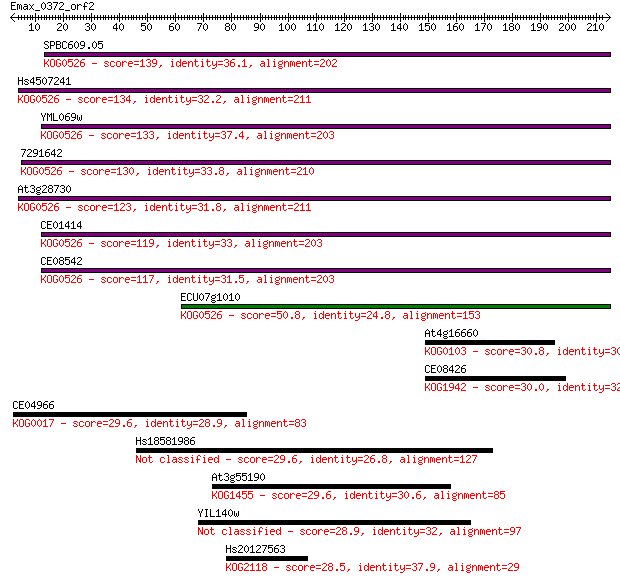
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Emax_0372_orf2
Length=214
Score E
Sequences producing significant alignments: (Bits) Value
SPBC609.05 139 5e-33
Hs4507241 134 2e-31
YML069w 133 2e-31
7291642 130 1e-30
At3g28730 123 2e-28
CE01414 119 3e-27
CE08542 117 2e-26
ECU07g1010 50.8 2e-06
At4g16660 30.8 1.9
CE08426 30.0 3.4
CE04966 29.6 4.6
Hs18581986 29.6 5.3
At3g55190 29.6 5.3
YIL140w 28.9 8.6
Hs20127563 28.5 9.6
> SPBC609.05
Length=512
Score = 139 bits (350), Expect = 5e-33, Method: Compositional matrix adjust.
Identities = 73/216 (33%), Positives = 126/216 (58%), Gaps = 14/216 (6%)
Query 13 KNDLAIELVQDDTR---DQTEDMLLEIRLFQP-------AAGDDETDSHLQSL-KEKLVK 61
KN++A+E D + D L+E+RL+ P AA +E + + +L E L +
Sbjct 143 KNEVALEFSTTDDKQIPSAQVDELVEMRLYVPGTTAKEDAADGEEVEQNAANLFYESLKE 202
Query 62 KSGVAETKMESIALLNDVPLLVPRGRYEIDIGRRALKFHGKSYDYTILYTNINRMFLVPR 121
++ + + ++I +++ LL PRGRY+ID+ ++ GK+YDY + Y++IN +FL+P+
Sbjct 203 RADIGQAAGDAIVSFSEILLLTPRGRYDIDMYETCMRLRGKTYDYKVEYSSINSLFLLPK 262
Query 122 PNSPHVNFILSLDNAMRQGQTSYPFVVMQFDSENVTSVEVNLEEAELKEKGLEK---QLE 178
P+ HV F++ L+ +RQGQT YPF+V QF + V++N+EE LKEK +K +
Sbjct 263 PDEQHVVFVIGLEPPLRQGQTRYPFLVTQFVRDEDMEVDLNIEETVLKEKYADKVKASYD 322
Query 179 GKTFDVITRILKSLIGKSVIVPGDFKSVKNQYGITC 214
F+V+++I + L G+ V P +F S + + C
Sbjct 323 QPAFEVVSQIFRGLTGRKVTTPAEFLSHEGHAAVKC 358
> Hs4507241
Length=709
Score = 134 bits (337), Expect = 2e-31, Method: Compositional matrix adjust.
Identities = 68/214 (31%), Positives = 126/214 (58%), Gaps = 8/214 (3%)
Query 4 DIVQVTTPSKNDLAIELVQDDTRDQTEDMLLEIRLFQPAAGDDETDSHLQSLKEKLVKKS 63
++ Q TT KN++ +E Q+D E L+E+R + P +D D +++ + ++ K+
Sbjct 135 NVSQCTT-GKNEVTLEFHQND---DAEVSLMEVRFYVPPTQEDGVDP-VEAFAQNVLSKA 189
Query 64 GVAETKMESIALLNDVPLLVPRGRYEIDIGRRALKFHGKSYDYTILYTNINRMFLVPRPN 123
V + ++I + ++ L PRGRY+I I L HGK++DY I YT + R+FL+P +
Sbjct 190 DVIQATGDAICIFRELQCLTPRGRYDIRIYPTFLHLHGKTFDYKIPYTTVLRLFLLPHKD 249
Query 124 SPHVNFILSLDNAMRQGQTSYPFVVMQFDSENVTSVEVNLEEAELK---EKGLEKQLEGK 180
+ F++SLD ++QGQT Y F+++ F + S+ +N+ E E++ E L K + G
Sbjct 250 QRQMFFVISLDPPIKQGQTRYHFLILLFSKDEDISLTLNMNEEEVEKRFEGRLTKNMSGS 309
Query 181 TFDVITRILKSLIGKSVIVPGDFKSVKNQYGITC 214
+++++R++K+L+ + + VPG+F+ ITC
Sbjct 310 LYEMVSRVMKALVNRKITVPGNFQGHSGAQCITC 343
> YML069w
Length=552
Score = 133 bits (335), Expect = 2e-31, Method: Compositional matrix adjust.
Identities = 76/240 (31%), Positives = 130/240 (54%), Gaps = 37/240 (15%)
Query 12 SKNDLAIEL-VQDDTRDQTEDMLLEIRLFQP----------------------------A 42
SKN++ IE +QD+ D L+E+R + P A
Sbjct 147 SKNEVGIEFNIQDEEYQPAGDELVEMRFYIPGVIQTNVDENMTKKEESSNEVVPKKEDGA 206
Query 43 AGDD-----ETDSHLQSLKEKLVKKSGVAETKMESIALLNDVPLLVPRGRYEIDIGRRAL 97
G+D E S ++ E+L +K+ + E ++I DV PRGRY+IDI + ++
Sbjct 207 EGEDVQMAVEEKSMAEAFYEELKEKADIGEVAGDAIVSFQDVFFTTPRGRYDIDIYKNSI 266
Query 98 KFHGKSYDYTILYTNINRMFLVPRPNSPHVNFILSLDNAMRQGQTSYPFVVMQFDSENVT 157
+ GK+Y+Y + + I R+ +P+ + H +L+++ +RQGQT+YPF+V+QF + T
Sbjct 267 RLRGKTYEYKLQHRQIQRIVSLPKADDIHHLLVLAIEPPLRQGQTTYPFLVLQFQKDEET 326
Query 158 SVEVNLEEAELKEK---GLEKQLEGKTFDVITRILKSLIGKSVIVPGDFKSVKNQYGITC 214
V++NLE+ + +E L+KQ + KT V++ +LK L + VIVPG++KS +Q ++C
Sbjct 327 EVQLNLEDEDYEENYKDKLKKQYDAKTHIVLSHVLKGLTDRRVIVPGEYKSKYDQCAVSC 386
> 7291642
Length=723
Score = 130 bits (328), Expect = 1e-30, Method: Compositional matrix adjust.
Identities = 71/213 (33%), Positives = 121/213 (56%), Gaps = 8/213 (3%)
Query 5 IVQVTTPSKNDLAIELVQDDTRDQTEDMLLEIRLFQPAAGDDETDSHLQSLKEKLVKKSG 64
+ Q T KN++ +E Q+D LLE+R PA E D + + ++ K+
Sbjct 136 VSQCVT-GKNEVTLEFHQND---DAPVGLLEMRFHIPAVESAEEDP-VDKFHQNVMSKAS 190
Query 65 VAETKMESIALLNDVPLLVPRGRYEIDIGRRALKFHGKSYDYTILYTNINRMFLVPRPNS 124
V ESIA+ ++ +L PRGRY+I I + HGK++DY I ++ R+F++P +S
Sbjct 191 VISASGESIAIFREIQILTPRGRYDIKIFSTFFQLHGKTFDYKIPMDSVLRLFMLPHKDS 250
Query 125 PHVNFILSLDNAMRQGQTSYPFVVMQFDSENVTSVEVNLEEAELKEK---GLEKQLEGKT 181
+ F+LSLD ++QGQT Y ++V+ F + T++E+ EAEL++K LEK++ G
Sbjct 251 RQMFFVLSLDPPIKQGQTRYHYLVLLFAPDEETTIELPFSEAELRDKYEGKLEKEISGPV 310
Query 182 FDVITRILKSLIGKSVIVPGDFKSVKNQYGITC 214
++V+ +++K LIG+ + PG+F + C
Sbjct 311 YEVMGKVMKVLIGRKITGPGNFIGHSGTAAVGC 343
> At3g28730
Length=646
Score = 123 bits (309), Expect = 2e-28, Method: Compositional matrix adjust.
Identities = 67/220 (30%), Positives = 119/220 (54%), Gaps = 10/220 (4%)
Query 4 DIVQVTTPSKNDLAIELVQDDTRDQTE-DMLLEIRLFQPAAG----DDETDSHLQSLKEK 58
D+ Q KND+ +E DDT E D L+EI P + DE Q +
Sbjct 135 DVSQTQLQGKNDVTLEFHVDDTAGANEKDSLMEISFHIPNSNTQFVGDENRPPSQVFNDT 194
Query 59 LVKKSGVAETKMESIALLNDVPLLVPRGRYEIDIGRRALKFHGKSYDYTILYTNINRMFL 118
+V + V+ +++ + +L PRGRY +++ L+ G++ D+ I Y+++ R+FL
Sbjct 195 IVAMADVSPGVEDAVVTFESIAILTPRGRYNVELHLSFLRLQGQANDFKIQYSSVVRLFL 254
Query 119 VPRPNSPHVNFILSLDNAMRQGQTSYPFVVMQFDSENVTSVEVNLEE----AELKEKGLE 174
+P+ N PH ++SLD +R+GQT YP +VMQF+++ V E+++ + + K+K LE
Sbjct 255 LPKSNQPHTFVVISLDPPIRKGQTMYPHIVMQFETDTVVESELSISDELMNTKFKDK-LE 313
Query 175 KQLEGKTFDVITRILKSLIGKSVIVPGDFKSVKNQYGITC 214
+ +G +V T +L+ L G + PG F+S ++ + +
Sbjct 314 RSYKGLIHEVFTTVLRWLSGAKITKPGKFRSSQDGFAVKS 353
> CE01414
Length=697
Score = 119 bits (299), Expect = 3e-27, Method: Compositional matrix adjust.
Identities = 67/207 (32%), Positives = 113/207 (54%), Gaps = 8/207 (3%)
Query 12 SKNDLAIELVQ-DDTRDQTEDMLLEIRLFQPAAGDDETDS-HLQSLKEKLVKKSGVAETK 69
+KN+ +E Q DD++ Q L+E+R P ++E D+ ++ K+ ++ +G+
Sbjct 141 NKNEAVLEFHQNDDSKVQ----LMEMRFHMPIDLENEEDADKVEEFKKAVLAYAGLEAET 196
Query 70 MESIALLNDVPLLVPRGRYEIDIGRRALKFHGKSYDYTILYTNINRMFLVPRPNSPHVNF 129
+ I LL D+ PRGRY+I + ++ HGK+YDY I +INR+FLVP + HV F
Sbjct 197 EQPICLLTDILCTTPRGRYDIKVYPTSIALHGKTYDYKIPIKSINRLFLVPHKDGRHVYF 256
Query 130 ILSLDNAMRQGQTSYPFVVMQFDSENVTSVEVNL--EEAELKEKGLEKQLEGKTFDVITR 187
+LSL+ +RQGQT Y +++ +F + +E+ L E+ E L + + G ++ I+
Sbjct 257 VLSLNPPIRQGQTRYSYLIFEFGKDEEQDLELALTDEQLESSNGNLRRDMTGPIYETISI 316
Query 188 ILKSLIGKSVIVPGDFKSVKNQYGITC 214
+ KS+ + VPG F I C
Sbjct 317 LFKSICNLKITVPGRFLGSSGTPAIQC 343
> CE08542
Length=689
Score = 117 bits (293), Expect = 2e-26, Method: Compositional matrix adjust.
Identities = 64/206 (31%), Positives = 114/206 (55%), Gaps = 6/206 (2%)
Query 12 SKNDLAIELVQDDTRDQTEDMLLEIRLFQPAAGDDETDS-HLQSLKEKLVKKSGVAETKM 70
+KN+ +E Q++ Q++ L+E+R P ++E D+ ++ K+ ++ +G+
Sbjct 141 NKNEAILEFHQNE---QSKVQLMEMRFHMPVDLENEEDTDKVEEFKKAVLAYAGLEAETE 197
Query 71 ESIALLNDVPLLVPRGRYEIDIGRRALKFHGKSYDYTILYTNINRMFLVPRPNSPHVNFI 130
+ I LL D+ PRGRY+I + ++ HGK+YDY I INR+FLVP + V F+
Sbjct 198 QPICLLTDILCTTPRGRYDIKVYPTSIALHGKTYDYKIPVKTINRLFLVPHKDGRQVYFV 257
Query 131 LSLDNAMRQGQTSYPFVVMQFDSENVTSVEVNLEEAELK--EKGLEKQLEGKTFDVITRI 188
LSL+ +RQGQT Y +++ +F + +E++L + +L L++++ G ++ I+ +
Sbjct 258 LSLNPPIRQGQTHYSYLIFEFGKDEEEDLELSLTDEQLDYFNGNLQREMTGPIYETISIL 317
Query 189 LKSLIGKSVIVPGDFKSVKNQYGITC 214
KS+ V VPG F I C
Sbjct 318 FKSICNLKVTVPGRFLGSSGTPAIQC 343
> ECU07g1010
Length=425
Score = 50.8 bits (120), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 38/157 (24%), Positives = 74/157 (47%), Gaps = 18/157 (11%)
Query 62 KSGVAETKMESIALLNDVPLLVPRGRYEIDIGRRALKFHGKSYDYTILYTNINRMFLVP- 120
K G + + + I + + L PRG++ R L+ G SYD+ I Y +I ++++
Sbjct 163 KGGCSSSVDDEILKMEGLSLAYPRGKFNFIFFRDYLRLVGSSYDHKIYYKSIKMLYVLEK 222
Query 121 ---RPNSPHVNFILSLDNAMRQGQTSYPFVVMQFDSENVTSVEVNLEEAELKEKGLEKQL 177
R +V ++ D +RQGQT Y VV+ FD VE L + +++L
Sbjct 223 GYIRDGERYV--VIGADPPIRQGQTRYDHVVVAFD-----DVERELSVS-------DERL 268
Query 178 EGKTFDVITRILKSLIGKSVIVPGDFKSVKNQYGITC 214
+G+ +++ I ++ ++ S +++ G+ C
Sbjct 269 KGEYSGLLSEIFAEVMEALCVIKAVRSSFESRDGMRC 305
> At4g16660
Length=400
Score = 30.8 bits (68), Expect = 1.9, Method: Compositional matrix adjust.
Identities = 14/46 (30%), Positives = 27/46 (58%), Gaps = 0/46 (0%)
Query 149 MQFDSENVTSVEVNLEEAELKEKGLEKQLEGKTFDVITRILKSLIG 194
+Q D+EN T+ EE + G EK+L+ +TF + ++++ +G
Sbjct 117 LQTDAENSTASNTTAEEPAVASLGTEKKLKKRTFRIPLKVVEKTVG 162
> CE08426
Length=458
Score = 30.0 bits (66), Expect = 3.4, Method: Compositional matrix adjust.
Identities = 16/50 (32%), Positives = 27/50 (54%), Gaps = 6/50 (12%)
Query 149 MQFDSENVTSVEVNLEEAELKEKGLEKQLEGKTFDVITRILKSLIGKSVI 198
M+++ E++ + V+ EAE Q E K FD++TR+ G+ VI
Sbjct 382 MKYNEEDIRKILVHRTEAE------NVQFEEKAFDLLTRLCAQTCGREVI 425
> CE04966
Length=2037
Score = 29.6 bits (65), Expect = 4.6, Method: Compositional matrix adjust.
Identities = 24/84 (28%), Positives = 38/84 (45%), Gaps = 13/84 (15%)
Query 2 AQDIVQVTTPSKNDLAIELVQDDTRDQTEDMLLEIRLFQPAAGDDETDSHLQSLKEKLVK 61
++D PS+ DL +EL + D D T D+ + H +S L++
Sbjct 1961 SEDFTSADPPSEGDL-VELEEHDCDDPTSDIPTQAYF-----------EHSRSTARTLLR 2008
Query 62 KSG-VAETKMESIALLNDVPLLVP 84
S V+E + S+ + D PLLVP
Sbjct 2009 VSPRVSEIRCLSLGRVTDSPLLVP 2032
> Hs18581986
Length=222
Score = 29.6 bits (65), Expect = 5.3, Method: Compositional matrix adjust.
Identities = 34/138 (24%), Positives = 60/138 (43%), Gaps = 12/138 (8%)
Query 46 DETDSHLQSLKEKLVK------KSGVAETKMESIALLND--VPLLVPRGRYEIDIGRRAL 97
DE D S K L+ + +A K ++++L D +L R+E+DI L
Sbjct 10 DELDEPTHSGKTNLMANTLSEIQGNLAHLKSDNMSLTGDFAASILPDDRRFEVDISALHL 69
Query 98 KFHGKSYDYTILYTNINRMFLVPRPNSPHVNFILSLDNAMRQGQTS---YPFVVMQFDSE 154
GK + +++T F V P+ ++ I ++ ++R S P M+ DS
Sbjct 70 DLLGKGHKPPVVHTGAVLDFRVHAPHLKSLSVIAAVSPSVRAQHISKGDAPGCCME-DSM 128
Query 155 NVTSVEVNLEEAELKEKG 172
+V+ + N E E+ G
Sbjct 129 SVSEGQGNDEAVEVMFSG 146
> At3g55190
Length=319
Score = 29.6 bits (65), Expect = 5.3, Method: Compositional matrix adjust.
Identities = 26/88 (29%), Positives = 43/88 (48%), Gaps = 6/88 (6%)
Query 73 IALLNDVPLLVPRGRYEI---DIGRRALKFHGKSYDYTILYTNINRMFLVPRPNSPHVNF 129
I+++N V L+P + I DI A+K K ++ + TN N PR + F
Sbjct 159 ISMINMVTNLIPSWKSIIHGPDILNSAIKLPEKRHE---IRTNPNCYNGWPRMKTMSELF 215
Query 130 ILSLDNAMRQGQTSYPFVVMQFDSENVT 157
+SLD R + + PF+V+ + + VT
Sbjct 216 RISLDLENRLNEVTMPFIVLHGEDDKVT 243
> YIL140w
Length=823
Score = 28.9 bits (63), Expect = 8.6, Method: Compositional matrix adjust.
Identities = 31/117 (26%), Positives = 50/117 (42%), Gaps = 20/117 (17%)
Query 68 TKMESIALLNDVPLLVPRGRYEIDIGRRALKFHGK-----SYDYTIL--------YTNIN 114
T SI+L +D LL Y G+ ALK ++D ++ Y +
Sbjct 125 TNRPSISLSSDFNLLALLKNYGYTNGKNALKLDPNEVFNVTFDRSMFTNEESIVSYYGRS 184
Query 115 RMFLVPRPN-----SPHVNFILSLD--NAMRQGQTSYPFVVMQFDSENVTSVEVNLE 164
+++ P PN S + F + N+ +TSY FV++ D E ++VEV E
Sbjct 185 QLYNAPLPNWLFFDSGELKFTGTAPVINSAIAPETSYSFVIIATDIEGFSAVEVEFE 241
> Hs20127563
Length=382
Score = 28.5 bits (62), Expect = 9.6, Method: Compositional matrix adjust.
Identities = 11/29 (37%), Positives = 19/29 (65%), Gaps = 0/29 (0%)
Query 78 DVPLLVPRGRYEIDIGRRALKFHGKSYDY 106
D +L+ +GR E++IG+ LKF ++ Y
Sbjct 231 DYFILILQGRVEVEIGKEGLKFENGAFTY 259
Lambda K H
0.316 0.134 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 3933248660
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40