bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_2095_orf2
Length=104
Score E
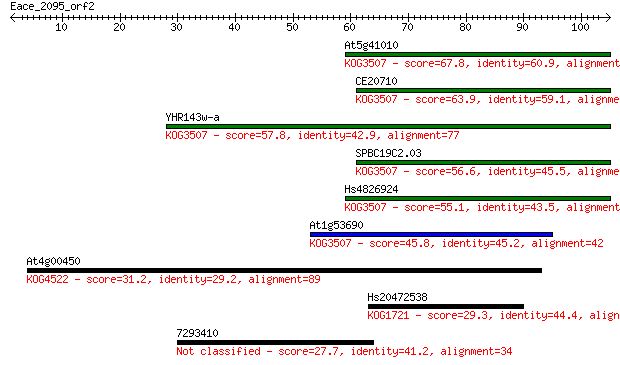
Sequences producing significant alignments: (Bits) Value
At5g41010 67.8 5e-12
CE20710 63.9 7e-11
YHR143w-a 57.8 5e-09
SPBC19C2.03 56.6 1e-08
Hs4826924 55.1 3e-08
At1g53690 45.8 2e-05
At4g00450 31.2 0.46
Hs20472538 29.3 2.1
7293410 27.7 4.7
> At5g41010
Length=51
Score = 67.8 bits (164), Expect = 5e-12, Method: Compositional matrix adjust.
Identities = 28/46 (60%), Positives = 38/46 (82%), Gaps = 0/46 (0%)
Query 59 EPVTYICGECGCDVTLQPTAAVRCRSCGSRILYKKRSRKMMQYEAR 104
EPVTY+CG+CG + TL+ ++CR CG RILYKKR+R+++QYEAR
Sbjct 6 EPVTYVCGDCGQENTLKSGDVIQCRECGYRILYKKRTRRVVQYEAR 51
> CE20710
Length=62
Score = 63.9 bits (154), Expect = 7e-11, Method: Compositional matrix adjust.
Identities = 26/44 (59%), Positives = 33/44 (75%), Gaps = 0/44 (0%)
Query 61 VTYICGECGCDVTLQPTAAVRCRSCGSRILYKKRSRKMMQYEAR 104
+ YICGEC + ++P A+RCR CG RILYKKR RK+M Y+AR
Sbjct 19 MIYICGECHAENEIKPKDAIRCRECGYRILYKKRCRKLMVYDAR 62
> YHR143w-a
Length=70
Score = 57.8 bits (138), Expect = 5e-09, Method: Compositional matrix adjust.
Identities = 33/77 (42%), Positives = 43/77 (55%), Gaps = 15/77 (19%)
Query 28 PTTAGATAAAGTTTAAAAAAKMEIEDNNTQLEPVTYICGECGCDVTLQPTAAVRCRSCGS 87
PT A AAAGT+ A A K YIC EC ++L T AVRC+ CG
Sbjct 9 PTNLDA-AAAGTSQARTATLK--------------YICAECSSKLSLSRTDAVRCKDCGH 53
Query 88 RILYKKRSRKMMQYEAR 104
RIL K R+++++Q+EAR
Sbjct 54 RILLKARTKRLVQFEAR 70
> SPBC19C2.03
Length=63
Score = 56.6 bits (135), Expect = 1e-08, Method: Compositional matrix adjust.
Identities = 20/44 (45%), Positives = 32/44 (72%), Gaps = 0/44 (0%)
Query 61 VTYICGECGCDVTLQPTAAVRCRSCGSRILYKKRSRKMMQYEAR 104
+ Y+C +CG T+Q +RCR CG R++YK R+++M+Q+EAR
Sbjct 20 MIYLCADCGARNTIQAKEVIRCRECGHRVMYKMRTKRMVQFEAR 63
> Hs4826924
Length=58
Score = 55.1 bits (131), Expect = 3e-08, Method: Compositional matrix adjust.
Identities = 20/46 (43%), Positives = 34/46 (73%), Gaps = 0/46 (0%)
Query 59 EPVTYICGECGCDVTLQPTAAVRCRSCGSRILYKKRSRKMMQYEAR 104
+P+ YICGEC + ++ +RCR CG RI+YKKR+++++ ++AR
Sbjct 13 QPMIYICGECHTENEIKSRDPIRCRECGYRIMYKKRTKRLVVFDAR 58
> At1g53690
Length=61
Score = 45.8 bits (107), Expect = 2e-05, Method: Compositional matrix adjust.
Identities = 19/42 (45%), Positives = 26/42 (61%), Gaps = 0/42 (0%)
Query 53 DNNTQLEPVTYICGECGCDVTLQPTAAVRCRSCGSRILYKKR 94
D+ + V Y+CG+CG + L+ +CR CG RILYKKR
Sbjct 9 DDKQPEQLVIYVCGDCGQENILKRGDVFQCRDCGFRILYKKR 50
> At4g00450
Length=2124
Score = 31.2 bits (69), Expect = 0.46, Method: Composition-based stats.
Identities = 26/95 (27%), Positives = 43/95 (45%), Gaps = 8/95 (8%)
Query 4 GLSTSAATVATATAAARGSAAAGTPTTAGATAAAGTTT------AAAAAAKMEIEDNNTQ 57
G ++S + +++ + R G+P+ A ++A T T AA A+ +
Sbjct 1823 GTASSGSEISSNKGSTRKGLRGGSPSLARRSSANTTDTSPPPSPAALRASMSLRLQFLLR 1882
Query 58 LEPVTYICGECGCDVTLQPTAAVRCRSCGSRILYK 92
L PV ICGE T A+ R GSR++Y+
Sbjct 1883 LLPV--ICGEPSFKNTRHALASTIVRLLGSRVVYE 1915
> Hs20472538
Length=273
Score = 29.3 bits (64), Expect = 2.1, Method: Compositional matrix adjust.
Identities = 12/27 (44%), Positives = 17/27 (62%), Gaps = 0/27 (0%)
Query 63 YICGECGCDVTLQPTAAVRCRSCGSRI 89
Y+CG+CG T + + AV RSC R+
Sbjct 245 YLCGQCGKSFTQRGSLAVHQRSCSQRL 271
> 7293410
Length=531
Score = 27.7 bits (60), Expect = 4.7, Method: Compositional matrix adjust.
Identities = 14/34 (41%), Positives = 22/34 (64%), Gaps = 0/34 (0%)
Query 30 TAGATAAAGTTTAAAAAAKMEIEDNNTQLEPVTY 63
+AG+TAA G+TTA A A+ +N+ + P +Y
Sbjct 131 SAGSTAAGGSTTAGATASSTATNNNHIYVNPHSY 164
Lambda K H
0.312 0.119 0.335
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1174332180
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40