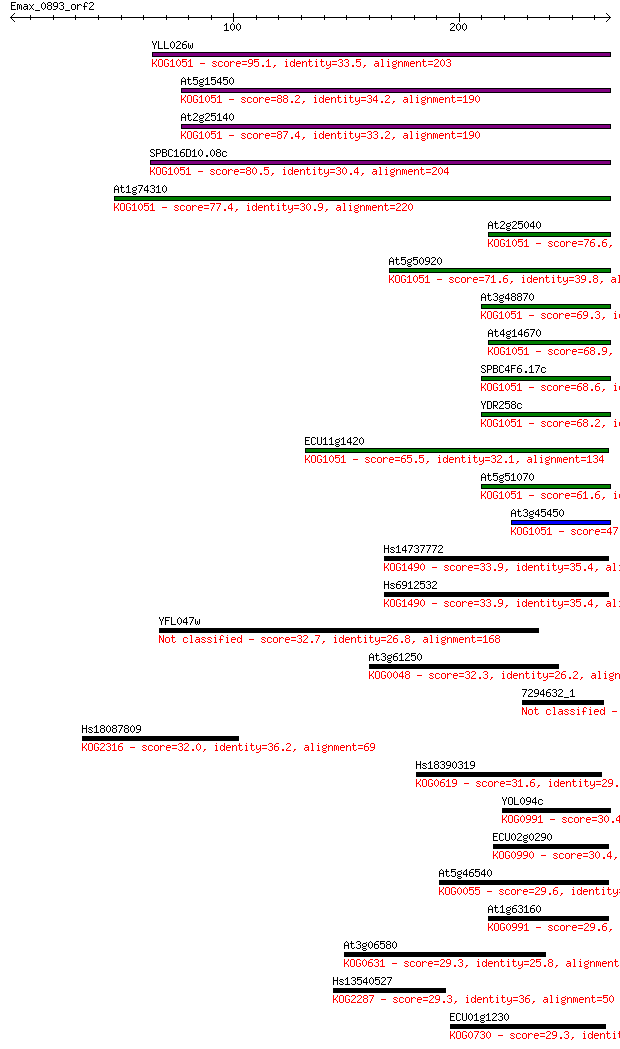
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Emax_0893_orf2
Length=266
Score E
Sequences producing significant alignments: (Bits) Value
YLL026w 95.1 1e-19
At5g15450 88.2 2e-17
At2g25140 87.4 2e-17
SPBC16D10.08c 80.5 3e-15
At1g74310 77.4 3e-14
At2g25040 76.6 5e-14
At5g50920 71.6 2e-12
At3g48870 69.3 8e-12
At4g14670 68.9 1e-11
SPBC4F6.17c 68.6 1e-11
YDR258c 68.2 2e-11
ECU11g1420 65.5 1e-10
At5g51070 61.6 2e-09
At3g45450 47.4 4e-05
Hs14737772 33.9 0.35
Hs6912532 33.9 0.36
YFL047w 32.7 0.78
At3g61250 32.3 1.0
7294632_1 32.0 1.4
Hs18087809 32.0 1.4
Hs18390319 31.6 1.6
YOL094c 30.4 3.6
ECU02g0290 30.4 3.8
At5g46540 29.6 6.0
At1g63160 29.6 6.8
At3g06580 29.3 8.3
Hs13540527 29.3 9.3
ECU01g1230 29.3 9.6
> YLL026w
Length=908
Score = 95.1 bits (235), Expect = 1e-19, Method: Compositional matrix adjust.
Identities = 68/214 (31%), Positives = 110/214 (51%), Gaps = 22/214 (10%)
Query 64 LAWHLHHDRLALAHLIMAMLEGSRLETPRE--------VITKCHGDFSQVRGVIKAVLVR 115
LA H +L H++ A +E TP + +I K D+ + V+ LVR
Sbjct 21 LASDHQHPQLQPIHILAAFIE-----TPEDGSVPYLQNLIEKGRYDYDLFKKVVNRNLVR 75
Query 116 LPRTPVTVPLGGAKVTP--AVTEVFLRARRI-AEAEGSLTGMQHLFLALLYCPTFYKILV 172
+P+ P A++TP A+ +V A +I + + S H+ AL + +I
Sbjct 76 IPQQQ-PAP---AEITPSYALGKVLQDAAKIQKQQKDSFIAQDHILFALFNDSSIQQIFK 131
Query 173 NSGCDIELFKEELALAYKNTPMPNTTGTTEQLTKDFDITQYGEDLTEKAEKGKLQPVIGR 232
+ DIE K++ AL + ++ G ++ +++Y D+TE+A +GKL PVIGR
Sbjct 132 EAQVDIEAIKQQ-ALELRGNTRIDSRGADTNTPLEY-LSKYAIDMTEQARQGKLDPVIGR 189
Query 233 DEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIE 266
+EEI ++L+R + P LIGEPG+GKTA+IE
Sbjct 190 EEEIRSTIRVLARRIKSNPCLIGEPGIGKTAIIE 223
> At5g15450
Length=968
Score = 88.2 bits (217), Expect = 2e-17, Method: Compositional matrix adjust.
Identities = 65/191 (34%), Positives = 98/191 (51%), Gaps = 7/191 (3%)
Query 77 HLIMAMLEGSRLETPREVITKCHGDFSQVRGVIKAVLVRLPRTPVTVPLGGAKVTPAVTE 136
HL+ A+LE + R + +K D ++V + + R P+ V G+ + +
Sbjct 109 HLMKALLE-QKNGLARRIFSKIGVDNTKVLEATEKFIQRQPK--VYGDAAGSMLGRDLEA 165
Query 137 VFLRARRIA-EAEGSLTGMQHLFLALLYCPTFYKILVNSGCDIELFKEELALAYKNTPMP 195
+F RAR+ + + S ++HL LA F K L D ++ + L A ++
Sbjct 166 LFQRARQFKKDLKDSYVSVEHLVLAFADDKRFGKQLFK---DFQISERSLKSAIESIRGK 222
Query 196 NTTGTTEQLTKDFDITQYGEDLTEKAEKGKLQPVIGRDEEIDRIAQILSRMMTKAPLLIG 255
+ + K + +YG+DLT A +GKL PVIGRD+EI R QILSR P+LIG
Sbjct 223 QSVIDQDPEGKYEALEKYGKDLTAMAREGKLDPVIGRDDEIRRCIQILSRRTKNNPVLIG 282
Query 256 EPGVGKTAVIE 266
EPGVGKTA+ E
Sbjct 283 EPGVGKTAISE 293
> At2g25140
Length=874
Score = 87.4 bits (215), Expect = 2e-17, Method: Compositional matrix adjust.
Identities = 63/193 (32%), Positives = 98/193 (50%), Gaps = 12/193 (6%)
Query 77 HLIMAMLEGSRLETPREVITKCHGDFSQVRGVIKAVLVRLPRTPVTVPLGGAKVTPAVTE 136
HL+ A+LE + R++ TK D S V++A + + + P G ++ +++
Sbjct 25 HLMKALLE-QKDGMARKIFTKAGIDNS---SVLQATDLFISKQPTVSDASGQRLGSSLSV 80
Query 137 VFLRARR-IAEAEGSLTGMQHLFLALLYCPTFY--KILVNSGCDIELFKEELALAYKNTP 193
+ A+R + S ++H LA Y T + + + DI++ K+ A K+
Sbjct 81 ILENAKRHKKDMLDSYVSVEHFLLAY-YSDTRFGQEFFRDMKLDIQVLKD----AIKDVR 135
Query 194 MPNTTGTTEQLTKDFDITQYGEDLTEKAEKGKLQPVIGRDEEIDRIAQILSRMMTKAPLL 253
+K + +YG DLTE A +GKL PVIGRD+EI R QIL R P++
Sbjct 136 GDQRVTDRNPESKYQALEKYGNDLTEMARRGKLDPVIGRDDEIRRCIQILCRRTKNNPVI 195
Query 254 IGEPGVGKTAVIE 266
IGEPGVGKTA+ E
Sbjct 196 IGEPGVGKTAIAE 208
> SPBC16D10.08c
Length=905
Score = 80.5 bits (197), Expect = 3e-15, Method: Compositional matrix adjust.
Identities = 62/210 (29%), Positives = 98/210 (46%), Gaps = 14/210 (6%)
Query 63 TLAWHLHHDRLALAHLIMAMLEGSRLETP---REVITKCHGDFSQVRGVIKAVLVRLP-R 118
++A H +L H+ A+L S R ++ K GD + + + LVRLP +
Sbjct 19 SIAQSYGHSQLTPIHIAAALLSDSDSNGTTLLRTIVDKAGGDGQKFERSVTSRLVRLPAQ 78
Query 119 TPVTVPLGGAKVTPAVTEVFLRARRIAEAE-GSLTGMQHLFLALLYCPTFYKILVNSGCD 177
P P ++P ++ A + + + S H T +L +G
Sbjct 79 DP---PPEQVTLSPESAKLLRNAHELQKTQKDSYIAQDHFIAVFTKDDTLKSLLAEAGVT 135
Query 178 IELFKEELALAYKNTPMPNTTGTTEQLTKDFD-ITQYGEDLTEKAEKGKLQPVIGRDEEI 236
+ F E A+ N N ++ + FD + ++ DLTE A G+L PVIGR++EI
Sbjct 136 PKAF--EFAV---NNVRGNKRIDSKNAEEGFDALNKFTVDLTELARNGQLDPVIGREDEI 190
Query 237 DRIAQILSRMMTKAPLLIGEPGVGKTAVIE 266
R ++LSR P+LIGEPGVGKT++ E
Sbjct 191 RRTIRVLSRRTKNNPVLIGEPGVGKTSIAE 220
> At1g74310
Length=911
Score = 77.4 bits (189), Expect = 3e-14, Method: Compositional matrix adjust.
Identities = 68/228 (29%), Positives = 103/228 (45%), Gaps = 20/228 (8%)
Query 47 PSELNFDAQLMVETMDTLAWHLHHDRLALAHLIMAMLEGSRLETPREVITKCHGDFSQ-V 105
P + + T LA + H + HL A++ P+ + + + +Q
Sbjct 3 PEKFTHKTNETIATAHELAVNAGHAQFTPLHLAGALISDPTGIFPQAISSAGGENAAQSA 62
Query 106 RGVIKAVLVRLPRT---PVTVPLGGAKVTPAVTEVFLRARRIAEAEGSLT-GMQHLFLAL 161
VI L +LP P +P + ++ +V RA+ ++ G + L + L
Sbjct 63 ERVINQALKKLPSQSPPPDDIP-----ASSSLIKVIRRAQAAQKSRGDTHLAVDQLIMGL 117
Query 162 LYCPTFYKILVNSGCDIELFKEELA--LAYKNTPMPNTTGTTEQLTKDFD-ITQYGEDLT 218
L +L G K E+ + + + +G T +F + YG DL
Sbjct 118 LEDSQIRDLLNEVGVATARVKSEVEKLRGKEGKKVESASGDT-----NFQALKTYGRDLV 172
Query 219 EKAEKGKLQPVIGRDEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIE 266
E+A GKL PVIGRDEEI R+ +ILSR P+LIGEPGVGKTAV+E
Sbjct 173 EQA--GKLDPVIGRDEEIRRVVRILSRRTKNNPVLIGEPGVGKTAVVE 218
> At2g25040
Length=135
Score = 76.6 bits (187), Expect = 5e-14, Method: Compositional matrix adjust.
Identities = 33/54 (61%), Positives = 43/54 (79%), Gaps = 0/54 (0%)
Query 213 YGEDLTEKAEKGKLQPVIGRDEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIE 266
YG DLT+ A +GKL P+IGRD+E++R QIL RM P++IGEPGVGKTA++E
Sbjct 72 YGSDLTKMARQGKLPPLIGRDDEVNRCIQILCRMTESNPVIIGEPGVGKTAIVE 125
> At5g50920
Length=929
Score = 71.6 bits (174), Expect = 2e-12, Method: Compositional matrix adjust.
Identities = 39/99 (39%), Positives = 57/99 (57%), Gaps = 1/99 (1%)
Query 169 KILVNSGCDIELFKEE-LALAYKNTPMPNTTGTTEQLTKDFDITQYGEDLTEKAEKGKLQ 227
++L N G D + + + + +N + G K + +YG +LT+ AE+GKL
Sbjct 215 RVLENLGADPSNIRTQVIRMVGENNEVTANVGGGSSSNKMPTLEEYGTNLTKLAEEGKLD 274
Query 228 PVIGRDEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIE 266
PV+GR +I+R+ QIL R P LIGEPGVGKTA+ E
Sbjct 275 PVVGRQPQIERVVQILGRRTKNNPCLIGEPGVGKTAIAE 313
> At3g48870
Length=952
Score = 69.3 bits (168), Expect = 8e-12, Method: Compositional matrix adjust.
Identities = 32/57 (56%), Positives = 43/57 (75%), Gaps = 0/57 (0%)
Query 210 ITQYGEDLTEKAEKGKLQPVIGRDEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIE 266
+ +YG +LT+ AE+GKL PV+GR +I+R+ QIL+R P LIGEPGVGKTA+ E
Sbjct 278 LEEYGTNLTKLAEEGKLDPVVGRQPQIERMVQILARRTKNNPCLIGEPGVGKTAIAE 334
> At4g14670
Length=623
Score = 68.9 bits (167), Expect = 1e-11, Method: Compositional matrix adjust.
Identities = 34/54 (62%), Positives = 40/54 (74%), Gaps = 2/54 (3%)
Query 213 YGEDLTEKAEKGKLQPVIGRDEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIE 266
YG DL E+A GKL PVIGR EI R+ ++LSR P+LIGEPGVGKTAV+E
Sbjct 132 YGTDLVEQA--GKLDPVIGRHREIRRVIEVLSRRTKNNPVLIGEPGVGKTAVVE 183
> SPBC4F6.17c
Length=803
Score = 68.6 bits (166), Expect = 1e-11, Method: Compositional matrix adjust.
Identities = 33/57 (57%), Positives = 41/57 (71%), Gaps = 0/57 (0%)
Query 210 ITQYGEDLTEKAEKGKLQPVIGRDEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIE 266
+ +YG DLT A++GKL PVIGR+EEI R QILSR P L+G GVGKTA++E
Sbjct 94 LEEYGTDLTALAKQGKLDPVIGREEEIQRTIQILSRRTKNNPALVGPAGVGKTAIME 150
> YDR258c
Length=811
Score = 68.2 bits (165), Expect = 2e-11, Method: Compositional matrix adjust.
Identities = 34/57 (59%), Positives = 40/57 (70%), Gaps = 0/57 (0%)
Query 210 ITQYGEDLTEKAEKGKLQPVIGRDEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIE 266
+ Q+G +LT+ A GKL PVIGRDEEI R QILSR P LIG GVGKTA+I+
Sbjct 98 LEQFGTNLTKLARDGKLDPVIGRDEEIARAIQILSRRTKNNPCLIGRAGVGKTALID 154
> ECU11g1420
Length=851
Score = 65.5 bits (158), Expect = 1e-10, Method: Compositional matrix adjust.
Identities = 43/138 (31%), Positives = 71/138 (51%), Gaps = 14/138 (10%)
Query 132 PAVTEVFLRARRIAEA--EGSLTGMQHLFLALLYCPTFYKILVNSGCDIELFKEELALAY 189
P V E + AE+ EG + L L+LL + +++ NS E ++++
Sbjct 78 PRVGETLAKVLMAAESKGEGEYVSVNALVLSLLEVESIRELIANS----EDIRKKVEEQT 133
Query 190 KNTPMP--NTTGTTEQLTKDFDITQYGEDLTEKAEKGKLQPVIGRDEEIDRIAQILSRMM 247
KN N T+ +++ + D+ E+A + PVIGRDEEI R+ +ILS+
Sbjct 134 KNKRFDSRNADDGTDVMSR------FAVDMVEQARQNVFDPVIGRDEEIRRVLEILSKKT 187
Query 248 TKAPLLIGEPGVGKTAVI 265
+L+G+PGVGKTA++
Sbjct 188 KSNAILVGKPGVGKTAIV 205
> At5g51070
Length=945
Score = 61.6 bits (148), Expect = 2e-09, Method: Compositional matrix adjust.
Identities = 28/57 (49%), Positives = 40/57 (70%), Gaps = 0/57 (0%)
Query 210 ITQYGEDLTEKAEKGKLQPVIGRDEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIE 266
+ Q+ DLT +A +G + PVIGR++E+ R+ QIL R P+L+GE GVGKTA+ E
Sbjct 271 LEQFCVDLTARASEGLIDPVIGREKEVQRVIQILCRRTKNNPILLGEAGVGKTAIAE 327
> At3g45450
Length=341
Score = 47.4 bits (111), Expect = 4e-05, Method: Compositional matrix adjust.
Identities = 23/45 (51%), Positives = 31/45 (68%), Gaps = 1/45 (2%)
Query 223 KGKLQPVIGRDEEIDRIAQILSRMMTK-APLLIGEPGVGKTAVIE 266
+GKL PV+GR +I R+ QIL+R + LIG+PGVGK A+ E
Sbjct 150 RGKLDPVVGRQPQIKRVVQILARRTCRNNACLIGKPGVGKRAIAE 194
> Hs14737772
Length=634
Score = 33.9 bits (76), Expect = 0.35, Method: Compositional matrix adjust.
Identities = 35/114 (30%), Positives = 55/114 (48%), Gaps = 21/114 (18%)
Query 167 FYKILVNSGCDIELFKEELALAYKNTPMPNTTGTTEQLTKDF-DITQYGEDLT-----EK 220
FY L+N D + +K LAL N + + KD+ + +YG+ L ++
Sbjct 78 FYADLMNILYDKDHYK--LALGQINI----AKNLVDNVAKDYVRLMKYGDSLYRCKQLKR 131
Query 221 AEKGKLQPVIGRD----EEIDRIAQILSRMMTKAP-----LLIGEPGVGKTAVI 265
A G++ VI R E ++++ Q LSR+ T P LL G P VGK++ I
Sbjct 132 AALGRMCTVIKRQKQSLEYLEQVRQHLSRLPTIDPNTRTLLLCGYPNVGKSSFI 185
> Hs6912532
Length=633
Score = 33.9 bits (76), Expect = 0.36, Method: Compositional matrix adjust.
Identities = 35/114 (30%), Positives = 55/114 (48%), Gaps = 21/114 (18%)
Query 167 FYKILVNSGCDIELFKEELALAYKNTPMPNTTGTTEQLTKDF-DITQYGEDLT-----EK 220
FY L+N D + +K LAL N + + KD+ + +YG+ L ++
Sbjct 78 FYADLMNILYDKDHYK--LALGQINI----AKNLVDNVAKDYVRLMKYGDSLYRCKQLKR 131
Query 221 AEKGKLQPVIGRD----EEIDRIAQILSRMMTKAP-----LLIGEPGVGKTAVI 265
A G++ VI R E ++++ Q LSR+ T P LL G P VGK++ I
Sbjct 132 AALGRMCTVIKRQKQSLEYLEQVRQHLSRLPTIDPNTRTLLLCGYPNVGKSSFI 185
> YFL047w
Length=714
Score = 32.7 bits (73), Expect = 0.78, Method: Compositional matrix adjust.
Identities = 45/188 (23%), Positives = 78/188 (41%), Gaps = 31/188 (16%)
Query 67 HLHHDRLALAHLIMAMLEGSRLETPREVITKCHGDFSQVRGV----------IKAVLVRL 116
H + D LA A + + + RE K H F+++ G+ ++ +L +L
Sbjct 161 HFNEDELANAMKSLKIQNKYEEDVARE---KDHRFFNRIAGIDFDYKTMKETLQLLLTKL 217
Query 117 PRTPVTVPL---------GGAKVTPAVTEVFLRARRIAEAEGSLTGMQHLFLALL-YCPT 166
P+T +PL G ++T + + + + I +AE G L L L YC
Sbjct 218 PKTDYKLPLISYSLSNTNNGGEITKFLLD-HMSLKDIDQAET--FGQDLLNLGFLKYCNG 274
Query 167 FYKILVNSGCDIELFKEELALAYKNTPMPNTTGTTEQLTKDFDITQYGEDLTEKAEKGKL 226
VNS + + A + N PMP G+ E T + I+++ + + K +
Sbjct 275 VGNTFVNSK-KFQYQWKNTAYMFANVPMP---GSEEPTTGESLISRFN-NWDGSSAKEII 329
Query 227 QPVIGRDE 234
Q IG D+
Sbjct 330 QSKIGNDQ 337
> At3g61250
Length=299
Score = 32.3 bits (72), Expect = 1.0, Method: Compositional matrix adjust.
Identities = 22/100 (22%), Positives = 42/100 (42%), Gaps = 16/100 (16%)
Query 160 ALLYCPTFYKILVNSGCD--IELFKEELALAYKN------------TPMPNTTGTTEQLT 205
++L+ P+FY +V + CD + ++ E+ A++N +P T T+ L
Sbjct 165 SMLFSPSFYSGVVKTECDHFLRIWNSEIGEAFRNLAPLDESTITSQSPCSRATSTSSALL 224
Query 206 KDFDITQYGEDLTEKAEKGKLQPVIG--RDEEIDRIAQIL 243
K + G+++T P D+ D Q+L
Sbjct 225 KSSTNSWGGKEVTVAIHGSDYSPYSNDLEDDSTDSALQLL 264
> 7294632_1
Length=537
Score = 32.0 bits (71), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 14/36 (38%), Positives = 22/36 (61%), Gaps = 0/36 (0%)
Query 228 PVIGRDEEIDRIAQILSRMMTKAPLLIGEPGVGKTA 263
P +GR + +++ IL T+ L+ G+PG GKTA
Sbjct 459 PYVGRQWLVQQLSNILLGTETRVVLINGQPGTGKTA 494
> Hs18087809
Length=267
Score = 32.0 bits (71), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 25/69 (36%), Positives = 34/69 (49%), Gaps = 5/69 (7%)
Query 33 VATLPLRPTPEQRAPSELNFDAQLMVETMDTLAWHLHHDRLALAHLIMAMLEGSRLETPR 92
VA LRP Q EL+ M +T+ A L+ + +AL L + G L+T R
Sbjct 28 VALANLRPAENQVGSDELD---SYMYQTVGHHAIDLYAEAMALP-LYRRTIRGRSLDT-R 82
Query 93 EVITKCHGD 101
+V TKC GD
Sbjct 83 QVYTKCEGD 91
> Hs18390319
Length=2828
Score = 31.6 bits (70), Expect = 1.6, Method: Compositional matrix adjust.
Identities = 24/85 (28%), Positives = 39/85 (45%), Gaps = 10/85 (11%)
Query 181 FKEELA-LAYKNTPM--PNTTGTTEQLTKDFDITQYGEDLTEKAEKGKLQPVIGRDEEID 237
FKEE + + + TP P+ T +L D +T GE+LT+ P++ E++D
Sbjct 1365 FKEESSPVGFPGTPTWNPSRTAQPGRLQTDIPVTTSGENLTDP-------PLLKELEDVD 1417
Query 238 RIAQILSRMMTKAPLLIGEPGVGKT 262
++ LS + P E G T
Sbjct 1418 FTSEFLSSLTVSTPFHQEEAGSSTT 1442
> YOL094c
Length=323
Score = 30.4 bits (67), Expect = 3.6, Method: Compositional matrix adjust.
Identities = 16/48 (33%), Positives = 23/48 (47%), Gaps = 0/48 (0%)
Query 219 EKAEKGKLQPVIGRDEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVIE 266
EK L ++G E IDR+ QI ++ G PG+GKT +
Sbjct 13 EKYRPQVLSDIVGNKETIDRLQQIAKDGNMPHMIISGMPGIGKTTSVH 60
> ECU02g0290
Length=305
Score = 30.4 bits (67), Expect = 3.8, Method: Compositional matrix adjust.
Identities = 19/51 (37%), Positives = 23/51 (45%), Gaps = 0/51 (0%)
Query 215 EDLTEKAEKGKLQPVIGRDEEIDRIAQILSRMMTKAPLLIGEPGVGKTAVI 265
+ LTEK G L V+G E + + I S L G PG GKT I
Sbjct 3 QQLTEKYRPGSLLEVVGNREVVSSLQSISSAGRIPNMLFYGPPGTGKTTSI 53
> At5g46540
Length=1248
Score = 29.6 bits (65), Expect = 6.0, Method: Compositional matrix adjust.
Identities = 22/89 (24%), Positives = 37/89 (41%), Gaps = 14/89 (15%)
Query 191 NTPMPNTTGTTEQLTKDFDITQYGEDLTEKAEKGKLQPVIGRDEEIDRIA---------Q 241
+T P+ + FDI + +EKG + P++ D E+ ++ Q
Sbjct 962 STMAPDINKAKDSAASIFDILDSKPKIDSSSEKGTILPIVHGDIELQHVSFRYPMRPDIQ 1021
Query 242 ILSRMM-----TKAPLLIGEPGVGKTAVI 265
I S + + L+GE G GK+ VI
Sbjct 1022 IFSDLCLTISSGQTVALVGESGSGKSTVI 1050
> At1g63160
Length=333
Score = 29.6 bits (65), Expect = 6.8, Method: Compositional matrix adjust.
Identities = 17/54 (31%), Positives = 29/54 (53%), Gaps = 2/54 (3%)
Query 213 YGEDLTEKAEKGKLQPVIGRDEEIDRIAQILSRMMTKAPLLI-GEPGVGKTAVI 265
Y E EK K+ ++G ++ + R+ Q+++R L++ G PG GKT I
Sbjct 13 YNEPWVEKYRPSKVVDIVGNEDAVSRL-QVIARDGNMPNLILSGPPGTGKTTSI 65
> At3g06580
Length=496
Score = 29.3 bits (64), Expect = 8.3, Method: Compositional matrix adjust.
Identities = 23/92 (25%), Positives = 43/92 (46%), Gaps = 11/92 (11%)
Query 149 GSLTGMQHLFLALLY---CPTFYKILVNSGCDIELFKEELALAYKNTPMPNTTGTTEQLT 205
G L H ++LY CP ++ +++ KE AL + T G L
Sbjct 396 GDLMNESHYSCSVLYECSCPELEEL-------VQVCKENGALGARLTG-AGWGGCAVALV 447
Query 206 KDFDITQYGEDLTEKAEKGKLQPVIGRDEEID 237
K+FD+TQ+ + EK K +++ + + E+++
Sbjct 448 KEFDVTQFIPAVKEKYYKKRVEKGVVKKEDME 479
> Hs13540527
Length=378
Score = 29.3 bits (64), Expect = 9.3, Method: Compositional matrix adjust.
Identities = 18/50 (36%), Positives = 24/50 (48%), Gaps = 3/50 (6%)
Query 144 IAEAEGSLTGMQHLFLALLYCPTFYKILVNSGCDIELFKEELALAYKNTP 193
++ A SL LFL +C F +L SGC + F L LA K+ P
Sbjct 82 VSSASLSLPSRHRLFLTYRHCRNFSILLEPSGCSKDTF---LLLAIKSQP 128
> ECU01g1230
Length=780
Score = 29.3 bits (64), Expect = 9.6, Method: Compositional matrix adjust.
Identities = 21/72 (29%), Positives = 40/72 (55%), Gaps = 6/72 (8%)
Query 196 NTTGTTEQLTKDFDITQYGEDLTEKAEKGKLQPVIGRDEEIDRIAQILSRMMTKAP---L 252
+ T + E++ K+F++ Y + +A+ K++ ++ E R +Q+ S++ K P L
Sbjct 190 DETISREEVEKEFNMVGYDDVGGCRAQMAKIRELV---ELPLRHSQLYSKIGVKPPKGIL 246
Query 253 LIGEPGVGKTAV 264
L G PG GKT +
Sbjct 247 LYGPPGTGKTLI 258
Lambda K H
0.320 0.135 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 5631469314
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40