bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Emax_0170_orf1
Length=110
Score E
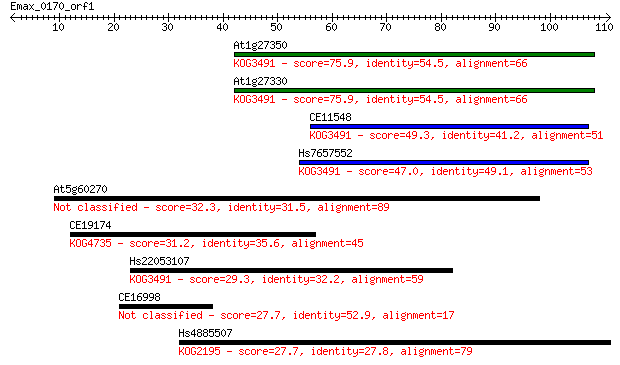
Sequences producing significant alignments: (Bits) Value
At1g27350 75.9 2e-14
At1g27330 75.9 2e-14
CE11548 49.3 2e-06
Hs7657552 47.0 8e-06
At5g60270 32.3 0.20
CE19174 31.2 0.44
Hs22053107 29.3 2.1
CE16998 27.7 4.8
Hs4885507 27.7 6.2
> At1g27350
Length=68
Score = 75.9 bits (185), Expect = 2e-14, Method: Compositional matrix adjust.
Identities = 36/66 (54%), Positives = 46/66 (69%), Gaps = 0/66 (0%)
Query 42 MAGLNRKYSKKIEAFDRNITKRGQVSESKQKRGKGYPVGPIVLGLFLFVVVGSAIIQIIS 101
M R +KIE FD+NI KRG V E+ K+GK YPVGPI+LG F+FVV+GS++ QII
Sbjct 1 MTTSKRLADRKIEKFDKNILKRGFVPETTTKKGKDYPVGPILLGFFVFVVIGSSLFQIIR 60
Query 102 SAQRGA 107
+A G
Sbjct 61 TATSGG 66
> At1g27330
Length=68
Score = 75.9 bits (185), Expect = 2e-14, Method: Compositional matrix adjust.
Identities = 36/66 (54%), Positives = 46/66 (69%), Gaps = 0/66 (0%)
Query 42 MAGLNRKYSKKIEAFDRNITKRGQVSESKQKRGKGYPVGPIVLGLFLFVVVGSAIIQIIS 101
M R +KIE FD+NI KRG V E+ K+GK YPVGPI+LG F+FVV+GS++ QII
Sbjct 1 MTTSKRLADRKIEKFDKNILKRGFVPETTTKKGKDYPVGPILLGFFVFVVIGSSLFQIIR 60
Query 102 SAQRGA 107
+A G
Sbjct 61 TATSGG 66
> CE11548
Length=65
Score = 49.3 bits (116), Expect = 2e-06, Method: Compositional matrix adjust.
Identities = 21/51 (41%), Positives = 32/51 (62%), Gaps = 0/51 (0%)
Query 56 FDRNITKRGQVSESKQKRGKGYPVGPIVLGLFLFVVVGSAIIQIISSAQRG 106
F +N+ RG V++S + YP P ++GLF+FVV GSA+ +II + G
Sbjct 14 FSKNVNNRGNVAKSLKPAEDKYPAAPWLIGLFVFVVCGSAVFEIIRYVKMG 64
> Hs7657552
Length=66
Score = 47.0 bits (110), Expect = 8e-06, Method: Compositional matrix adjust.
Identities = 26/54 (48%), Positives = 35/54 (64%), Gaps = 1/54 (1%)
Query 54 EAFDRNITKRGQVSE-SKQKRGKGYPVGPIVLGLFLFVVVGSAIIQIISSAQRG 106
E +NIT+RG V++ S+ + VGP +L LF+FVV GSAI QII S + G
Sbjct 12 EKHSKNITQRGNVAKTSRNAPEEKASVGPWLLALFIFVVCGSAIFQIIQSIRMG 65
> At5g60270
Length=668
Score = 32.3 bits (72), Expect = 0.20, Method: Composition-based stats.
Identities = 28/101 (27%), Positives = 45/101 (44%), Gaps = 12/101 (11%)
Query 9 LLPLQNRVPCN---RKHRHRLHHFQKGFRQFVPISKMAG--------LNRKYSKKIEAF- 56
L PL+N+ P L +G R FV S G L +SK + +
Sbjct 204 LAPLRNKKPSRPLLSSTSINLTDILQGRRMFVGFSGSTGSSMSYQYILGWSFSKSMASLP 263
Query 57 DRNITKRGQVSESKQKRGKGYPVGPIVLGLFLFVVVGSAII 97
+ +I+K +V S K+ PV ++LGL F+V+G ++
Sbjct 264 NIDISKLPKVPHSSTKKKSTSPVLSVLLGLIAFIVLGILVV 304
> CE19174
Length=479
Score = 31.2 bits (69), Expect = 0.44, Method: Composition-based stats.
Identities = 16/48 (33%), Positives = 25/48 (52%), Gaps = 3/48 (6%)
Query 12 LQNRVPCNRKHRHRLHHFQKGFRQFVPI--SKMAG-LNRKYSKKIEAF 56
+Q RVPC + H + L H PI S +A LN+++ ++E F
Sbjct 338 VQTRVPCRQAHFYHLRHSHNTVPSPTPINMSPLADMLNKQWQTRVEGF 385
> Hs22053107
Length=117
Score = 29.3 bits (64), Expect = 2.1, Method: Compositional matrix adjust.
Identities = 19/59 (32%), Positives = 29/59 (49%), Gaps = 1/59 (1%)
Query 23 RHRLHHFQKGFRQFVPISKMAGLNRKYSKKIEAFDRNITKRGQVSESKQKRGKGYPVGP 81
R RL + +G F M R E +NIT+RG V+++ + + + YPVGP
Sbjct 60 RSRLPRWPRGVGAFQSAQAMVAKQR-IRMANEKHSKNITQRGNVAKTLRPQEEKYPVGP 117
> CE16998
Length=669
Score = 27.7 bits (60), Expect = 4.8, Method: Composition-based stats.
Identities = 9/17 (52%), Positives = 13/17 (76%), Gaps = 0/17 (0%)
Query 21 KHRHRLHHFQKGFRQFV 37
KH H L HF+K F++F+
Sbjct 635 KHPHNLEHFKKNFKEFL 651
> Hs4885507
Length=740
Score = 27.7 bits (60), Expect = 6.2, Method: Composition-based stats.
Identities = 22/89 (24%), Positives = 41/89 (46%), Gaps = 10/89 (11%)
Query 32 GFRQFVPISKMAG-----LNRKYSKKIEAFDRNITKRGQVSESKQKRGK-----GYPVGP 81
GF +++ AG L+ + ++ D + T R + ++Q G +GP
Sbjct 561 GFSSHQAVARTAGSVILRLSDSFFLPLKVSDYSETLRSFLQAAQQDLGALLEQHSISLGP 620
Query 82 IVLGLFLFVVVGSAIIQIISSAQRGAPLP 110
+V + F +A+ Q IS+ Q+G+P P
Sbjct 621 LVTAVEKFEAEAAALGQRISTLQKGSPDP 649
Lambda K H
0.326 0.143 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1195973986
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40