bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Emax_0825_orf2
Length=144
Score E
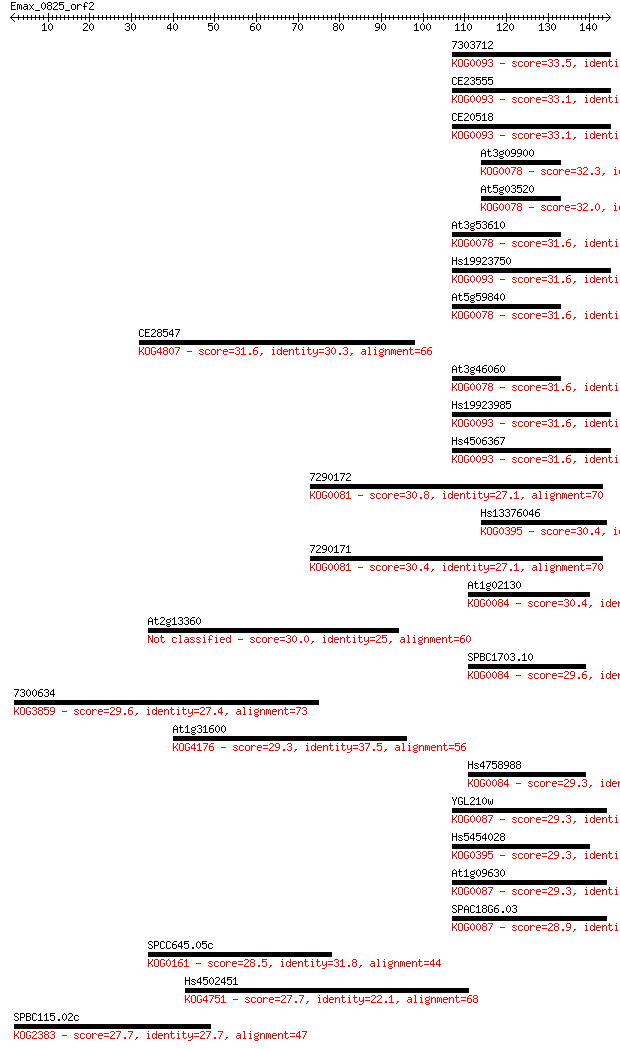
Sequences producing significant alignments: (Bits) Value
7303712 33.5 0.14
CE23555 33.1 0.16
CE20518 33.1 0.17
At3g09900 32.3 0.35
At5g03520 32.0 0.39
At3g53610 31.6 0.46
Hs19923750 31.6 0.47
At5g59840 31.6 0.47
CE28547 31.6 0.49
At3g46060 31.6 0.50
Hs19923985 31.6 0.55
Hs4506367 31.6 0.58
7290172 30.8 0.97
Hs13376046 30.4 1.1
7290171 30.4 1.1
At1g02130 30.4 1.1
At2g13360 30.0 1.5
SPBC1703.10 29.6 1.9
7300634 29.6 2.0
At1g31600 29.3 2.5
Hs4758988 29.3 2.6
YGL210w 29.3 2.8
Hs5454028 29.3 2.9
At1g09630 29.3 2.9
SPAC18G6.03 28.9 3.6
SPCC645.05c 28.5 5.0
Hs4502451 27.7 7.5
SPBC115.02c 27.7 8.6
> 7303712
Length=220
Score = 33.5 bits (75), Expect = 0.14, Method: Compositional matrix adjust.
Identities = 14/42 (33%), Positives = 27/42 (64%), Gaps = 4/42 (9%)
Query 107 DQYRSIA----HGRMGFLIVFDVTDPASFNTATELLTLTREF 144
++YR+I G MGF++++DVT+ SFN+ + +T + +
Sbjct 81 ERYRTITTAYYRGAMGFILMYDVTNEDSFNSVQDWVTQIKTY 122
> CE23555
Length=233
Score = 33.1 bits (74), Expect = 0.16, Method: Compositional matrix adjust.
Identities = 13/42 (30%), Positives = 26/42 (61%), Gaps = 4/42 (9%)
Query 107 DQYRSIA----HGRMGFLIVFDVTDPASFNTATELLTLTREF 144
++YR+I G MGF++++D+T+ SFN+ + T + +
Sbjct 96 ERYRTITTAYYRGAMGFILMYDITNEESFNSVQDWCTQIKTY 137
> CE20518
Length=219
Score = 33.1 bits (74), Expect = 0.17, Method: Compositional matrix adjust.
Identities = 13/42 (30%), Positives = 26/42 (61%), Gaps = 4/42 (9%)
Query 107 DQYRSIA----HGRMGFLIVFDVTDPASFNTATELLTLTREF 144
++YR+I G MGF++++D+T+ SFN+ + T + +
Sbjct 82 ERYRTITTAYYRGAMGFILMYDITNEESFNSVQDWCTQIKTY 123
> At3g09900
Length=218
Score = 32.3 bits (72), Expect = 0.35, Method: Compositional matrix adjust.
Identities = 12/19 (63%), Positives = 15/19 (78%), Gaps = 0/19 (0%)
Query 114 HGRMGFLIVFDVTDPASFN 132
G MG L+V+DVTD +SFN
Sbjct 86 RGAMGILLVYDVTDESSFN 104
> At5g03520
Length=216
Score = 32.0 bits (71), Expect = 0.39, Method: Compositional matrix adjust.
Identities = 12/19 (63%), Positives = 15/19 (78%), Gaps = 0/19 (0%)
Query 114 HGRMGFLIVFDVTDPASFN 132
G MG L+V+DVTD +SFN
Sbjct 86 RGAMGILLVYDVTDESSFN 104
> At3g53610
Length=216
Score = 31.6 bits (70), Expect = 0.46, Method: Compositional matrix adjust.
Identities = 14/30 (46%), Positives = 21/30 (70%), Gaps = 4/30 (13%)
Query 107 DQYRSIA----HGRMGFLIVFDVTDPASFN 132
+++R+I G MG L+V+DVTD +SFN
Sbjct 75 ERFRTITTAYYRGAMGILLVYDVTDESSFN 104
> Hs19923750
Length=219
Score = 31.6 bits (70), Expect = 0.47, Method: Compositional matrix adjust.
Identities = 13/42 (30%), Positives = 25/42 (59%), Gaps = 4/42 (9%)
Query 107 DQYRSIA----HGRMGFLIVFDVTDPASFNTATELLTLTREF 144
++YR+I G MGF++++D+T+ SFN + T + +
Sbjct 82 ERYRTITTAYYRGAMGFILMYDITNEESFNAVQDWATQIKTY 123
> At5g59840
Length=216
Score = 31.6 bits (70), Expect = 0.47, Method: Compositional matrix adjust.
Identities = 14/30 (46%), Positives = 21/30 (70%), Gaps = 4/30 (13%)
Query 107 DQYRSIA----HGRMGFLIVFDVTDPASFN 132
+++R+I G MG L+V+DVTD +SFN
Sbjct 75 ERFRTITTAYYRGAMGILLVYDVTDESSFN 104
> CE28547
Length=984
Score = 31.6 bits (70), Expect = 0.49, Method: Compositional matrix adjust.
Identities = 20/66 (30%), Positives = 34/66 (51%), Gaps = 2/66 (3%)
Query 32 LYRVMALPIDEGDEQTQKVICEVEDTFSPSKQNEKKSIRSLVDMKRRKLLNLPRGVKDFT 91
LY L + DEQ +I E E + S +N+K +RS +D K +++ +L R + ++
Sbjct 851 LYSAKCLENSQLDEQLSNLISEKEASVELSIENDK--LRSEIDRKSKEIGDLQRKMNNYE 908
Query 92 PFSLWR 97
S R
Sbjct 909 KLSQSR 914
> At3g46060
Length=216
Score = 31.6 bits (70), Expect = 0.50, Method: Compositional matrix adjust.
Identities = 14/30 (46%), Positives = 21/30 (70%), Gaps = 4/30 (13%)
Query 107 DQYRSIA----HGRMGFLIVFDVTDPASFN 132
+++R+I G MG L+V+DVTD +SFN
Sbjct 75 ERFRTITTAYYRGAMGILLVYDVTDESSFN 104
> Hs19923985
Length=227
Score = 31.6 bits (70), Expect = 0.55, Method: Compositional matrix adjust.
Identities = 13/42 (30%), Positives = 25/42 (59%), Gaps = 4/42 (9%)
Query 107 DQYRSIA----HGRMGFLIVFDVTDPASFNTATELLTLTREF 144
++YR+I G MGF++++D+T+ SFN + T + +
Sbjct 90 ERYRTITTAYYRGAMGFILMYDITNEESFNAVQDWSTQIKTY 131
> Hs4506367
Length=220
Score = 31.6 bits (70), Expect = 0.58, Method: Compositional matrix adjust.
Identities = 13/42 (30%), Positives = 25/42 (59%), Gaps = 4/42 (9%)
Query 107 DQYRSIA----HGRMGFLIVFDVTDPASFNTATELLTLTREF 144
++YR+I G MGF++++D+T+ SFN + T + +
Sbjct 82 ERYRTITTAYYRGAMGFILMYDITNEESFNAVQDWSTQIKTY 123
> 7290172
Length=236
Score = 30.8 bits (68), Expect = 0.97, Method: Compositional matrix adjust.
Identities = 19/74 (25%), Positives = 33/74 (44%), Gaps = 9/74 (12%)
Query 73 VDMKRRKLLNLPRGVKDFTPFSLWRPPIIPHHEGDQYRSIAHG----RMGFLIVFDVTDP 128
+D + ++LL RG + +W +++RS+ MGFL++FD+T
Sbjct 52 IDFREKRLLYNSRGRRHRIHLQIWDTA-----GQERFRSLTTAFYRDAMGFLLIFDLTSE 106
Query 129 ASFNTATELLTLTR 142
SF L+ R
Sbjct 107 KSFLETANWLSQLR 120
> Hs13376046
Length=205
Score = 30.4 bits (67), Expect = 1.1, Method: Compositional matrix adjust.
Identities = 13/30 (43%), Positives = 19/30 (63%), Gaps = 0/30 (0%)
Query 114 HGRMGFLIVFDVTDPASFNTATELLTLTRE 143
H GF+IV+D++D +SF A L+ RE
Sbjct 75 HWADGFVIVYDISDRSSFAFAKALIYRIRE 104
> 7290171
Length=249
Score = 30.4 bits (67), Expect = 1.1, Method: Compositional matrix adjust.
Identities = 19/74 (25%), Positives = 33/74 (44%), Gaps = 9/74 (12%)
Query 73 VDMKRRKLLNLPRGVKDFTPFSLWRPPIIPHHEGDQYRSIAHG----RMGFLIVFDVTDP 128
+D + ++LL RG + +W +++RS+ MGFL++FD+T
Sbjct 65 IDFREKRLLYNSRGRRHRIHLQIWDTA-----GQERFRSLTTAFYRDAMGFLLIFDLTSE 119
Query 129 ASFNTATELLTLTR 142
SF L+ R
Sbjct 120 KSFLETANWLSQLR 133
> At1g02130
Length=203
Score = 30.4 bits (67), Expect = 1.1, Method: Compositional matrix adjust.
Identities = 13/29 (44%), Positives = 17/29 (58%), Gaps = 0/29 (0%)
Query 111 SIAHGRMGFLIVFDVTDPASFNTATELLT 139
S G G +IV+DVTD SFN + L+
Sbjct 76 SYYRGAHGIIIVYDVTDEESFNNVKQWLS 104
> At2g13360
Length=401
Score = 30.0 bits (66), Expect = 1.5, Method: Compositional matrix adjust.
Identities = 15/60 (25%), Positives = 28/60 (46%), Gaps = 0/60 (0%)
Query 34 RVMALPIDEGDEQTQKVICEVEDTFSPSKQNEKKSIRSLVDMKRRKLLNLPRGVKDFTPF 93
+V+A + + + T K IC V + + N+ ++R+L+D + L L GV
Sbjct 124 QVLASKLSQDENHTIKAICIVHNETATGVTNDISAVRTLLDHYKHPALLLVDGVSSICAL 183
> SPBC1703.10
Length=203
Score = 29.6 bits (65), Expect = 1.9, Method: Compositional matrix adjust.
Identities = 13/28 (46%), Positives = 16/28 (57%), Gaps = 0/28 (0%)
Query 111 SIAHGRMGFLIVFDVTDPASFNTATELL 138
S G G +IV+DVTD SFN + L
Sbjct 76 SYYRGAHGIIIVYDVTDQDSFNNVKQWL 103
> 7300634
Length=419
Score = 29.6 bits (65), Expect = 2.0, Method: Compositional matrix adjust.
Identities = 20/74 (27%), Positives = 34/74 (45%), Gaps = 6/74 (8%)
Query 2 GCGKTSIADAFVNNVIARSNPPEPTTLPRLLYRVMALPIDEGDEQTQKVICEVEDTFSPS 61
G GK+++ D N + P P TLP + + + E + + + IC DT
Sbjct 53 GLGKSTLMDTLFNTSF--ESTPSPHTLPSVKLKAHTYELQESNVRLKLTIC---DTVGYG 107
Query 62 KQ-NEKKSIRSLVD 74
Q N+ S +++VD
Sbjct 108 DQINKDDSFKAVVD 121
> At1g31600
Length=410
Score = 29.3 bits (64), Expect = 2.5, Method: Compositional matrix adjust.
Identities = 21/62 (33%), Positives = 25/62 (40%), Gaps = 6/62 (9%)
Query 40 IDEGDEQTQKVICEVEDTFSPSKQNEKKSIRSLVDMKRRKL------LNLPRGVKDFTPF 93
+ D+ +VI D FS E S R D+K R L L LP V D P
Sbjct 130 VYAADDSGVRVIVSFADPFSAKAALEALSGRPCPDLKGRSLHIRYSVLQLPSEVNDCVPV 189
Query 94 SL 95
SL
Sbjct 190 SL 191
> Hs4758988
Length=205
Score = 29.3 bits (64), Expect = 2.6, Method: Compositional matrix adjust.
Identities = 12/28 (42%), Positives = 16/28 (57%), Gaps = 0/28 (0%)
Query 111 SIAHGRMGFLIVFDVTDPASFNTATELL 138
S G G ++V+DVTD SFN + L
Sbjct 79 SYYRGAHGIIVVYDVTDQESFNNVKQWL 106
> YGL210w
Length=222
Score = 29.3 bits (64), Expect = 2.8, Method: Compositional matrix adjust.
Identities = 14/41 (34%), Positives = 23/41 (56%), Gaps = 4/41 (9%)
Query 107 DQYRSIA----HGRMGFLIVFDVTDPASFNTATELLTLTRE 143
++YR+I G +G LIV+D++ +S+ LT RE
Sbjct 73 ERYRAITSAYYRGAVGALIVYDISKSSSYENCNHWLTELRE 113
> Hs5454028
Length=218
Score = 29.3 bits (64), Expect = 2.9, Method: Compositional matrix adjust.
Identities = 14/33 (42%), Positives = 18/33 (54%), Gaps = 3/33 (9%)
Query 107 DQYRSIAHGRMGFLIVFDVTDPASFNTATELLT 139
+QY HG FL+VF + D SFN +L T
Sbjct 95 EQYMRAGHG---FLLVFAINDRQSFNEVGKLFT 124
> At1g09630
Length=217
Score = 29.3 bits (64), Expect = 2.9, Method: Compositional matrix adjust.
Identities = 14/41 (34%), Positives = 23/41 (56%), Gaps = 4/41 (9%)
Query 107 DQYRSIA----HGRMGFLIVFDVTDPASFNTATELLTLTRE 143
++YR+I G +G L+V+DVT P +F + L R+
Sbjct 72 ERYRAITSAYYRGALGALLVYDVTKPTTFENVSRWLKELRD 112
> SPAC18G6.03
Length=214
Score = 28.9 bits (63), Expect = 3.6, Method: Compositional matrix adjust.
Identities = 15/41 (36%), Positives = 23/41 (56%), Gaps = 4/41 (9%)
Query 107 DQYRSIA----HGRMGFLIVFDVTDPASFNTATELLTLTRE 143
++YR+I G +G LIV+D+T +SF+ L RE
Sbjct 70 ERYRAITSAYYRGAVGALIVYDITKQSSFDNVGRWLKELRE 110
> SPCC645.05c
Length=1526
Score = 28.5 bits (62), Expect = 5.0, Method: Composition-based stats.
Identities = 14/44 (31%), Positives = 25/44 (56%), Gaps = 0/44 (0%)
Query 34 RVMALPIDEGDEQTQKVICEVEDTFSPSKQNEKKSIRSLVDMKR 77
R +AL +E +TQ+ + +ED+FS +KQ + R +K+
Sbjct 872 RAIALDKEEILRRTQERLANIEDSFSETKQQNENLQRESASLKQ 915
> Hs4502451
Length=3418
Score = 27.7 bits (60), Expect = 7.5, Method: Compositional matrix adjust.
Identities = 15/71 (21%), Positives = 34/71 (47%), Gaps = 3/71 (4%)
Query 43 GDEQTQKVICEVEDTFSPSKQNEKKSIRSLVDMKRRKLLNLPRGVKDFTPFSLWRPPIIP 102
D++ ++ E+ ++Q E+ R + + + ++++ + KD S+WRP
Sbjct 2937 NDKKQAQIQLEIRKAMESAEQKEQGLSRDVTTVWKLRIVSYSKKEKDSVILSIWRPSSDL 2996
Query 103 HH---EGDQYR 110
+ EG +YR
Sbjct 2997 YSLLTEGKRYR 3007
> SPBC115.02c
Length=454
Score = 27.7 bits (60), Expect = 8.6, Method: Compositional matrix adjust.
Identities = 13/47 (27%), Positives = 21/47 (44%), Gaps = 5/47 (10%)
Query 2 GCGKTSIADAFVNNVIARSNPPEPTTLPRLLYRVMALPIDEGDEQTQ 48
GCGKT++ D F +N+ PP T R+ + + + Q
Sbjct 128 GCGKTALMDLFYHNL-----PPNVTRSQRIHFHAFMMQVHRTSHDLQ 169
Lambda K H
0.319 0.137 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1675978996
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40