bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
164,496 sequences; 82,071,388 total letters
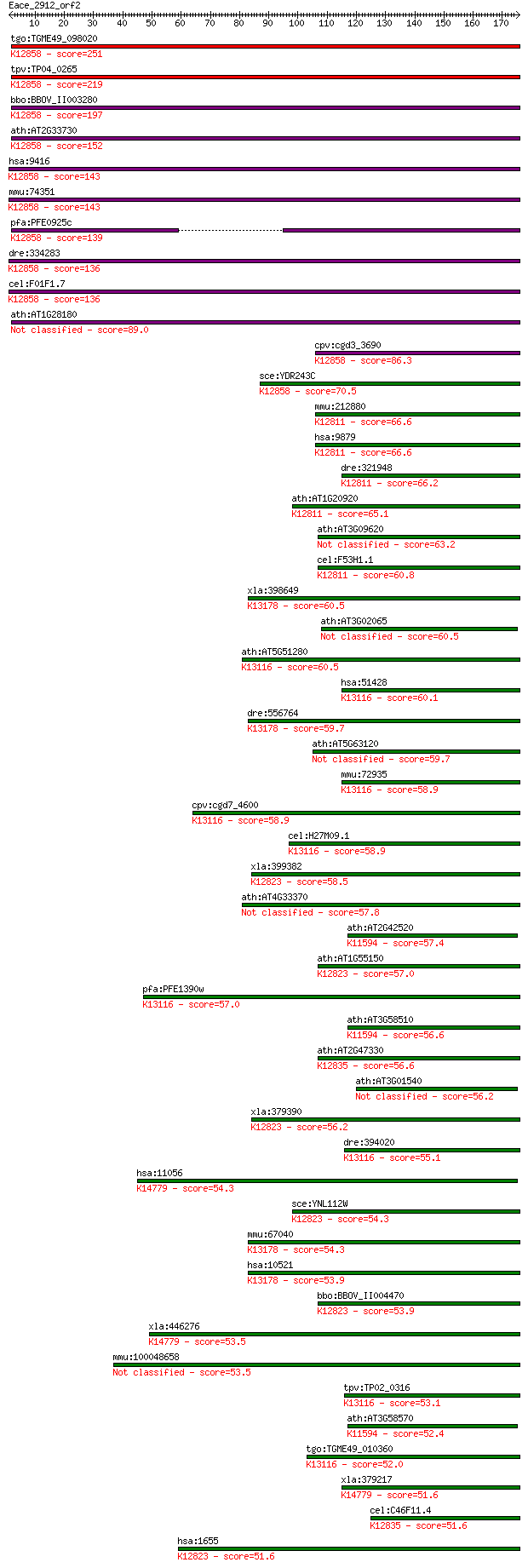
Query= Eace_2912_orf2
Length=175
Score E
Sequences producing significant alignments: (Bits) Value
tgo:TGME49_098020 DEAD-box ATP-dependent RNA helicase, putativ... 251 7e-67
tpv:TP04_0265 small nuclear ribonucleoprotein; K12858 ATP-depe... 219 5e-57
bbo:BBOV_II003280 18.m06276; DEAD box RNA helicase; K12858 ATP... 197 2e-50
ath:AT2G33730 DEAD box RNA helicase, putative; K12858 ATP-depe... 152 6e-37
hsa:9416 DDX23, MGC8416, PRPF28, U5-100K, U5-100KD, prp28; DEA... 143 3e-34
mmu:74351 Ddx23, 3110082M05Rik, 4921506D17Rik; DEAD (Asp-Glu-A... 143 3e-34
pfa:PFE0925c snrnp protein, putative; K12858 ATP-dependent RNA... 139 5e-33
dre:334283 ddx23, wu:fi39b12, zgc:63742; DEAD (Asp-Glu-Ala-Asp... 136 4e-32
cel:F01F1.7 ddx-23; DEAD boX helicase homolog family member (d... 136 5e-32
ath:AT1G28180 ATP binding / ATP-dependent helicase/ helicase/ ... 89.0 8e-18
cpv:cgd3_3690 U5 snRNP 100 kD protein ; K12858 ATP-dependent R... 86.3 5e-17
sce:YDR243C PRP28; Prp28p (EC:3.6.1.-); K12858 ATP-dependent R... 70.5 3e-12
mmu:212880 Ddx46, 2200005K02Rik, 8430438J23Rik, AI325430, AI95... 66.6 4e-11
hsa:9879 DDX46, FLJ25329, KIAA0801, MGC9936, PRPF5, Prp5; DEAD... 66.6 4e-11
dre:321948 ddx46, fb39a03, wu:fb39a03; DEAD (Asp-Glu-Ala-Asp) ... 66.2 5e-11
ath:AT1G20920 DEAD box RNA helicase, putative; K12811 ATP-depe... 65.1 1e-10
ath:AT3G09620 DEAD/DEAH box helicase, putative 63.2 4e-10
cel:F53H1.1 hypothetical protein; K12811 ATP-dependent RNA hel... 60.8 2e-09
xla:398649 ddx17, MGC80019; DEAD (Asp-Glu-Ala-Asp) box polypep... 60.5 3e-09
ath:AT3G02065 DEAD/DEAH box helicase family protein 60.5 3e-09
ath:AT5G51280 DEAD-box protein abstrakt, putative; K13116 ATP-... 60.5 3e-09
hsa:51428 DDX41, ABS, MGC8828; DEAD (Asp-Glu-Ala-Asp) box poly... 60.1 4e-09
dre:556764 similar to Probable RNA-dependent helicase p72 (DEA... 59.7 4e-09
ath:AT5G63120 ethylene-responsive DEAD box RNA helicase, putat... 59.7 5e-09
mmu:72935 Ddx41, 2900024F02Rik, AA958953, ABS, AI324246; DEAD ... 58.9 8e-09
cpv:cgd7_4600 abstrakt protein SF II helicase + Znknuckle C2HC... 58.9 9e-09
cel:H27M09.1 hypothetical protein; K13116 ATP-dependent RNA he... 58.9 9e-09
xla:399382 ddx5, MGC81559; DEAD (Asp-Glu-Ala-Asp) box polypept... 58.5 1e-08
ath:AT4G33370 DEAD-box protein abstrakt, putative 57.8 2e-08
ath:AT2G42520 DEAD box RNA helicase, putative; K11594 ATP-depe... 57.4 3e-08
ath:AT1G55150 DEAD box RNA helicase, putative (RH20); K12823 A... 57.0 3e-08
pfa:PFE1390w RNA helicase-1; K13116 ATP-dependent RNA helicase... 57.0 3e-08
ath:AT3G58510 DEAD box RNA helicase, putative (RH11); K11594 A... 56.6 4e-08
ath:AT2G47330 DEAD/DEAH box helicase, putative; K12835 ATP-dep... 56.6 5e-08
ath:AT3G01540 DRH1; DRH1 (DEAD BOX RNA HELICASE 1); ATP-depend... 56.2 5e-08
xla:379390 MGC53795; similar to DEAD/H (Asp-Glu-Ala-Asp/His) b... 56.2 6e-08
dre:394020 ddx41, MGC55896, wu:fb92e02, zgc:55896; DEAD (Asp-G... 55.1 1e-07
hsa:11056 DDX52, HUSSY19, ROK1; DEAD (Asp-Glu-Ala-Asp) box pol... 54.3 2e-07
sce:YNL112W DBP2; Dbp2p (EC:3.6.1.-); K12823 ATP-dependent RNA... 54.3 2e-07
mmu:67040 Ddx17, 2610007K22Rik, A430025E01Rik, AI047725, C8092... 54.3 2e-07
hsa:10521 DDX17, DKFZp761H2016, P72, RH70; DEAD (Asp-Glu-Ala-A... 53.9 3e-07
bbo:BBOV_II004470 18.m06373; p68-like protein; K12823 ATP-depe... 53.9 3e-07
xla:446276 ddx52; DEAD (Asp-Glu-Ala-Asp) box polypeptide 52; K... 53.5 4e-07
mmu:100048658 Ddx43, OTTMUSG00000019690; DEAD (Asp-Glu-Ala-Asp... 53.5 4e-07
tpv:TP02_0316 RNA helicase-1; K13116 ATP-dependent RNA helicas... 53.1 5e-07
ath:AT3G58570 DEAD box RNA helicase, putative; K11594 ATP-depe... 52.4 7e-07
tgo:TGME49_010360 DEAD/DEAH box helicase, putative (EC:5.99.1.... 52.0 1e-06
xla:379217 MGC53409; similar to ATP-dependent, RNA helicase; K... 51.6 1e-06
cel:C46F11.4 hypothetical protein; K12835 ATP-dependent RNA he... 51.6 1e-06
hsa:1655 DDX5, DKFZp434E109, DKFZp686J01190, G17P1, HLR1, HUMP... 51.6 1e-06
> tgo:TGME49_098020 DEAD-box ATP-dependent RNA helicase, putative
(EC:2.7.11.25); K12858 ATP-dependent RNA helicase DDX23/PRP28
[EC:3.6.4.13]
Length=1158
Score = 251 bits (642), Expect = 7e-67, Method: Compositional matrix adjust.
Identities = 128/174 (73%), Positives = 145/174 (83%), Gaps = 1/174 (0%)
Query 2 SEDTMRGDNNPLYQNRLEPQLLFGRGFRAGMDIREQRKANNFYDELVKRRQEYELKQRQR 61
+EDT +GDNNPLYQ R+EPQLLFGRGFRAGMDIREQRK NNFYDELVKRRQE++ K
Sbjct 607 AEDTCKGDNNPLYQERMEPQLLFGRGFRAGMDIREQRKQNNFYDELVKRRQEHQ-KAEAS 665
Query 62 GAEEAAAAAATAAAEAARASDKKKVKDMEEEPELEGHWSTKKREEMTDRDWRIFREDFEI 121
AAAAAT A AAR + ++++ E+ + GHW+TKKREEM +RDWRIFREDFEI
Sbjct 666 RGAAEAAAAATEAVRAARDAQASRLREKEDAEDNRGHWTTKKREEMNERDWRIFREDFEI 725
Query 122 YLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
Y+KGGRVPPPIRTWAE+ LPWELIEAVK ANY+RPTPIQMQAIPIALE RDLIG
Sbjct 726 YIKGGRVPPPIRTWAESALPWELIEAVKHANYDRPTPIQMQAIPIALEQRDLIG 779
> tpv:TP04_0265 small nuclear ribonucleoprotein; K12858 ATP-dependent
RNA helicase DDX23/PRP28 [EC:3.6.4.13]
Length=744
Score = 219 bits (557), Expect = 5e-57, Method: Compositional matrix adjust.
Identities = 107/175 (61%), Positives = 124/175 (70%), Gaps = 22/175 (12%)
Query 2 SEDTMRGDNNPLYQNRLEPQLLFGRGFRAGMDIREQRKANNFYDELVKRRQEYELKQRQR 61
SEDT + +NNP+YQ+R EPQLLFGRGFRAG+D+REQRK NNFYDEL ++R E +
Sbjct 218 SEDTTKFENNPIYQDRPEPQLLFGRGFRAGIDVREQRKKNNFYDELSRKRAELPQTTPKP 277
Query 62 GAEEAAAAAATAAAEAARASDKKKVKDMEEEPE-LEGHWSTKKREEMTDRDWRIFREDFE 120
EEE E L HW+ KK EMT+RDWRIFREDFE
Sbjct 278 PEPTPKH---------------------EEESEVLSNHWTKKKLSEMTERDWRIFREDFE 316
Query 121 IYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
IY+KGGRVPPPIRTWAE+ LPWEL+EA+K+A Y +PTPIQMQAIPIALEMRDLIG
Sbjct 317 IYIKGGRVPPPIRTWAESPLPWELLEAIKKAGYIKPTPIQMQAIPIALEMRDLIG 371
> bbo:BBOV_II003280 18.m06276; DEAD box RNA helicase; K12858 ATP-dependent
RNA helicase DDX23/PRP28 [EC:3.6.4.13]
Length=714
Score = 197 bits (501), Expect = 2e-50, Method: Compositional matrix adjust.
Identities = 97/178 (54%), Positives = 120/178 (67%), Gaps = 25/178 (14%)
Query 2 SEDTMRGDNNPLYQNRLEPQLLFGRGFRAGMDIREQRKANNFYDELVKRRQEYELKQRQR 61
S+DT R DNNP+YQNR EPQLLFGRG RAGMD +EQRK +FYD+L K R
Sbjct 183 SDDTSRNDNNPIYQNRPEPQLLFGRGCRAGMDPKEQRKHADFYDKLSKLR---------- 232
Query 62 GAEEAAAAAATAAAEAARASDKKKVKDMEE----EPELEGHWSTKKREEMTDRDWRIFRE 117
++AA + D + +D E + +++ HWS K +E MT RDWRIFRE
Sbjct 233 -----------TGSDAAFSRDSRSNEDRTEPTSVDDDVDTHWSAKTKENMTQRDWRIFRE 281
Query 118 DFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
DF+IY+KG RVPPP+RTWAE+ LP EL+ A+K A ++ PTPIQMQAIPI L MRDLIG
Sbjct 282 DFDIYVKGTRVPPPMRTWAESNLPSELLRAIKDAGFKSPTPIQMQAIPIGLGMRDLIG 339
> ath:AT2G33730 DEAD box RNA helicase, putative; K12858 ATP-dependent
RNA helicase DDX23/PRP28 [EC:3.6.4.13]
Length=733
Score = 152 bits (384), Expect = 6e-37, Method: Compositional matrix adjust.
Identities = 86/176 (48%), Positives = 113/176 (64%), Gaps = 15/176 (8%)
Query 2 SEDTMRGDNNPLYQNRLEPQLLFGRGFRAGMDIREQRKANNFYDELVKRRQEYELKQRQR 61
+EDT R D N LYQN E QLLFGRGFRAGMD REQ+K + E E++ R
Sbjct 193 TEDTSR-DMNVLYQNPHEAQLLFGRGFRAGMDRREQKKQAA--------KHEKEMRDEIR 243
Query 62 GAEEAAAAAATAAAEAAR--ASDKKKVKDMEEEPELEGHWSTKKREEMTDRDWRIFREDF 119
+ AAA+ R A+D DM ++ HWS K+ EEMT+RDWRIFREDF
Sbjct 244 KKDGIVEKPEEAAAQRVREEAADTYDSFDMR----VDRHWSDKRLEEMTERDWRIFREDF 299
Query 120 EIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
I KG R+P P+R+W E++L EL++AV++A Y++P+PIQM AIP+ L+ RD+IG
Sbjct 300 NISYKGSRIPRPMRSWEESKLTSELLKAVERAGYKKPSPIQMAAIPLGLQQRDVIG 355
> hsa:9416 DDX23, MGC8416, PRPF28, U5-100K, U5-100KD, prp28; DEAD
(Asp-Glu-Ala-Asp) box polypeptide 23 (EC:3.6.4.13); K12858
ATP-dependent RNA helicase DDX23/PRP28 [EC:3.6.4.13]
Length=820
Score = 143 bits (361), Expect = 3e-34, Method: Compositional matrix adjust.
Identities = 80/176 (45%), Positives = 110/176 (62%), Gaps = 21/176 (11%)
Query 1 ASEDTMRGDNNPLYQNRLEPQLLFGRGFRAGMDIREQ-RKANNFYDELVKRRQEYELKQR 59
ASEDT D NPLY+ R + QLL GRGF AG+D+++Q R+ + FY +L+++R+ E K++
Sbjct 278 ASEDTSI-DYNPLYKERHQVQLL-GRGFIAGIDLKQQKREQSRFYGDLMEKRRTLEEKEQ 335
Query 60 QRGAEEAAAAAATAAAEAARASDKKKVKDMEEEPELEGHWSTKKREEMTDRDWRIFREDF 119
EEA R D+ HWS KK +EMTDRDWRIFRED+
Sbjct 336 ----EEARLRKLRKKEAKQRWDDR--------------HWSQKKLDEMTDRDWRIFREDY 377
Query 120 EIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
I KGG++P PIR+W ++ LP ++E + + Y+ PTPIQ QAIPI L+ RD+IG
Sbjct 378 SITTKGGKIPNPIRSWKDSSLPPHILEVIDKCGYKEPTPIQRQAIPIGLQNRDIIG 433
> mmu:74351 Ddx23, 3110082M05Rik, 4921506D17Rik; DEAD (Asp-Glu-Ala-Asp)
box polypeptide 23 (EC:3.6.1.-); K12858 ATP-dependent
RNA helicase DDX23/PRP28 [EC:3.6.4.13]
Length=819
Score = 143 bits (360), Expect = 3e-34, Method: Compositional matrix adjust.
Identities = 80/176 (45%), Positives = 110/176 (62%), Gaps = 21/176 (11%)
Query 1 ASEDTMRGDNNPLYQNRLEPQLLFGRGFRAGMDIREQ-RKANNFYDELVKRRQEYELKQR 59
ASEDT D NPLY+ R + QLL GRGF AG+D+++Q R+ + FY +L+++R+ E K++
Sbjct 277 ASEDTSI-DYNPLYKERHQVQLL-GRGFIAGIDLKQQKREQSRFYGDLMEKRRTLEEKEQ 334
Query 60 QRGAEEAAAAAATAAAEAARASDKKKVKDMEEEPELEGHWSTKKREEMTDRDWRIFREDF 119
EEA R D+ HWS KK +EMTDRDWRIFRED+
Sbjct 335 ----EEARLRKLRKKEAKQRWDDR--------------HWSQKKLDEMTDRDWRIFREDY 376
Query 120 EIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
I KGG++P PIR+W ++ LP ++E + + Y+ PTPIQ QAIPI L+ RD+IG
Sbjct 377 SITTKGGKIPNPIRSWKDSSLPPHILEVIDKCGYKEPTPIQRQAIPIGLQNRDIIG 432
> pfa:PFE0925c snrnp protein, putative; K12858 ATP-dependent RNA
helicase DDX23/PRP28 [EC:3.6.4.13]
Length=1123
Score = 139 bits (350), Expect = 5e-33, Method: Compositional matrix adjust.
Identities = 61/81 (75%), Positives = 69/81 (85%), Gaps = 0/81 (0%)
Query 95 LEGHWSTKKREEMTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYE 154
E HWS K REEMTDRDWRIFRED EIY+KGG VPPPIR W E+ L +L++A+K+A YE
Sbjct 660 CEKHWSQKSREEMTDRDWRIFREDNEIYIKGGVVPPPIRKWEESNLSNDLLKAIKKAKYE 719
Query 155 RPTPIQMQAIPIALEMRDLIG 175
+PTPIQMQAIPIALEMRDLIG
Sbjct 720 KPTPIQMQAIPIALEMRDLIG 740
Score = 90.5 bits (223), Expect = 3e-18, Method: Compositional matrix adjust.
Identities = 40/57 (70%), Positives = 47/57 (82%), Gaps = 0/57 (0%)
Query 2 SEDTMRGDNNPLYQNRLEPQLLFGRGFRAGMDIREQRKANNFYDELVKRRQEYELKQ 58
SEDT R D+NPLYQNRLEPQLLFGRG+ AG+D+REQRK NNFYD+LV+ R L +
Sbjct 488 SEDTSRNDSNPLYQNRLEPQLLFGRGYIAGIDVREQRKKNNFYDKLVQNRLNMSLNK 544
> dre:334283 ddx23, wu:fi39b12, zgc:63742; DEAD (Asp-Glu-Ala-Asp)
box polypeptide 23 (EC:3.6.1.-); K12858 ATP-dependent RNA
helicase DDX23/PRP28 [EC:3.6.4.13]
Length=807
Score = 136 bits (342), Expect = 4e-32, Method: Compositional matrix adjust.
Identities = 76/176 (43%), Positives = 106/176 (60%), Gaps = 21/176 (11%)
Query 1 ASEDTMRGDNNPLYQNRLEPQLLFGRGFRAGMDIREQ-RKANNFYDELVKRRQEYELKQR 59
ASEDT D NP+Y+ + + L +GRGF AG+D+++Q R + FY +L+++R+ E K++
Sbjct 265 ASEDTSI-DYNPIYKEKHQVHL-YGRGFIAGIDLKQQKRDQSRFYGDLMEKRRTNEEKEQ 322
Query 60 QRGAEEAAAAAATAAAEAARASDKKKVKDMEEEPELEGHWSTKKREEMTDRDWRIFREDF 119
EE R D+ HWS KK +EMTDRDWRIFRED+
Sbjct 323 ----EEQRLKKVRKKEAKQRWDDR--------------HWSQKKLDEMTDRDWRIFREDY 364
Query 120 EIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
I KGG++P PIR W E LP ++E +++ Y+ PTPIQ QAIPI L+ RD+IG
Sbjct 365 SITTKGGKIPNPIRNWKEYSLPPHILEVIEKCGYKDPTPIQRQAIPIGLQNRDIIG 420
> cel:F01F1.7 ddx-23; DEAD boX helicase homolog family member
(ddx-23); K12858 ATP-dependent RNA helicase DDX23/PRP28 [EC:3.6.4.13]
Length=730
Score = 136 bits (342), Expect = 5e-32, Method: Compositional matrix adjust.
Identities = 79/176 (44%), Positives = 101/176 (57%), Gaps = 21/176 (11%)
Query 1 ASEDTMRGDNNPLYQNRLEPQLLFGRGFRAGMDIREQRK-ANNFYDELVKRRQEYELKQR 59
A EDT + D N LYQ+R E Q FGRG AG D+ Q+K N+FY E+++ R+ + K++
Sbjct 188 AGEDTSQ-DYNKLYQSRHEIQF-FGRGSVAGTDVNAQKKEKNSFYQEMMENRRTVDEKEQ 245
Query 60 QRGAEEAAAAAATAAAEAARASDKKKVKDMEEEPELEGHWSTKKREEMTDRDWRIFREDF 119
+ E A R HW K+ EM+DRDWRIFREDF
Sbjct 246 EMHRLEKELKKEKKVAHDDR------------------HWRMKELSEMSDRDWRIFREDF 287
Query 120 EIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
I +KGGRVP P+R W EA P E+ +AVK+ Y PTPIQ QAIPI L+ RD+IG
Sbjct 288 NISIKGGRVPRPLRNWEEAGFPDEVYQAVKEIGYLEPTPIQRQAIPIGLQNRDVIG 343
> ath:AT1G28180 ATP binding / ATP-dependent helicase/ helicase/
nucleic acid binding
Length=614
Score = 89.0 bits (219), Expect = 8e-18, Method: Compositional matrix adjust.
Identities = 59/174 (33%), Positives = 72/174 (41%), Gaps = 67/174 (38%)
Query 2 SEDTMRGDNNPLYQNRLEPQLLFGRGFRAGMDIREQRKANNFYDELVKRRQEYELKQRQR 61
+EDT+ G+ N LYQN E Q LFGRG RAG+D REQ+K
Sbjct 138 TEDTLSGEMNVLYQNPHEAQPLFGRGCRAGIDRREQKKL--------------------- 176
Query 62 GAEEAAAAAATAAAEAARASDKKKVKDMEEEPELEGHWSTKKREEMTDRDWRIFREDFEI 121
E DK HWS KK EEM +RDWRIF+EDF I
Sbjct 177 -------MTGKHEREKREEEDK--------------HWSEKKLEEMNERDWRIFKEDFNI 215
Query 122 YLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
+G ++P P+R W E IP+ LE RD+IG
Sbjct 216 SYRGSKIPHPMRNWEE-------------------------TIPLGLEQRDVIG 244
> cpv:cgd3_3690 U5 snRNP 100 kD protein ; K12858 ATP-dependent
RNA helicase DDX23/PRP28 [EC:3.6.4.13]
Length=529
Score = 86.3 bits (212), Expect = 5e-17, Method: Composition-based stats.
Identities = 36/70 (51%), Positives = 50/70 (71%), Gaps = 0/70 (0%)
Query 106 EMTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIP 165
+MT+RDW+IFRED+ I ++G VP PIR W + + E ++ YE+PTPIQMQ IP
Sbjct 115 DMTERDWKIFREDYSINVRGKDVPNPIRNWKDCHVLEIQTELIRNIGYEKPTPIQMQCIP 174
Query 166 IALEMRDLIG 175
I L++RD+IG
Sbjct 175 IGLKLRDMIG 184
> sce:YDR243C PRP28; Prp28p (EC:3.6.1.-); K12858 ATP-dependent
RNA helicase DDX23/PRP28 [EC:3.6.4.13]
Length=588
Score = 70.5 bits (171), Expect = 3e-12, Method: Compositional matrix adjust.
Identities = 37/94 (39%), Positives = 50/94 (53%), Gaps = 5/94 (5%)
Query 87 KDMEEEPELEGHWSTKKREEMTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAE-LPWELI 145
K+ E + HW+ K EM +RDWRI +ED+ I KGG V P+R W E +P +L+
Sbjct 126 KNAAESSYMGKHWTEKSLHEMNERDWRILKEDYAIVTKGGTVENPLRNWEELNIIPRDLL 185
Query 146 EAVKQ-ANYERPTPIQMQAIPIALEM---RDLIG 175
+ Q + PTPIQ IP M RD +G
Sbjct 186 RVIIQELRFPSPTPIQRITIPNVCNMKQYRDFLG 219
> mmu:212880 Ddx46, 2200005K02Rik, 8430438J23Rik, AI325430, AI957095,
MGC116676, MGC31579, mKIAA0801; DEAD (Asp-Glu-Ala-Asp)
box polypeptide 46 (EC:3.6.4.13); K12811 ATP-dependent RNA
helicase DDX46/PRP5 [EC:3.6.4.13]
Length=1031
Score = 66.6 bits (161), Expect = 4e-11, Method: Compositional matrix adjust.
Identities = 28/71 (39%), Positives = 46/71 (64%), Gaps = 1/71 (1%)
Query 106 EMTDRDWRIFREDFE-IYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAI 164
+M+ + +FR + E I +KG P PI++W + + +++ ++K+ YE+PTPIQ QAI
Sbjct 344 KMSQEEVNVFRLEMEGITVKGKGCPKPIKSWVQCGISMKILNSLKKHGYEKPTPIQTQAI 403
Query 165 PIALEMRDLIG 175
P + RDLIG
Sbjct 404 PAIMSGRDLIG 414
> hsa:9879 DDX46, FLJ25329, KIAA0801, MGC9936, PRPF5, Prp5; DEAD
(Asp-Glu-Ala-Asp) box polypeptide 46 (EC:3.6.4.13); K12811
ATP-dependent RNA helicase DDX46/PRP5 [EC:3.6.4.13]
Length=1031
Score = 66.6 bits (161), Expect = 4e-11, Method: Compositional matrix adjust.
Identities = 28/71 (39%), Positives = 46/71 (64%), Gaps = 1/71 (1%)
Query 106 EMTDRDWRIFREDFE-IYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAI 164
+M+ + +FR + E I +KG P PI++W + + +++ ++K+ YE+PTPIQ QAI
Sbjct 344 KMSQEEVNVFRLEMEGITVKGKGCPKPIKSWVQCGISMKILNSLKKHGYEKPTPIQTQAI 403
Query 165 PIALEMRDLIG 175
P + RDLIG
Sbjct 404 PAIMSGRDLIG 414
> dre:321948 ddx46, fb39a03, wu:fb39a03; DEAD (Asp-Glu-Ala-Asp)
box polypeptide 46 (EC:3.6.4.13); K12811 ATP-dependent RNA
helicase DDX46/PRP5 [EC:3.6.4.13]
Length=1018
Score = 66.2 bits (160), Expect = 5e-11, Method: Compositional matrix adjust.
Identities = 29/62 (46%), Positives = 42/62 (67%), Gaps = 1/62 (1%)
Query 115 FREDFE-IYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDL 173
+R + E I +KG P PI+TW + + +++ A+K+ NYE+PTPIQ QAIP + RDL
Sbjct 321 YRLELEGISVKGKGCPKPIKTWVQCGISMKVLNALKKHNYEKPTPIQAQAIPAIMSGRDL 380
Query 174 IG 175
IG
Sbjct 381 IG 382
> ath:AT1G20920 DEAD box RNA helicase, putative; K12811 ATP-dependent
RNA helicase DDX46/PRP5 [EC:3.6.4.13]
Length=828
Score = 65.1 bits (157), Expect = 1e-10, Method: Compositional matrix adjust.
Identities = 28/78 (35%), Positives = 46/78 (58%), Gaps = 0/78 (0%)
Query 98 HWSTKKREEMTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPT 157
+ K MT + +R++ E+ + G VP PI+ W + L ++++ +K+ NYE+P
Sbjct 156 YIEVKDISRMTQEEVNTYRKELELKVHGKDVPRPIKFWHQTGLTSKILDTMKKLNYEKPM 215
Query 158 PIQMQAIPIALEMRDLIG 175
PIQ QA+PI + RD IG
Sbjct 216 PIQTQALPIIMSGRDCIG 233
> ath:AT3G09620 DEAD/DEAH box helicase, putative
Length=989
Score = 63.2 bits (152), Expect = 4e-10, Method: Compositional matrix adjust.
Identities = 27/69 (39%), Positives = 43/69 (62%), Gaps = 0/69 (0%)
Query 107 MTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPI 166
MT +R++ E+ + G VP PI+ W + L ++++ +K+ NYE+P PIQ QA+PI
Sbjct 370 MTQDAVNAYRKELELKVHGKDVPRPIQFWHQTGLTSKILDTLKKLNYEKPMPIQAQALPI 429
Query 167 ALEMRDLIG 175
+ RD IG
Sbjct 430 IMSGRDCIG 438
> cel:F53H1.1 hypothetical protein; K12811 ATP-dependent RNA helicase
DDX46/PRP5 [EC:3.6.4.13]
Length=970
Score = 60.8 bits (146), Expect = 2e-09, Method: Compositional matrix adjust.
Identities = 27/70 (38%), Positives = 44/70 (62%), Gaps = 1/70 (1%)
Query 107 MTDRDWRIFREDFE-IYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIP 165
MT + + +RE+ + I +KG P PI+TWA+ + +++ +K+ Y +PT IQ QAIP
Sbjct 277 MTKAEVKAYREELDSITVKGIDCPKPIKTWAQCGVNLKMMNVLKKFEYSKPTSIQAQAIP 336
Query 166 IALEMRDLIG 175
+ RD+IG
Sbjct 337 SIMSGRDVIG 346
> xla:398649 ddx17, MGC80019; DEAD (Asp-Glu-Ala-Asp) box polypeptide
17; K13178 ATP-dependent RNA helicase DDX17 [EC:3.6.4.13]
Length=610
Score = 60.5 bits (145), Expect = 3e-09, Method: Compositional matrix adjust.
Identities = 31/95 (32%), Positives = 51/95 (53%), Gaps = 2/95 (2%)
Query 83 KKKVKDMEEEPELEGHWSTKKRE--EMTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAEL 140
+KK D+ E P+ E ++ T+ E MT D R EI ++G P P+ + +A
Sbjct 30 RKKRWDLNELPKFEKNFYTEHPEVARMTQHDVEELRRKKEITIRGVNCPKPLYAFHQANF 89
Query 141 PWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
P +++ + ++ PTPIQ Q P+AL RD++G
Sbjct 90 PQYVLDVLLDQRFKEPTPIQCQGFPLALSGRDMVG 124
> ath:AT3G02065 DEAD/DEAH box helicase family protein
Length=505
Score = 60.5 bits (145), Expect = 3e-09, Method: Compositional matrix adjust.
Identities = 27/69 (39%), Positives = 43/69 (62%), Gaps = 2/69 (2%)
Query 108 TDRDWRIFREDFEIYLKG--GRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIP 165
+ D ++ R +I+++G VPPP+ T+ LP +L+ ++ A Y+ PTPIQMQAIP
Sbjct 83 SSHDAQLLRRKLDIHVQGQGSAVPPPVLTFTSCGLPPKLLLNLETAGYDFPTPIQMQAIP 142
Query 166 IALEMRDLI 174
AL + L+
Sbjct 143 AALTGKSLL 151
> ath:AT5G51280 DEAD-box protein abstrakt, putative; K13116 ATP-dependent
RNA helicase DDX41 [EC:3.6.4.13]
Length=591
Score = 60.5 bits (145), Expect = 3e-09, Method: Compositional matrix adjust.
Identities = 31/103 (30%), Positives = 55/103 (53%), Gaps = 8/103 (7%)
Query 81 SDKKKVKDMEE--------EPELEGHWSTKKREEMTDRDWRIFREDFEIYLKGGRVPPPI 132
SDKK + + E EP L G +M+ + + R+ + I + G +PPPI
Sbjct 86 SDKKTLMSVRELAKGITYTEPLLTGWKPPLHIRKMSSKQRDLIRKQWHIIVNGDDIPPPI 145
Query 133 RTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
+ + + + P +++ +K+ +PTPIQ+Q +P+ L RD+IG
Sbjct 146 KNFKDMKFPRPVLDTLKEKGIVQPTPIQVQGLPVILAGRDMIG 188
> hsa:51428 DDX41, ABS, MGC8828; DEAD (Asp-Glu-Ala-Asp) box polypeptide
41 (EC:3.6.4.13); K13116 ATP-dependent RNA helicase
DDX41 [EC:3.6.4.13]
Length=622
Score = 60.1 bits (144), Expect = 4e-09, Method: Compositional matrix adjust.
Identities = 23/61 (37%), Positives = 38/61 (62%), Gaps = 0/61 (0%)
Query 115 FREDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLI 174
R+ + I ++G +PPPI+++ E + P ++ +K+ PTPIQ+Q IP L RD+I
Sbjct 163 VRKKYHILVEGDGIPPPIKSFKEMKFPAAILRGLKKKGIHHPTPIQIQGIPTILSGRDMI 222
Query 175 G 175
G
Sbjct 223 G 223
> dre:556764 similar to Probable RNA-dependent helicase p72 (DEAD-box
protein p72) (DEAD-box protein 17); K13178 ATP-dependent
RNA helicase DDX17 [EC:3.6.4.13]
Length=671
Score = 59.7 bits (143), Expect = 4e-09, Method: Compositional matrix adjust.
Identities = 29/95 (30%), Positives = 54/95 (56%), Gaps = 2/95 (2%)
Query 83 KKKVKDMEEEPELEGHWSTKKRE--EMTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAEL 140
+KK D+++ P+ E ++ + E M+ D +R EI ++G P P+ + +A+
Sbjct 43 RKKKWDLDQLPKFEKNFYNENPEVHHMSQYDVEEYRRKREITVRGSGCPKPVTNFHQAQF 102
Query 141 PWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
P +++ + Q N++ PT IQ Q P+AL RD++G
Sbjct 103 PQYVMDVLLQQNFKEPTAIQAQGFPLALSGRDMVG 137
> ath:AT5G63120 ethylene-responsive DEAD box RNA helicase, putative
(RH30)
Length=484
Score = 59.7 bits (143), Expect = 5e-09, Method: Compositional matrix adjust.
Identities = 27/71 (38%), Positives = 47/71 (66%), Gaps = 0/71 (0%)
Query 105 EEMTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAI 164
+ MT++D ++R + +I ++G VP P++ + +A P ++EA+ + + PTPIQ Q
Sbjct 137 QAMTEQDVAMYRTERDISVEGRDVPKPMKMFQDANFPDNILEAIAKLGFTEPTPIQAQGW 196
Query 165 PIALEMRDLIG 175
P+AL+ RDLIG
Sbjct 197 PMALKGRDLIG 207
> mmu:72935 Ddx41, 2900024F02Rik, AA958953, ABS, AI324246; DEAD
(Asp-Glu-Ala-Asp) box polypeptide 41 (EC:3.6.4.13); K13116
ATP-dependent RNA helicase DDX41 [EC:3.6.4.13]
Length=622
Score = 58.9 bits (141), Expect = 8e-09, Method: Compositional matrix adjust.
Identities = 23/61 (37%), Positives = 38/61 (62%), Gaps = 0/61 (0%)
Query 115 FREDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLI 174
R+ + I ++G +PPPI+++ E + P ++ +K+ PTPIQ+Q IP L RD+I
Sbjct 163 VRKKYHILVEGDGIPPPIKSFKEMKFPAAILRGLKKKGILHPTPIQIQGIPTILSGRDMI 222
Query 175 G 175
G
Sbjct 223 G 223
> cpv:cgd7_4600 abstrakt protein SF II helicase + Znknuckle C2HC
(PA) ; K13116 ATP-dependent RNA helicase DDX41 [EC:3.6.4.13]
Length=570
Score = 58.9 bits (141), Expect = 9e-09, Method: Compositional matrix adjust.
Identities = 34/113 (30%), Positives = 53/113 (46%), Gaps = 1/113 (0%)
Query 64 EEAAAAAATAAAEAARASDKKKVKDMEEEPELEGHWST-KKREEMTDRDWRIFREDFEIY 122
EE AA + A + + K + E W KK ++ + + R I
Sbjct 38 EEKLLAAVNQSYNAPLKAVHEIAKGITFSKREETSWRVPKKYSSLSASECQDLRSRLLIV 97
Query 123 LKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
+ G VPPPI ++ + P E+++A+ +P+ IQMQ +PI L RDLIG
Sbjct 98 VNGSDVPPPILSFKDMGFPQEILDALASKGISKPSQIQMQGLPIILMGRDLIG 150
> cel:H27M09.1 hypothetical protein; K13116 ATP-dependent RNA
helicase DDX41 [EC:3.6.4.13]
Length=630
Score = 58.9 bits (141), Expect = 9e-09, Method: Compositional matrix adjust.
Identities = 31/80 (38%), Positives = 45/80 (56%), Gaps = 6/80 (7%)
Query 97 GHWSTKKREEMTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAELPWELIEAV-KQANYER 155
GH + +E D+ I R+ I +G +PPPI ++ E + P L+E + KQ
Sbjct 158 GHIRRQSQE-----DYEIQRKRLGISCEGDHIPPPIGSFLEMKFPKSLLEFMQKQKGIVT 212
Query 156 PTPIQMQAIPIALEMRDLIG 175
PT IQ+Q IP+AL RD+IG
Sbjct 213 PTAIQIQGIPVALSGRDMIG 232
> xla:399382 ddx5, MGC81559; DEAD (Asp-Glu-Ala-Asp) box polypeptide
5; K12823 ATP-dependent RNA helicase DDX5/DBP2 [EC:3.6.4.13]
Length=608
Score = 58.5 bits (140), Expect = 1e-08, Method: Compositional matrix adjust.
Identities = 31/94 (32%), Positives = 54/94 (57%), Gaps = 2/94 (2%)
Query 84 KKVKDMEEEPELEGHWSTKKREEM--TDRDWRIFREDFEIYLKGGRVPPPIRTWAEAELP 141
KK +++E P+ E ++ + + + T ++ +R EI ++G P PI + EA P
Sbjct 41 KKKWNLDELPKFEKNFYQEHPDVVRRTPQECDQYRRSKEITVRGINCPKPILNFNEASFP 100
Query 142 WELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
++EA+K+ N+ PTPIQ Q P+AL D++G
Sbjct 101 ANVMEAIKRQNFTEPTPIQGQGWPVALSGLDMVG 134
> ath:AT4G33370 DEAD-box protein abstrakt, putative
Length=542
Score = 57.8 bits (138), Expect = 2e-08, Method: Composition-based stats.
Identities = 33/104 (31%), Positives = 53/104 (50%), Gaps = 10/104 (9%)
Query 81 SDKKKVKDMEE--------EPELEGHWSTKKR-EEMTDRDWRIFREDFEIYLKGGRVPPP 131
SDKKK+ + E EP L W +M+ + + R+ + I + G +PPP
Sbjct 37 SDKKKLMSVGELARGITYTEP-LSTWWKPPLHVRKMSTKQMDLIRKQWHITVNGEDIPPP 95
Query 132 IRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
I+ + + + P L+ +K PTPIQ+Q +P+ L RD+IG
Sbjct 96 IKNFMDMKFPSPLLRMLKDKGIMHPTPIQVQGLPVVLSGRDMIG 139
> ath:AT2G42520 DEAD box RNA helicase, putative; K11594 ATP-dependent
RNA helicase [EC:3.6.4.13]
Length=633
Score = 57.4 bits (137), Expect = 3e-08, Method: Compositional matrix adjust.
Identities = 27/58 (46%), Positives = 36/58 (62%), Gaps = 0/58 (0%)
Query 117 EDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLI 174
ED I G VPPP+ T+AE +L L +++ Y +PTP+Q AIPI LE RDL+
Sbjct 143 EDIPIETSGDNVPPPVNTFAEIDLGEALNLNIRRCKYVKPTPVQRHAIPILLEGRDLM 200
> ath:AT1G55150 DEAD box RNA helicase, putative (RH20); K12823
ATP-dependent RNA helicase DDX5/DBP2 [EC:3.6.4.13]
Length=501
Score = 57.0 bits (136), Expect = 3e-08, Method: Compositional matrix adjust.
Identities = 27/69 (39%), Positives = 45/69 (65%), Gaps = 0/69 (0%)
Query 107 MTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPI 166
MTD + +R+ EI ++G +P P++++ + P ++E VK+A + PTPIQ Q P+
Sbjct 73 MTDTEVEEYRKLREITVEGKDIPKPVKSFRDVGFPDYVLEEVKKAGFTEPTPIQSQGWPM 132
Query 167 ALEMRDLIG 175
A++ RDLIG
Sbjct 133 AMKGRDLIG 141
> pfa:PFE1390w RNA helicase-1; K13116 ATP-dependent RNA helicase
DDX41 [EC:3.6.4.13]
Length=665
Score = 57.0 bits (136), Expect = 3e-08, Method: Composition-based stats.
Identities = 36/130 (27%), Positives = 66/130 (50%), Gaps = 1/130 (0%)
Query 47 LVKRRQEYELKQRQRGAEEAAAAAATAAAEAARASDKKKVKDMEEEPELEGHWSTKKREE 106
L K+ E + + R EE A + A A S K++ K + + +E W K+ +
Sbjct 130 LEKKNDEIDETEEIRKREEKLLAQVSKALNAPLQSVKERAKGIVYKENVESIWKLPKKYK 189
Query 107 MTDRDW-RIFREDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIP 165
+ + + R F I + G +P PI+ + + + P +++ +++ N ++PT IQMQ +P
Sbjct 190 LLKKSYVEKIRRIFYIDVNGDDIPAPIKNFKDMKFPKAILKGLRKKNIKKPTQIQMQGLP 249
Query 166 IALEMRDLIG 175
L RD+IG
Sbjct 250 SILLGRDIIG 259
> ath:AT3G58510 DEAD box RNA helicase, putative (RH11); K11594
ATP-dependent RNA helicase [EC:3.6.4.13]
Length=612
Score = 56.6 bits (135), Expect = 4e-08, Method: Compositional matrix adjust.
Identities = 26/59 (44%), Positives = 36/59 (61%), Gaps = 0/59 (0%)
Query 117 EDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
ED + GG VPPP+ T+A+ +L L +++ Y RPTP+Q AIPI L RDL+
Sbjct 135 EDIPVETSGGDVPPPVNTFADIDLGDALNLNIRRCKYVRPTPVQRHAIPILLAERDLMA 193
> ath:AT2G47330 DEAD/DEAH box helicase, putative; K12835 ATP-dependent
RNA helicase DDX42 [EC:3.6.4.13]
Length=760
Score = 56.6 bits (135), Expect = 5e-08, Method: Compositional matrix adjust.
Identities = 25/69 (36%), Positives = 43/69 (62%), Gaps = 0/69 (0%)
Query 107 MTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPI 166
MT+++ +R+ I + G V P++T+ + +++ A+K+ YE+PT IQ QA+PI
Sbjct 202 MTEQETTDYRQRLGIRVSGFDVHRPVKTFEDCGFSSQIMSAIKKQAYEKPTAIQCQALPI 261
Query 167 ALEMRDLIG 175
L RD+IG
Sbjct 262 VLSGRDVIG 270
> ath:AT3G01540 DRH1; DRH1 (DEAD BOX RNA HELICASE 1); ATP-dependent
RNA helicase/ ATPase
Length=619
Score = 56.2 bits (134), Expect = 5e-08, Method: Compositional matrix adjust.
Identities = 24/55 (43%), Positives = 36/55 (65%), Gaps = 0/55 (0%)
Query 120 EIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLI 174
EI + GG+VPPP+ ++ P EL+ V A + PTPIQ Q+ PIA++ RD++
Sbjct 145 EITVSGGQVPPPLMSFEATGFPPELLREVLSAGFSAPTPIQAQSWPIAMQGRDIV 199
> xla:379390 MGC53795; similar to DEAD/H (Asp-Glu-Ala-Asp/His)
box polypeptide 5 (RNA helicase, 68kDa); K12823 ATP-dependent
RNA helicase DDX5/DBP2 [EC:3.6.4.13]
Length=607
Score = 56.2 bits (134), Expect = 6e-08, Method: Compositional matrix adjust.
Identities = 31/98 (31%), Positives = 52/98 (53%), Gaps = 10/98 (10%)
Query 84 KKVKDMEEEPELEGHWSTKKREEMTDRDWRI------FREDFEIYLKGGRVPPPIRTWAE 137
KK +++E P+ E ++ +E+ D R +R EI ++G P P+ + E
Sbjct 39 KKKWNLDELPKFEKNFY----QELPDVSRRTPQECDQYRRSKEITVRGLNCPKPVLNFHE 94
Query 138 AELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
A P ++E +K+ N+ PTPIQ Q P+AL D++G
Sbjct 95 ASFPANVMEVIKRLNFTEPTPIQGQGWPVALSGLDMVG 132
> dre:394020 ddx41, MGC55896, wu:fb92e02, zgc:55896; DEAD (Asp-Glu-Ala-Asp)
box polypeptide 41 (EC:3.6.4.13); K13116 ATP-dependent
RNA helicase DDX41 [EC:3.6.4.13]
Length=306
Score = 55.1 bits (131), Expect = 1e-07, Method: Compositional matrix adjust.
Identities = 22/60 (36%), Positives = 38/60 (63%), Gaps = 0/60 (0%)
Query 116 REDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
R+ + I ++G +P PI+++ E + P +++ +K+ PTPIQ+Q IP L RD+IG
Sbjct 155 RKKYHILVEGEGIPAPIKSFREMKFPQAILKGLKKKGIVHPTPIQIQGIPTILSGRDMIG 214
> hsa:11056 DDX52, HUSSY19, ROK1; DEAD (Asp-Glu-Ala-Asp) box polypeptide
52 (EC:3.6.4.13); K14779 ATP-dependent RNA helicase
DDX52/ROK1 [EC:3.6.4.13]
Length=599
Score = 54.3 bits (129), Expect = 2e-07, Method: Compositional matrix adjust.
Identities = 37/134 (27%), Positives = 71/134 (52%), Gaps = 7/134 (5%)
Query 45 DELVKRRQEYELKQRQRGAEEAAAAAATAAAEAARASDKKKVKDMEEEPELEGHWSTKKR 104
+ L +R++E K+R+ E A+ A + +S + K++D ++ + E ++ K
Sbjct 76 ESLTERKREQSKKKRKTMTSEIASQEEGATIQWM-SSVEAKIED--KKVQRESKLTSGKL 132
Query 105 EEMTDRDWRIFREDFEIYLKGGRVPPPIRTWA----EAELPWELIEAVKQANYERPTPIQ 160
E + R +I+++G +P PI T+ E ++ L++ + A ++ PTPIQ
Sbjct 133 ENLRKEKINFLRNKHKIHVQGTDLPDPIATFQQLDQEYKINSRLLQNILDAGFQMPTPIQ 192
Query 161 MQAIPIALEMRDLI 174
MQAIP+ L R+L+
Sbjct 193 MQAIPVMLHGRELL 206
> sce:YNL112W DBP2; Dbp2p (EC:3.6.1.-); K12823 ATP-dependent RNA
helicase DDX5/DBP2 [EC:3.6.4.13]
Length=546
Score = 54.3 bits (129), Expect = 2e-07, Method: Compositional matrix adjust.
Identities = 28/78 (35%), Positives = 45/78 (57%), Gaps = 3/78 (3%)
Query 98 HWSTKKREEMTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPT 157
H S + R +D + FR++ E+ + G +P PI T+ EA P ++ VK +++PT
Sbjct 81 HESVRDR---SDSEIAQFRKENEMTISGHDIPKPITTFDEAGFPDYVLNEVKAEGFDKPT 137
Query 158 PIQMQAIPIALEMRDLIG 175
IQ Q P+AL RD++G
Sbjct 138 GIQCQGWPMALSGRDMVG 155
> mmu:67040 Ddx17, 2610007K22Rik, A430025E01Rik, AI047725, C80929,
Gm926, MGC79147, p72; DEAD (Asp-Glu-Ala-Asp) box polypeptide
17 (EC:3.6.4.13); K13178 ATP-dependent RNA helicase DDX17
[EC:3.6.4.13]
Length=650
Score = 54.3 bits (129), Expect = 2e-07, Method: Compositional matrix adjust.
Identities = 30/96 (31%), Positives = 51/96 (53%), Gaps = 3/96 (3%)
Query 83 KKKVKDMEEEPELEGHWSTKKRE--EMTDRDWRIFREDFEIYLKGGRVPP-PIRTWAEAE 139
+KK D+ E P+ E ++ + E +T + R EI ++GG V P P+ + A
Sbjct 39 RKKKWDLSELPKFEKNFYVEHPEVARLTPYEVDELRRKKEITVRGGDVCPKPVFAFHHAN 98
Query 140 LPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
P +++ + ++ PTPIQ Q P+AL RD++G
Sbjct 99 FPQYVMDVLMDQHFTEPTPIQCQGFPLALSGRDMVG 134
> hsa:10521 DDX17, DKFZp761H2016, P72, RH70; DEAD (Asp-Glu-Ala-Asp)
box polypeptide 17 (EC:3.6.4.13); K13178 ATP-dependent
RNA helicase DDX17 [EC:3.6.4.13]
Length=731
Score = 53.9 bits (128), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 30/96 (31%), Positives = 51/96 (53%), Gaps = 3/96 (3%)
Query 83 KKKVKDMEEEPELEGHWSTKKRE--EMTDRDWRIFREDFEIYLKGGRVPP-PIRTWAEAE 139
+KK D+ E P+ E ++ + E +T + R EI ++GG V P P+ + A
Sbjct 118 RKKKWDLSELPKFEKNFYVEHPEVARLTPYEVDELRRKKEITVRGGDVCPKPVFAFHHAN 177
Query 140 LPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
P +++ + ++ PTPIQ Q P+AL RD++G
Sbjct 178 FPQYVMDVLMDQHFTEPTPIQCQGFPLALSGRDMVG 213
> bbo:BBOV_II004470 18.m06373; p68-like protein; K12823 ATP-dependent
RNA helicase DDX5/DBP2 [EC:3.6.4.13]
Length=529
Score = 53.9 bits (128), Expect = 3e-07, Method: Compositional matrix adjust.
Identities = 27/70 (38%), Positives = 42/70 (60%), Gaps = 1/70 (1%)
Query 107 MTDRDWRIFREDFEIYLKGGR-VPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIP 165
M+ D R++ EI + GR VP P+ ++ P +++A++ A + PTPIQ+Q P
Sbjct 81 MSSADVDRVRKEREITIIAGRDVPKPVVSFEHTSFPDYILKAIRAAGFTAPTPIQVQGWP 140
Query 166 IALEMRDLIG 175
IAL RD+IG
Sbjct 141 IALSGRDVIG 150
> xla:446276 ddx52; DEAD (Asp-Glu-Ala-Asp) box polypeptide 52;
K14779 ATP-dependent RNA helicase DDX52/ROK1 [EC:3.6.4.13]
Length=614
Score = 53.5 bits (127), Expect = 4e-07, Method: Composition-based stats.
Identities = 35/131 (26%), Positives = 68/131 (51%), Gaps = 4/131 (3%)
Query 49 KRRQEYELKQRQRGAEEAAAAAATAAAEAARASDKKKVKDMEEEPELEGHWSTKKREEMT 108
KR+++ ++R +E+ A +A E+ + K + + + + S +K ++
Sbjct 83 KRKKDLGRGVKKRKIQESDAGDVSAGDESILWMSSLEAKLQDGKGKSKKKLSIEKMKQQR 142
Query 109 DRDWRIFREDFEIYLKGGRVPPPIRTWAEAELPWELI----EAVKQANYERPTPIQMQAI 164
FR + +IY++G +P P T+ + E ++++ + VK A + PTPIQMQAI
Sbjct 143 LEKINHFRNEHKIYVQGTDIPEPAATFQQLEQEYKILSKIMQNVKDAGFHTPTPIQMQAI 202
Query 165 PIALEMRDLIG 175
PI L R+++
Sbjct 203 PIMLHDREILA 213
> mmu:100048658 Ddx43, OTTMUSG00000019690; DEAD (Asp-Glu-Ala-Asp)
box polypeptide 43 (EC:3.6.4.13)
Length=646
Score = 53.5 bits (127), Expect = 4e-07, Method: Compositional matrix adjust.
Identities = 51/168 (30%), Positives = 82/168 (48%), Gaps = 34/168 (20%)
Query 37 QRKANNFYDELVKRRQEYELKQR--------QRGAEEAAAAAAT---------AAAEAAR 79
Q KA D +VK++Q Y QR GA+ + + + T E A
Sbjct 120 QTKAKTVIDNVVKKQQNYIPGQRVGIIAFQPTVGADTSTSESVTDDQPLIDWDQIREDAL 179
Query 80 ASDKKKVKDMEEEPELEGHW-----STKKREEMTDRDWRIFREDFEIY---LKGGR---V 128
+KKK D+ P ++ ++ +T ++ +WR +E+F I LK G +
Sbjct 180 KWEKKKWADL---PPIKKNFYIESATTSSMSQVQIDNWR--KENFNITCDDLKDGEKRPI 234
Query 129 PPPIRTWAEAELPW-ELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
P PI + +A + E++E +K+A +++PTPIQ QA PI L+ DLIG
Sbjct 235 PNPICKFEDAFQSYPEVMENIKRAGFQKPTPIQSQAWPIVLQGIDLIG 282
> tpv:TP02_0316 RNA helicase-1; K13116 ATP-dependent RNA helicase
DDX41 [EC:3.6.4.13]
Length=598
Score = 53.1 bits (126), Expect = 5e-07, Method: Composition-based stats.
Identities = 25/60 (41%), Positives = 37/60 (61%), Gaps = 0/60 (0%)
Query 116 REDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
R I + G +VPPPI T+ + +LP +++A++ PT IQMQA+P L RD+IG
Sbjct 175 RNALVIDVSGDQVPPPILTFEDMKLPRPILKALRHKKIFEPTKIQMQAMPAVLLGRDVIG 234
> ath:AT3G58570 DEAD box RNA helicase, putative; K11594 ATP-dependent
RNA helicase [EC:3.6.4.13]
Length=646
Score = 52.4 bits (124), Expect = 7e-07, Method: Compositional matrix adjust.
Identities = 25/58 (43%), Positives = 34/58 (58%), Gaps = 0/58 (0%)
Query 117 EDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLI 174
ED I G VPPP+ T+AE +L L +++ Y +PTP+Q AIPI RDL+
Sbjct 130 EDIPIETSGDNVPPPVNTFAEIDLGEALNLNIQRCKYVKPTPVQRNAIPILAAGRDLM 187
> tgo:TGME49_010360 DEAD/DEAH box helicase, putative (EC:5.99.1.3);
K13116 ATP-dependent RNA helicase DDX41 [EC:3.6.4.13]
Length=657
Score = 52.0 bits (123), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 27/73 (36%), Positives = 39/73 (53%), Gaps = 0/73 (0%)
Query 103 KREEMTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQ 162
K EMT + RE F I + G PPP R + + P +++ +++ PT IQMQ
Sbjct 178 KYAEMTLAEANEVRERFFIDVSGEDPPPPFRNFKDMRFPQPILKGLQERGISYPTQIQMQ 237
Query 163 AIPIALEMRDLIG 175
IP L+ RD+IG
Sbjct 238 GIPAILQGRDIIG 250
> xla:379217 MGC53409; similar to ATP-dependent, RNA helicase;
K14779 ATP-dependent RNA helicase DDX52/ROK1 [EC:3.6.4.13]
Length=686
Score = 51.6 bits (122), Expect = 1e-06, Method: Composition-based stats.
Identities = 25/65 (38%), Positives = 41/65 (63%), Gaps = 4/65 (6%)
Query 115 FREDFEIYLKGGRVPPPIRTWAEAELPWEL----IEAVKQANYERPTPIQMQAIPIALEM 170
FR + +IY++G +P P T+ + E +++ ++ VK A + PTPIQMQAIPI L
Sbjct 149 FRNEQKIYIQGTDIPEPAATFQQLEQEYKIHSKIMQNVKDAGFHTPTPIQMQAIPIMLHG 208
Query 171 RDLIG 175
R+++
Sbjct 209 REILA 213
> cel:C46F11.4 hypothetical protein; K12835 ATP-dependent RNA
helicase DDX42 [EC:3.6.4.13]
Length=811
Score = 51.6 bits (122), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 24/52 (46%), Positives = 35/52 (67%), Gaps = 1/52 (1%)
Query 125 GGRVPP-PIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPIALEMRDLIG 175
GG PP P+ ++A L+EA++++ YE+PTPIQ AIP AL RD++G
Sbjct 256 GGLKPPRPVCSFAHFSFDKLLMEAIRKSEYEQPTPIQAMAIPSALSGRDVLG 307
> hsa:1655 DDX5, DKFZp434E109, DKFZp686J01190, G17P1, HLR1, HUMP68,
p68; DEAD (Asp-Glu-Ala-Asp) box polypeptide 5 (EC:3.6.4.13);
K12823 ATP-dependent RNA helicase DDX5/DBP2 [EC:3.6.4.13]
Length=614
Score = 51.6 bits (122), Expect = 1e-06, Method: Compositional matrix adjust.
Identities = 33/129 (25%), Positives = 59/129 (45%), Gaps = 12/129 (9%)
Query 59 RQRGAEEAAAAAATAAAEAARASDKK----------KVKDMEEEPELEGHWSTKKRE--E 106
R RG + A + A S KK K +++E P+ E ++ + +
Sbjct 8 RDRGRDRGFGAPRFGGSRAGPLSGKKFGNPGEKLVKKKWNLDELPKFEKNFYQEHPDLAR 67
Query 107 MTDRDWRIFREDFEIYLKGGRVPPPIRTWAEAELPWELIEAVKQANYERPTPIQMQAIPI 166
T ++ +R EI ++G P P+ + EA P +++ + + N+ PT IQ Q P+
Sbjct 68 RTAQEVETYRRSKEITVRGHNCPKPVLNFYEANFPANVMDVIARQNFTEPTAIQAQGWPV 127
Query 167 ALEMRDLIG 175
AL D++G
Sbjct 128 ALSGLDMVG 136
Lambda K H
0.315 0.132 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 4535951560
Database: egene_temp_file_orthology_annotation_similarity_blast_database_966
Posted date: Sep 16, 2011 8:45 PM
Number of letters in database: 82,071,388
Number of sequences in database: 164,496
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40