bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
164,496 sequences; 82,071,388 total letters
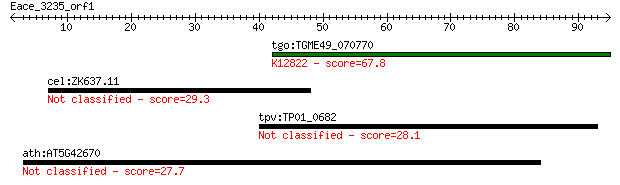
Query= Eace_3235_orf1
Length=94
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_070770 hypothetical protein ; K12822 RNA-binding pr... 67.8 8e-12
cel:ZK637.11 cdc-25.3; Cell Division Cycle related family memb... 29.3 3.5
tpv:TP01_0682 60S ribosomal protein L17 28.1 7.7
ath:AT5G42670 agenet domain-containing protein 27.7 9.5
> tgo:TGME49_070770 hypothetical protein ; K12822 RNA-binding
protein 25
Length=779
Score = 67.8 bits (164), Expect = 8e-12, Method: Compositional matrix adjust.
Identities = 30/53 (56%), Positives = 39/53 (73%), Gaps = 0/53 (0%)
Query 42 LRKLDPEAIAAAAENSIYRIAPDGTDRRTYIEEDYRPSRRELERLQNAGRREA 94
L+ ++PE AA+ S+YR+ PDG DRR YI EDYRP RRELERL + RR++
Sbjct 375 LKAMNPELAEAASSCSVYRVGPDGVDRRRYITEDYRPCRRELERLHHLERRDS 427
> cel:ZK637.11 cdc-25.3; Cell Division Cycle related family member
(cdc-25.3)
Length=316
Score = 29.3 bits (64), Expect = 3.5, Method: Compositional matrix adjust.
Identities = 15/42 (35%), Positives = 24/42 (57%), Gaps = 1/42 (2%)
Query 7 QHSSHKKESSKDKHKYSSSSKTAATDTSAGSAFHRLRK-LDP 47
+H HKK SK K+SS++ + T++G+ LR+ DP
Sbjct 268 KHQFHKKNVSKPMKKWSSTTSVISILTTSGTRISTLRQTCDP 309
> tpv:TP01_0682 60S ribosomal protein L17
Length=260
Score = 28.1 bits (61), Expect = 7.7, Method: Compositional matrix adjust.
Identities = 17/57 (29%), Positives = 28/57 (49%), Gaps = 4/57 (7%)
Query 40 HRLRKLDPEAIA----AAAENSIYRIAPDGTDRRTYIEEDYRPSRRELERLQNAGRR 92
++LR+ D +A EN IY P G D+ +I + +RR+ + N GR+
Sbjct 124 YKLRQRDSAPLAYIELVNTENEIYPAKPVGIDKLNHILNLMKSTRRDYRKYHNYGRK 180
> ath:AT5G42670 agenet domain-containing protein
Length=294
Score = 27.7 bits (60), Expect = 9.5, Method: Composition-based stats.
Identities = 23/94 (24%), Positives = 41/94 (43%), Gaps = 13/94 (13%)
Query 3 ELNQQHSSHKKESSKDKHKYSS---SSKTAATDTSAGSAFHRLRKLDPEAIAAAA---EN 56
+LN+ S + ++K + + K T+ +L +D E+I AA E
Sbjct 201 QLNEDQVSEAESMEEEKQRGAEPKEEEKQRPTEKLKSDEALQLNIMDSESIEAAVLDLEE 260
Query 57 SIYR-------IAPDGTDRRTYIEEDYRPSRREL 83
I R + PD +++ ++I EDY PS +
Sbjct 261 MIVRFEWLKDILTPDSSEKNSWIYEDYHPSSSRM 294
Lambda K H
0.306 0.121 0.324
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 2069995292
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
Posted date: Sep 16, 2011 8:45 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40