bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Emax_0103_orf1
Length=73
Score E
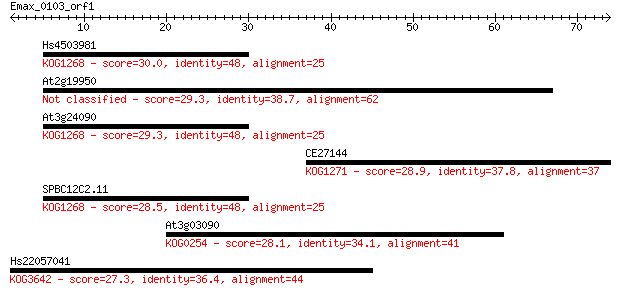
Sequences producing significant alignments: (Bits) Value
Hs4503981 30.0 1.1
At2g19950 29.3 1.8
At3g24090 29.3 1.9
CE27144 28.9 2.6
SPBC12C2.11 28.5 2.8
At3g03090 28.1 4.1
Hs22057041 27.3 7.8
> Hs4503981
Length=681
Score = 30.0 bits (66), Expect = 1.1, Method: Composition-based stats.
Identities = 12/25 (48%), Positives = 15/25 (60%), Gaps = 0/25 (0%)
Query 5 HEQTSHSLWAFTGEPRPLNLCPQLS 29
H +H+ WA GEP P+N PQ S
Sbjct 88 HLGIAHTRWATHGEPSPVNSHPQRS 112
> At2g19950
Length=713
Score = 29.3 bits (64), Expect = 1.8, Method: Composition-based stats.
Identities = 24/66 (36%), Positives = 31/66 (46%), Gaps = 8/66 (12%)
Query 5 HEQTSHSLWAFTGEPRPLNLCPQLSMF----TVLFLLKAFLVLFRNGVKLAEGFLVFFCG 60
HE +H TG R L +LS F T++F L AF + +N VKL + V
Sbjct 607 HEAQTHGYSEHTG--RDLGAHYELSAFSFNFTLMFALFAFCLQLQNAVKLLDSGAVR--A 662
Query 61 FDFLWR 66
FLWR
Sbjct 663 TRFLWR 668
> At3g24090
Length=691
Score = 29.3 bits (64), Expect = 1.9, Method: Composition-based stats.
Identities = 12/25 (48%), Positives = 14/25 (56%), Gaps = 0/25 (0%)
Query 5 HEQTSHSLWAFTGEPRPLNLCPQLS 29
H +H+ WA GEP P N PQ S
Sbjct 81 HAGIAHTRWATHGEPAPRNSHPQSS 105
> CE27144
Length=236
Score = 28.9 bits (63), Expect = 2.6, Method: Composition-based stats.
Identities = 14/37 (37%), Positives = 19/37 (51%), Gaps = 6/37 (16%)
Query 37 LKAFLVLFRNGVKLAEGFLVFFCGFDFLWRGNAEEMC 73
LKA+L NG+ F++F C F F +EMC
Sbjct 162 LKAYLGFLDNGLSAGGRFVIFSCNFTF------DEMC 192
> SPBC12C2.11
Length=696
Score = 28.5 bits (62), Expect = 2.8, Method: Composition-based stats.
Identities = 12/25 (48%), Positives = 14/25 (56%), Gaps = 0/25 (0%)
Query 5 HEQTSHSLWAFTGEPRPLNLCPQLS 29
H SH+ WA G P P+N PQ S
Sbjct 80 HCAISHTRWATHGIPSPINCHPQRS 104
> At3g03090
Length=342
Score = 28.1 bits (61), Expect = 4.1, Method: Composition-based stats.
Identities = 14/41 (34%), Positives = 22/41 (53%), Gaps = 0/41 (0%)
Query 20 RPLNLCPQLSMFTVLFLLKAFLVLFRNGVKLAEGFLVFFCG 60
RPL LC M LFLL ++ + ++N +A L+ + G
Sbjct 210 RPLLLCGVSGMVISLFLLGSYYMFYKNVPAVAVAALLLYVG 250
> Hs22057041
Length=377
Score = 27.3 bits (59), Expect = 7.8, Method: Compositional matrix adjust.
Identities = 16/44 (36%), Positives = 21/44 (47%), Gaps = 0/44 (0%)
Query 1 AHSRHEQTSHSLWAFTGEPRPLNLCPQLSMFTVLFLLKAFLVLF 44
AH ++H+ A P PL L PQL M + L +A L F
Sbjct 218 AHDLKVGSNHTSKAAVRSPFPLGLSPQLIMIVQIALHQALLTGF 261
Lambda K H
0.332 0.144 0.489
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1187734408
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40