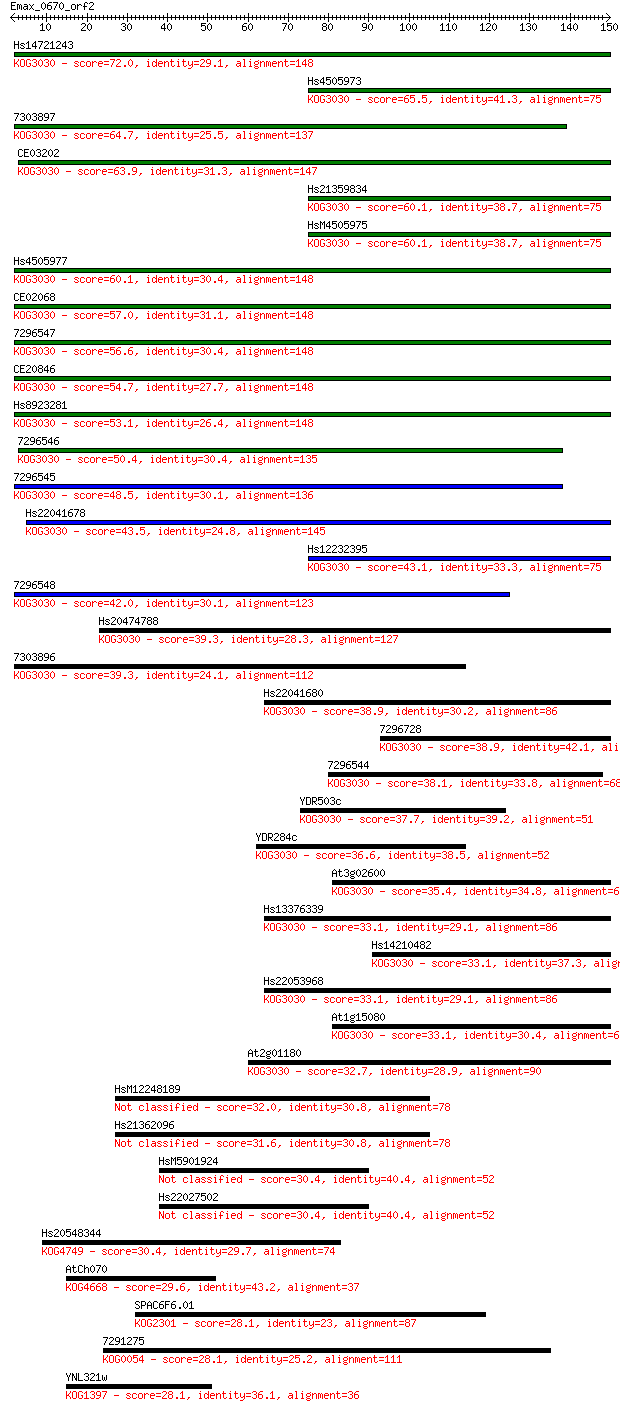
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Emax_0670_orf2
Length=149
Score E
Sequences producing significant alignments: (Bits) Value
Hs14721243 72.0 4e-13
Hs4505973 65.5 4e-11
7303897 64.7 7e-11
CE03202 63.9 1e-10
Hs21359834 60.1 2e-09
HsM4505975 60.1 2e-09
Hs4505977 60.1 2e-09
CE02068 57.0 1e-08
7296547 56.6 2e-08
CE20846 54.7 6e-08
Hs8923281 53.1 2e-07
7296546 50.4 1e-06
7296545 48.5 5e-06
Hs22041678 43.5 1e-04
Hs12232395 43.1 2e-04
7296548 42.0 4e-04
Hs20474788 39.3 0.003
7303896 39.3 0.003
Hs22041680 38.9 0.003
7296728 38.9 0.004
7296544 38.1 0.007
YDR503c 37.7 0.008
YDR284c 36.6 0.021
At3g02600 35.4 0.043
Hs13376339 33.1 0.19
Hs14210482 33.1 0.20
Hs22053968 33.1 0.22
At1g15080 33.1 0.23
At2g01180 32.7 0.29
HsM12248189 32.0 0.46
Hs21362096 31.6 0.57
HsM5901924 30.4 1.3
Hs22027502 30.4 1.3
Hs20548344 30.4 1.4
AtCh070 29.6 2.2
SPAC6F6.01 28.1 5.7
7291275 28.1 5.9
YNL321w 28.1 7.3
> Hs14721243
Length=285
Score = 72.0 bits (175), Expect = 4e-13, Method: Compositional matrix adjust.
Identities = 43/148 (29%), Positives = 67/148 (45%), Gaps = 20/148 (13%)
Query 2 FCGEDDIALPYMPASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVV 61
FC ++ I PY +++++ + LP I++ E L +V S +
Sbjct 40 FCKDNSINYPYHDSTVTSTVLILVGVGLPISSIILGETL--SVYCNLLHSNSFIRNNYIA 97
Query 62 VVLDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCR 121
+ Y G+ A+ LT+ K ++G LRPHFLDVC PDWS+++C
Sbjct 98 TI------------YKAIGTFLFGAAASQSLTDIAKYSIGRLRPHFLDVCDPDWSKINCS 145
Query 122 DSSGLYDVYIPDFFCTGDPHRVEEARRS 149
D YI + C G+ RV+E R S
Sbjct 146 DG------YIEYYICRGNAERVKEGRLS 167
> Hs4505973
Length=289
Score = 65.5 bits (158), Expect = 4e-11, Method: Compositional matrix adjust.
Identities = 31/75 (41%), Positives = 43/75 (57%), Gaps = 6/75 (8%)
Query 75 LYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCRDSSGLYDVYIPDF 134
+Y G+ A+ LT+ K ++G LRPHFLDVC PDWS+++C D YI +
Sbjct 103 IYKAIGTFLFGAAASQSLTDIAKYSIGRLRPHFLDVCDPDWSKINCSDG------YIEYY 156
Query 135 FCTGDPHRVEEARRS 149
C G+ RV+E R S
Sbjct 157 ICRGNAERVKEGRLS 171
> 7303897
Length=372
Score = 64.7 bits (156), Expect = 7e-11, Method: Compositional matrix adjust.
Identities = 35/137 (25%), Positives = 67/137 (48%), Gaps = 3/137 (2%)
Query 2 FCGEDDIALPYMPASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVV 61
FC ++ + P+ +++ ++I ++P +I +VEV++ +A Q+ + +
Sbjct 112 FCDDESLKHPFHDSTVRNWMLYFIGAVIPVGVIFIVEVIISQNKAKQDNGNATSRR---Y 168
Query 62 VVLDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCR 121
V ++ +P+ + Y G A L T+ K ++G LRPHF+ VC+P + S
Sbjct 169 VFMNYELPDWMIECYKKIGIYAFGAVLSQLTTDIAKYSIGRLRPHFIAVCQPQMADGSTC 228
Query 122 DSSGLYDVYIPDFFCTG 138
D + YI +F C G
Sbjct 229 DDAINAGKYIQEFTCKG 245
> CE03202
Length=318
Score = 63.9 bits (154), Expect = 1e-10, Method: Compositional matrix adjust.
Identities = 46/148 (31%), Positives = 62/148 (41%), Gaps = 20/148 (13%)
Query 3 CGEDDIALPYMPASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVVV 62
CG+ I P+ ++ I+ P I+ +VE +L K K
Sbjct 84 CGDISIQQPFKENTVGLKHLLVITLGSPFLIVALVEAIL------HFKSKGSNR------ 131
Query 63 VLDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCRD 122
L + T + TY L M A F + LK VG LRPHF VC+PDWS+V C D
Sbjct 132 -LAKFFSATTI---TYLKYLLMYAACTFAM-EFLKCYVGRLRPHFFSVCKPDWSKVDCTD 186
Query 123 SSGLYDVYIPDFFCTG-DPHRVEEARRS 149
D D CT +P ++ AR S
Sbjct 187 KQSFIDS--SDLVCTNPNPRKIRTARTS 212
> Hs21359834
Length=311
Score = 60.1 bits (144), Expect = 2e-09, Method: Compositional matrix adjust.
Identities = 29/75 (38%), Positives = 42/75 (56%), Gaps = 6/75 (8%)
Query 75 LYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCRDSSGLYDVYIPDF 134
LY G A T+ K ++G LRPHFL VC PD+S+++C + YI ++
Sbjct 126 LYKQVGCFLFGCAISQSFTDIAKVSIGRLRPHFLSVCNPDFSQINCSEG------YIQNY 179
Query 135 FCTGDPHRVEEARRS 149
C GD +V+EAR+S
Sbjct 180 RCRGDDSKVQEARKS 194
> HsM4505975
Length=311
Score = 60.1 bits (144), Expect = 2e-09, Method: Compositional matrix adjust.
Identities = 29/75 (38%), Positives = 42/75 (56%), Gaps = 6/75 (8%)
Query 75 LYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCRDSSGLYDVYIPDF 134
LY G A T+ K ++G LRPHFL VC PD+S+++C + YI ++
Sbjct 126 LYKQVGCFLFGCAISQSFTDIAKVSIGRLRPHFLSVCNPDFSQINCSEG------YIQNY 179
Query 135 FCTGDPHRVEEARRS 149
C GD +V+EAR+S
Sbjct 180 RCRGDDSKVQEARKS 194
> Hs4505977
Length=288
Score = 60.1 bits (144), Expect = 2e-09, Method: Compositional matrix adjust.
Identities = 45/149 (30%), Positives = 64/149 (42%), Gaps = 23/149 (15%)
Query 2 FCGEDDIALPYMPASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVV 61
+CG+D I PY P +I+ + A +I+V + V + +S+
Sbjct 37 YCGDDSIRYPYRPDTITHG--LMAGVTITATVILVSAGEAYLVYTDRLYSRSDFNNYVAA 94
Query 62 VVLDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCR 121
V Y G+ A LT+ K +G LRP+FL VC PDWSRV+C
Sbjct 95 V-------------YKVLGTFLFGAAVSQSLTDLAKYMIGRLRPNFLAVCDPDWSRVNC- 140
Query 122 DSSGLYDVYIP-DFFCTGDPHRVEEARRS 149
VY+ + C G+P V EAR S
Sbjct 141 ------SVYVQLEKVCRGNPADVTEARLS 163
> CE02068
Length=341
Score = 57.0 bits (136), Expect = 1e-08, Method: Compositional matrix adjust.
Identities = 46/148 (31%), Positives = 61/148 (41%), Gaps = 11/148 (7%)
Query 2 FCGEDDIALPYMPASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVV 61
FC +D I Y +I+A + +L A ++ VE + + R + +
Sbjct 56 FCDDDSIRYEYRKDTITAVQLMLYNLVLNAATVLFVEYYRMQKVESNINNPRYRWRNNHL 115
Query 62 VVLDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCR 121
VL V L TY G + L K VG LRPHFLDVC+
Sbjct 116 HVL-------FVRLLTYFGYSQIGFVMNIALNIVTKHVVGRLRPHFLDVCKLANDTCVTG 168
Query 122 DSSGLYDVYIPDFFCTGDPHRVEEARRS 149
DS YI D+ CTG P V EAR+S
Sbjct 169 DSHR----YITDYTCTGPPELVLEARKS 192
> 7296547
Length=340
Score = 56.6 bits (135), Expect = 2e-08, Method: Compositional matrix adjust.
Identities = 45/149 (30%), Positives = 69/149 (46%), Gaps = 11/149 (7%)
Query 2 FCGEDDIALPYMPASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVV 61
FC ++ I+ P+ +I+ I +LPA ++VVVE V + D S A V
Sbjct 67 FCDDESISYPFQDNTITPVMLGLIVGLLPALVMVVVEY----VSHLRAGDIS-----ATV 117
Query 62 VVLDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVS-C 120
+L R+ V L + F T K +G LRPHFL VC+P + S C
Sbjct 118 DLLGWRVSTWYVELGRQSTYFCFGLLLTFDATEVGKYTIGRLRPHFLAVCQPQIADGSMC 177
Query 121 RDSSGLYDVYIPDFFCTGDPHRVEEARRS 149
D L+ Y+ ++ C G+ VE+ R++
Sbjct 178 SDPVNLHR-YMENYDCAGEGFTVEDVRQA 205
> CE20846
Length=346
Score = 54.7 bits (130), Expect = 6e-08, Method: Compositional matrix adjust.
Identities = 41/149 (27%), Positives = 72/149 (48%), Gaps = 10/149 (6%)
Query 2 FCGEDDIALPYMPASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVV 61
+C ++ I P+ + ++ + ++P +I+ E L+ A ++K ++E + V
Sbjct 41 YCDDESIRYPFRDSKVTRQMLIVVGLLIPILLILATE--LFRTLAWEKKCETEFKTYHV- 97
Query 62 VVLDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCR 121
+ + V LY + G + V L+ + K +G RPHF+DVCRPD +C
Sbjct 98 --RNHSVHRLVVRLYCFIGYFFVGVCFNQLMVDIAKYTIGRQRPHFMDVCRPDIGYQTCS 155
Query 122 DSSGLYDVYIPDFFC-TGDPHRVEEARRS 149
D+YI DF C T D ++ EA+ S
Sbjct 156 QP----DLYITDFKCTTTDTKKIHEAQLS 180
> Hs8923281
Length=325
Score = 53.1 bits (126), Expect = 2e-07, Method: Compositional matrix adjust.
Identities = 39/158 (24%), Positives = 73/158 (46%), Gaps = 21/158 (13%)
Query 2 FCGEDDIALPYMPAS-----ISAAGAFWISTMLPAFIIVVVEVLLWAVRATQE----KDK 52
FC + D+ PY P + I+ + + P II + E+ ++ +++T+E ++K
Sbjct 49 FCQDGDLMKPY-PGTEEESFITPLVLYCVLAATPTAIIFIGEISMYFIKSTRESLIAREK 107
Query 53 SEREQQAVVVV-LDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVC 111
+ + + L RRI + G A + + + N+ + G L P+FL VC
Sbjct 108 TILTGECCYLNPLLRRIIR-------FTGVFAFGLFATDIFVNAGQVVTGHLTPYFLTVC 160
Query 112 RPDWSRVSCRDSSGLYDVYIPDFFCTGDPHRVEEARRS 149
+P+++ C+ + CTGD +E+ARRS
Sbjct 161 KPNYTSADCQAHHQFIN---NGNICTGDLEVIEKARRS 195
> 7296546
Length=305
Score = 50.4 bits (119), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 41/141 (29%), Positives = 64/141 (45%), Gaps = 14/141 (9%)
Query 3 CGEDDIALPYMPASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVVV 62
C + + PY ++ LPA ++VVE+L RA +E Q+ V V
Sbjct 44 CSDTSLKYPYRQPWLTKVHLTIAVVALPAAFVLVVEML----RAAVVPSSTELTQRFVFV 99
Query 63 V--LDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSR--- 117
+ R I E + Y L +++A++ L +S G LRP+F D+C+P W
Sbjct 100 GVRIPRFISECYKAIGVYLFGLGLTLAAIRLTKHS----TGRLRPYFFDICQPTWGTEGG 155
Query 118 VSCRDSSGLYD-VYIPDFFCT 137
SC D + +Y+ DF CT
Sbjct 156 ESCSDVTAQNSTLYLEDFSCT 176
> 7296545
Length=341
Score = 48.5 bits (114), Expect = 5e-06, Method: Compositional matrix adjust.
Identities = 41/139 (29%), Positives = 62/139 (44%), Gaps = 14/139 (10%)
Query 2 FCGEDDIALPYMPASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVV 61
FCG++ ++ P +IS+ I +P +IVVVE+ + +R+ +
Sbjct 39 FCGDETLSYPARDGTISSKVIIAIVLGVPNAVIVVVELFRQLPGGPLREAGGKRDSCRIA 98
Query 62 VVLDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCR---PDWSRV 118
L + +LY +A V T K +G LRPHFL VC+ PD S
Sbjct 99 HRLGVLYRQVIFYLY--------GLAMVTFTTMLTKLCLGRLRPHFLAVCQPMLPDGS-- 148
Query 119 SCRDSSGLYDVYIPDFFCT 137
SC+D+ L YI F C+
Sbjct 149 SCQDAQNL-GRYIDSFTCS 166
> Hs22041678
Length=321
Score = 43.5 bits (101), Expect = 1e-04, Method: Compositional matrix adjust.
Identities = 36/147 (24%), Positives = 65/147 (44%), Gaps = 16/147 (10%)
Query 5 EDDIALPYMPASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVVVVL 64
ED A+P + AAG +P +I+V E AV Q + Q+ ++
Sbjct 58 EDSSAVPPVLLYSLAAG-------VPVLVIIVGET---AVFCLQLATRDFENQEKTILTG 107
Query 65 DR-RIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCRDS 123
D I + G + + + N+ + G+L PHFL +C+P+++ + C+
Sbjct 108 DCCYINPLVRRTVRFLGIYTFGLFATDIFVNAGQVVTGNLAPHFLALCKPNYTALGCQQ- 166
Query 124 SGLYDVYIP-DFFCTGDPHRVEEARRS 149
Y +I + CTG+P + AR++
Sbjct 167 ---YTQFISGEEACTGNPDLIMRARKT 190
> Hs12232395
Length=343
Score = 43.1 bits (100), Expect = 2e-04, Method: Compositional matrix adjust.
Identities = 25/81 (30%), Positives = 38/81 (46%), Gaps = 8/81 (9%)
Query 75 LYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCRDSSGLYDVYIPDF 134
L + G + + + + N+ + G+ PHFL VCRP+++ + C S D PD
Sbjct 126 LVRFLGVYSFGLFTTTIFANAGQVVTGNPTPHFLSVCRPNYTALGCLPPSP--DRPGPDR 183
Query 135 F------CTGDPHRVEEARRS 149
F C G P V ARR+
Sbjct 184 FVTDQGACAGSPSLVAAARRA 204
> 7296548
Length=305
Score = 42.0 bits (97), Expect = 4e-04, Method: Compositional matrix adjust.
Identities = 37/124 (29%), Positives = 48/124 (38%), Gaps = 20/124 (16%)
Query 2 FCGEDDIALPYMPASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVV 61
FC ++ + PY ++S W+ LP +VV+E L S R+ A
Sbjct 43 FCDDESLMYPYHENTVSPTLLHWLGLYLPLISLVVLESFL-----------SHRKDMAPW 91
Query 62 VVLDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDW-SRVSC 120
L LY Y S LL K A+G LRPHF VC P + SC
Sbjct 92 PTLWPVYNTVRWFLYGY--------VSNDLLKGIGKQALGRLRPHFFAVCSPHFPDGSSC 143
Query 121 RDSS 124
D S
Sbjct 144 LDES 147
> Hs20474788
Length=271
Score = 39.3 bits (90), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 36/127 (28%), Positives = 53/127 (41%), Gaps = 40/127 (31%)
Query 23 FWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVVVVLDRRIPETCVHLYTYCGSL 82
F IS + P +I VV+++ + DK+E ++ + V SL
Sbjct 55 FAISFLTPLAVICVVKII-------RRTDKTEIKEAFLAV------------------SL 89
Query 83 AMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCRDSSGLYDVYIPDFFCTGDPHR 142
A+++ V TN++K VG RP F C PD V + CTGDP
Sbjct 90 ALALNGV--CTNTIKLIVGRPRPDFFYRCFPD-------------GVMNSEMHCTGDPDL 134
Query 143 VEEARRS 149
V E R+S
Sbjct 135 VSEGRKS 141
> 7303896
Length=246
Score = 39.3 bits (90), Expect = 0.003, Method: Compositional matrix adjust.
Identities = 27/115 (23%), Positives = 49/115 (42%), Gaps = 7/115 (6%)
Query 2 FCGEDDIALPYMPASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVV 61
FC + + PY +++ + + + LP +++VVE R ++ S + +
Sbjct 16 FCDDSSLRHPYRDSTMPSWILYLMCGALPLTVMLVVEFF----RGQDKRLHSPFPKSTMC 71
Query 62 V---VLDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRP 113
+ +P V Y G + L TN K ++G LRPHF +C+P
Sbjct 72 SGYHLCHLELPTWLVECYHRMGIFIFGLGVEQLSTNIAKYSIGRLRPHFYTLCQP 126
> Hs22041680
Length=763
Score = 38.9 bits (89), Expect = 0.003, Method: Composition-based stats.
Identities = 26/89 (29%), Positives = 46/89 (51%), Gaps = 16/89 (17%)
Query 64 LDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWS--RVSCR 121
L R + VH++ C S L+T+ ++ + G P+FL VC+P+++ VSC+
Sbjct 172 LRRAVRFVGVHVFGLC--------STALITDIIQLSTGYQAPYFLTVCKPNYTSLNVSCK 223
Query 122 DSSGLYDVYIPDFFCTG-DPHRVEEARRS 149
++S YI + C+G D + R+S
Sbjct 224 ENS-----YIVEDICSGSDLTVINSGRKS 247
> 7296728
Length=412
Score = 38.9 bits (89), Expect = 0.004, Method: Compositional matrix adjust.
Identities = 24/57 (42%), Positives = 30/57 (52%), Gaps = 1/57 (1%)
Query 93 TNSLKAAVGSLRPHFLDVCRPDWSRVSCRDSSGLYDVYIPDFFCTGDPHRVEEARRS 149
T+ LK VG RP + C PD V S+G+ D I DF CTG P + E R+S
Sbjct 212 TSVLKITVGRPRPDYFYRCFPDGVMVLNTTSNGV-DTSILDFNCTGLPGDINEGRKS 267
> 7296544
Length=334
Score = 38.1 bits (87), Expect = 0.007, Method: Compositional matrix adjust.
Identities = 23/71 (32%), Positives = 33/71 (46%), Gaps = 6/71 (8%)
Query 80 GSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRP---DWSRVSCRDSSGLYDVYIPDFFC 136
+ + + +L T K AVG LRPHF C+P D S SC D ++Y+ F C
Sbjct 67 ATFSFGFIATYLTTELAKHAVGRLRPHFFHGCQPRLDDGS--SCSDLQNA-ELYVEQFHC 123
Query 137 TGDPHRVEEAR 147
T + + R
Sbjct 124 TNNNLSTRQIR 134
> YDR503c
Length=274
Score = 37.7 bits (86), Expect = 0.008, Method: Compositional matrix adjust.
Identities = 20/51 (39%), Positives = 30/51 (58%), Gaps = 2/51 (3%)
Query 73 VHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCRDS 123
+H C L +S+ + LT +LK +G+LRP F+D C PD ++S DS
Sbjct 114 MHTSILCLMLIISINAA--LTGALKLIIGNLRPDFVDRCIPDLQKMSDSDS 162
> YDR284c
Length=289
Score = 36.6 bits (83), Expect = 0.021, Method: Compositional matrix adjust.
Identities = 20/52 (38%), Positives = 28/52 (53%), Gaps = 2/52 (3%)
Query 62 VVLDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRP 113
++ DRR LYT L+++ S TN +K +G LRP FLD C+P
Sbjct 85 ILADRR--HLIFILYTSLLGLSLAWFSTSFFTNFIKNWIGRLRPDFLDRCQP 134
> At3g02600
Length=314
Score = 35.4 bits (80), Expect = 0.043, Method: Compositional matrix adjust.
Identities = 24/69 (34%), Positives = 31/69 (44%), Gaps = 9/69 (13%)
Query 81 SLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCRDSSGLYDVYIPDFFCTGDP 140
L SV +LT+++K AVG RP F C P D LYD + D C GD
Sbjct 102 GLLYSVLVTAVLTDAIKNAVGRPRPDFFWRCFP--------DGKALYDS-LGDVICHGDK 152
Query 141 HRVEEARRS 149
+ E +S
Sbjct 153 SVIREGHKS 161
> Hs13376339
Length=746
Score = 33.1 bits (74), Expect = 0.19, Method: Compositional matrix adjust.
Identities = 25/89 (28%), Positives = 42/89 (47%), Gaps = 16/89 (17%)
Query 64 LDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWS--RVSCR 121
L R + VH++ C + L+T+ ++ A G P FL VC+P+++ SC
Sbjct 127 LRRTVRFVGVHVFGLC--------ATALVTDVIQLATGYHTPFFLTVCKPNYTLLGTSCE 178
Query 122 DSSGLYDVYIPDFFCTG-DPHRVEEARRS 149
+ YI C+G D H + AR++
Sbjct 179 -----VNPYITQDICSGHDIHAILSARKT 202
> Hs14210482
Length=175
Score = 33.1 bits (74), Expect = 0.20, Method: Compositional matrix adjust.
Identities = 22/59 (37%), Positives = 26/59 (44%), Gaps = 13/59 (22%)
Query 91 LLTNSLKAAVGSLRPHFLDVCRPDWSRVSCRDSSGLYDVYIPDFFCTGDPHRVEEARRS 149
+ TN++K VG RP F C PD GL D CTGD V E R+S
Sbjct 61 VFTNTIKLIVGRPRPDFFYRCFPD----------GLAHS---DLMCTGDKDVVNEGRKS 106
> Hs22053968
Length=686
Score = 33.1 bits (74), Expect = 0.22, Method: Compositional matrix adjust.
Identities = 25/89 (28%), Positives = 42/89 (47%), Gaps = 16/89 (17%)
Query 64 LDRRIPETCVHLYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWS--RVSCR 121
L R + VH++ C + L+T+ ++ A G P FL VC+P+++ SC
Sbjct 67 LRRTVRFVGVHVFGLC--------ATALVTDVIQLATGYHTPFFLTVCKPNYTLLGTSCE 118
Query 122 DSSGLYDVYIPDFFCTG-DPHRVEEARRS 149
+ YI C+G D H + AR++
Sbjct 119 -----VNPYITQDICSGHDIHAILSARKT 142
> At1g15080
Length=290
Score = 33.1 bits (74), Expect = 0.23, Method: Compositional matrix adjust.
Identities = 21/69 (30%), Positives = 31/69 (44%), Gaps = 8/69 (11%)
Query 81 SLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCRDSSGLYDVYIPDFFCTGDP 140
L SV ++T+++K AVG RP F C P D G++ + CTG
Sbjct 102 GLLFSVLITGVITDAIKDAVGRPRPDFFWRCFP--------DGIGIFHNVTKNVLCTGAK 153
Query 141 HRVEEARRS 149
V+E +S
Sbjct 154 DVVKEGHKS 162
> At2g01180
Length=302
Score = 32.7 bits (73), Expect = 0.29, Method: Compositional matrix adjust.
Identities = 26/91 (28%), Positives = 41/91 (45%), Gaps = 10/91 (10%)
Query 60 VVVVLDRRIPETCVH-LYTYCGSLAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRV 118
++V + + TCV+ L+ L +V ++T+S+K A G RP+F C P
Sbjct 80 IIVFVCFYLKRTCVYDLHHSILGLLFAVLITGVITDSIKVATGRPRPNFYWRCFP----- 134
Query 119 SCRDSSGLYDVYIPDFFCTGDPHRVEEARRS 149
D LYD + C G V+E +S
Sbjct 135 ---DGKELYDA-LGGVVCHGKAAEVKEGHKS 161
> HsM12248189
Length=403
Score = 32.0 bits (71), Expect = 0.46, Method: Compositional matrix adjust.
Identities = 24/80 (30%), Positives = 41/80 (51%), Gaps = 3/80 (3%)
Query 27 TMLPAFIIVVVEVLLWAV-RATQEKDKSEREQ-QAVVVVLDRRIPETCVHLYTYCGSLAM 84
T PA +V++ LW + R + +S RE + V V +D+ +P++C+ ++ L
Sbjct 78 TYGPAPDLVIINSCLWDLSRYGRCSMESYRENLERVFVRMDQVLPDSCLLVWNMAMPLGE 137
Query 85 SVASVFLLTNSLKAAVGSLR 104
+ FLL L+ GSLR
Sbjct 138 RITGGFLLP-ELQPLAGSLR 156
> Hs21362096
Length=454
Score = 31.6 bits (70), Expect = 0.57, Method: Compositional matrix adjust.
Identities = 24/80 (30%), Positives = 41/80 (51%), Gaps = 3/80 (3%)
Query 27 TMLPAFIIVVVEVLLWAV-RATQEKDKSEREQ-QAVVVVLDRRIPETCVHLYTYCGSLAM 84
T PA +V++ LW + R + +S RE + V V +D+ +P++C+ ++ L
Sbjct 129 TYGPAPDLVIINSCLWDLSRYGRCSMESYRENLERVFVRMDQVLPDSCLLVWNMAMPLGE 188
Query 85 SVASVFLLTNSLKAAVGSLR 104
+ FLL L+ GSLR
Sbjct 189 RITGGFLLP-ELQPLAGSLR 207
> HsM5901924
Length=994
Score = 30.4 bits (67), Expect = 1.3, Method: Composition-based stats.
Identities = 21/53 (39%), Positives = 29/53 (54%), Gaps = 1/53 (1%)
Query 38 EVLLWAVRATQEKDKSE-REQQAVVVVLDRRIPETCVHLYTYCGSLAMSVASV 89
EVLL A RA EK KS+ QQ + VLDR++ L+ GS+ + A +
Sbjct 241 EVLLQAKRAELEKLKSQVTSQQQEMAVLDRQLGHKKEELHLLQGSMVQAKADL 293
> Hs22027502
Length=994
Score = 30.4 bits (67), Expect = 1.3, Method: Composition-based stats.
Identities = 21/53 (39%), Positives = 29/53 (54%), Gaps = 1/53 (1%)
Query 38 EVLLWAVRATQEKDKSE-REQQAVVVVLDRRIPETCVHLYTYCGSLAMSVASV 89
EVLL A RA EK KS+ QQ + VLDR++ L+ GS+ + A +
Sbjct 241 EVLLQAKRAELEKLKSQVTSQQQEMAVLDRQLGHKKEELHLLQGSMVQAKADL 293
> Hs20548344
Length=190
Score = 30.4 bits (67), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 22/76 (28%), Positives = 37/76 (48%), Gaps = 7/76 (9%)
Query 9 ALPYMPASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEKDKSEREQQAVVVVLD--R 66
AL +MPAS + +++ LPA ++ WA T+EK SE Q + ++D +
Sbjct 7 ALWFMPAS-----SLFLALKLPARHLLHQRFAEWANECTEEKKTSEEIFQHLQNIVDFGK 61
Query 67 RIPETCVHLYTYCGSL 82
+ E Y +CG +
Sbjct 62 NVMEFLGENYVHCGEV 77
> AtCh070
Length=746
Score = 29.6 bits (65), Expect = 2.2, Method: Composition-based stats.
Identities = 16/48 (33%), Positives = 26/48 (54%), Gaps = 11/48 (22%)
Query 15 ASISAAGAFWISTMLPAFIIV-----------VVEVLLWAVRATQEKD 51
A++ AAG F ++ +LP FI++ ++ VLL A A +KD
Sbjct 267 ATMVAAGIFLVARLLPLFIVIPSIMYIISLIGIITVLLGATLALAQKD 314
> SPAC6F6.01
Length=1854
Score = 28.1 bits (61), Expect = 5.7, Method: Composition-based stats.
Identities = 20/87 (22%), Positives = 38/87 (43%), Gaps = 5/87 (5%)
Query 32 FIIVVVEVLLWAVRATQEKDKSEREQQAVVVVLDRRIPETCVHLYTYCGSLAMSVASVFL 91
F+++V+ +L +R+ DKS+ ++V+ E + +Y + G L +S +
Sbjct 192 FVVIVLHAVLLMIRSDDPHDKSQTIDYLIIVIGILYTLEMLLKIYLF-GFLYDGSSSFYD 250
Query 92 LTNSLKAAVGSLRPHFLDVCRPDWSRV 118
NS P + R W+RV
Sbjct 251 FINSYTKKT----PRTMSYLRHSWNRV 273
> 7291275
Length=1283
Score = 28.1 bits (61), Expect = 5.9, Method: Compositional matrix adjust.
Identities = 28/113 (24%), Positives = 54/113 (47%), Gaps = 5/113 (4%)
Query 24 WISTMLPAFIIVVVEVLLWAVRATQE-KDKSEREQQAVVVVLDRRIPE-TCVHLYTYCGS 81
W S + AFI++++ + WA +A+ +S + + A V +++ I + +Y + S
Sbjct 202 WTSLIGIAFIVILIPLQAWAAKASASFGTRSAKHRDARVKLMNEIIGAIQVIKMYAWEKS 261
Query 82 LAMSVASVFLLTNSLKAAVGSLRPHFLDVCRPDWSRVSCRDSSGLYDVYIPDF 134
+A+V + +KA GS+ + C S++S S Y VY+ D
Sbjct 262 FGRLIAAV--RQSEVKAIRGSMSIYAALQCTNMISKISLFLSLVAY-VYVGDL 311
> YNL321w
Length=908
Score = 28.1 bits (61), Expect = 7.3, Method: Composition-based stats.
Identities = 13/36 (36%), Positives = 21/36 (58%), Gaps = 0/36 (0%)
Query 15 ASISAAGAFWISTMLPAFIIVVVEVLLWAVRATQEK 50
ASISA + + ++ AF +VE+ L+ V Q+K
Sbjct 551 ASISAQTSMGVGAVINAFFSTIVEIFLYCVALQQKK 586
Lambda K H
0.325 0.137 0.430
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1858150626
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40