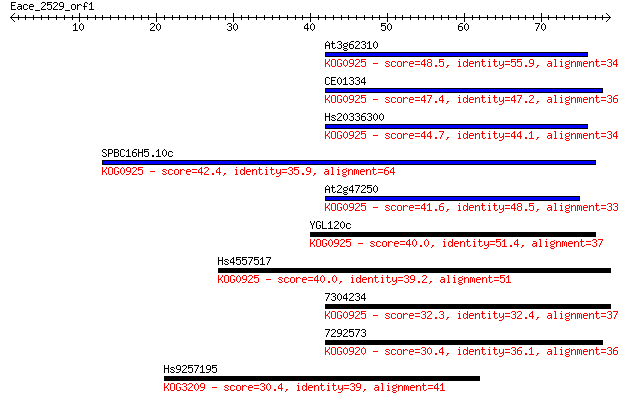
bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_2529_orf1
Length=78
Score E
Sequences producing significant alignments: (Bits) Value
At3g62310 48.5 3e-06
CE01334 47.4 7e-06
Hs20336300 44.7 4e-05
SPBC16H5.10c 42.4 2e-04
At2g47250 41.6 4e-04
YGL120c 40.0 0.001
Hs4557517 40.0 0.001
7304234 32.3 0.24
7292573 30.4 0.74
Hs9257195 30.4 0.85
> At3g62310
Length=726
Score = 48.5 bits (114), Expect = 3e-06, Method: Composition-based stats.
Identities = 19/34 (55%), Positives = 24/34 (70%), Gaps = 0/34 (0%)
Query 42 INPFNGRPFSDRYYRILEGRKKLPAWGSQRHFLK 75
IN +NG+P+S RYY ILE R+ LP W + FLK
Sbjct 39 INKWNGKPYSQRYYDILEKRRTLPVWLQKEEFLK 72
> CE01334
Length=739
Score = 47.4 bits (111), Expect = 7e-06, Method: Composition-based stats.
Identities = 17/36 (47%), Positives = 26/36 (72%), Gaps = 0/36 (0%)
Query 42 INPFNGRPFSDRYYRILEGRKKLPAWGSQRHFLKLV 77
INP+N +PFS+RY+ I E R +LP W + F++L+
Sbjct 54 INPYNNQPFSNRYWAIWEKRSQLPVWEYKEKFMELL 89
> Hs20336300
Length=743
Score = 44.7 bits (104), Expect = 4e-05, Method: Composition-based stats.
Identities = 15/34 (44%), Positives = 25/34 (73%), Gaps = 0/34 (0%)
Query 42 INPFNGRPFSDRYYRILEGRKKLPAWGSQRHFLK 75
+NPF+G P+S RYY++L+ R+ LP W + F++
Sbjct 40 LNPFDGLPYSSRYYKLLKEREDLPIWKEKYSFME 73
> SPBC16H5.10c
Length=735
Score = 42.4 bits (98), Expect = 2e-04, Method: Composition-based stats.
Identities = 23/64 (35%), Positives = 31/64 (48%), Gaps = 15/64 (23%)
Query 13 AAAAAAAAAAVNGPIGPTGPTSGPEPADGINPFNGRPFSDRYYRILEGRKKLPAWGSQRH 72
A A AA A GP N FN +PFS Y++ILE R++LP + +
Sbjct 39 ATTVAQAAKAEEGPN---------------NFFNDKPFSQNYFKILETRRELPVYQQREE 83
Query 73 FLKL 76
FLK+
Sbjct 84 FLKI 87
> At2g47250
Length=729
Score = 41.6 bits (96), Expect = 4e-04, Method: Composition-based stats.
Identities = 16/33 (48%), Positives = 22/33 (66%), Gaps = 0/33 (0%)
Query 42 INPFNGRPFSDRYYRILEGRKKLPAWGSQRHFL 74
IN +NG+ +S RY+ ILE R+ LP W + FL
Sbjct 43 INKWNGKAYSQRYFEILEKRRDLPVWLQKDDFL 75
> YGL120c
Length=767
Score = 40.0 bits (92), Expect = 0.001, Method: Composition-based stats.
Identities = 19/38 (50%), Positives = 25/38 (65%), Gaps = 1/38 (2%)
Query 40 DG-INPFNGRPFSDRYYRILEGRKKLPAWGSQRHFLKL 76
DG INPF GR F+ +Y IL+ R++LP + FLKL
Sbjct 68 DGKINPFTGREFTPKYVDILKIRRELPVHAQRDEFLKL 105
> Hs4557517
Length=813
Score = 40.0 bits (92), Expect = 0.001, Method: Composition-based stats.
Identities = 20/51 (39%), Positives = 27/51 (52%), Gaps = 3/51 (5%)
Query 28 GPTGPTSGPEPADGINPFNGRPFSDRYYRILEGRKKLPAWGSQRHFLKLVA 78
G G TS P+ INPF P + RYY IL+ R +LP W + F ++
Sbjct 104 GHAGHTSLPQ---CINPFTNLPHTPRYYDILKKRLQLPVWEYKDRFTDILG 151
> 7304234
Length=729
Score = 32.3 bits (72), Expect = 0.24, Method: Composition-based stats.
Identities = 12/37 (32%), Positives = 21/37 (56%), Gaps = 0/37 (0%)
Query 42 INPFNGRPFSDRYYRILEGRKKLPAWGSQRHFLKLVA 78
+NP P+S RY + + R LP + Q F++L++
Sbjct 50 MNPLTNTPYSQRYQNLYKKRIALPVFEYQADFMRLLS 86
> 7292573
Length=1280
Score = 30.4 bits (67), Expect = 0.74, Method: Composition-based stats.
Identities = 13/36 (36%), Positives = 21/36 (58%), Gaps = 0/36 (0%)
Query 42 INPFNGRPFSDRYYRILEGRKKLPAWGSQRHFLKLV 77
+ F R +RY +I++GRK+LPA+ L L+
Sbjct 423 LQQFVERRKEERYQKIIDGRKQLPAFAEIERILALI 458
> Hs9257195
Length=1256
Score = 30.4 bits (67), Expect = 0.85, Method: Composition-based stats.
Identities = 16/41 (39%), Positives = 23/41 (56%), Gaps = 1/41 (2%)
Query 21 AAVNGPIGPTGPTSGPEPADGINPFNGRPFSDRYYRILEGR 61
+A+ P+ P P S PEPA + P G+PF R L+G+
Sbjct 430 SALVPPVIPNHPPSNPEPAREV-PLQGKPFFTRNPSELKGK 469
Lambda K H
0.316 0.135 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1171925608
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40