bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_0290_orf1
Length=315
Score E
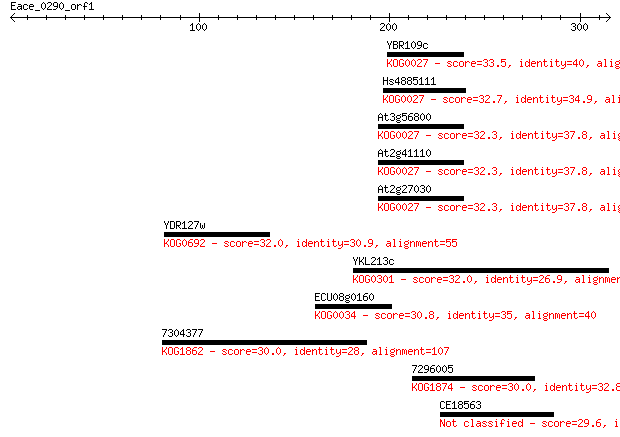
Sequences producing significant alignments: (Bits) Value
YBR109c 33.5 0.57
Hs4885111 32.7 0.94
At3g56800 32.3 1.4
At2g41110 32.3 1.4
At2g27030 32.3 1.4
YDR127w 32.0 1.8
YKL213c 32.0 2.0
ECU08g0160 30.8 4.2
7304377 30.0 6.0
7296005 30.0 7.2
CE18563 29.6 8.9
> YBR109c
Length=147
Score = 33.5 bits (75), Expect = 0.57, Method: Compositional matrix adjust.
Identities = 16/43 (37%), Positives = 27/43 (62%), Gaps = 3/43 (6%)
Query 199 ELIRCWNDYDK---GLYSTALLRHVFRAMRRIVSDAHVDELLK 238
EL+ + +DK GL S A L+HV ++ ++DA VD++L+
Sbjct 85 ELLEAFKVFDKNGDGLISAAELKHVLTSIGEKLTDAEVDDMLR 127
> Hs4885111
Length=149
Score = 32.7 bits (73), Expect = 0.94, Method: Compositional matrix adjust.
Identities = 15/43 (34%), Positives = 23/43 (53%), Gaps = 0/43 (0%)
Query 197 VSELIRCWNDYDKGLYSTALLRHVFRAMRRIVSDAHVDELLKA 239
+ E R ++ G S A LRHV + +SD VDE+++A
Sbjct 86 IREAFRVFDKDGNGFVSAAELRHVMTRLGEKLSDEEVDEMIRA 128
> At3g56800
Length=149
Score = 32.3 bits (72), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 17/48 (35%), Positives = 26/48 (54%), Gaps = 3/48 (6%)
Query 194 TNAVSELIRCWNDYDK---GLYSTALLRHVFRAMRRIVSDAHVDELLK 238
T++ EL + +DK G S A LRHV + ++D VDE++K
Sbjct 80 TDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIK 127
> At2g41110
Length=149
Score = 32.3 bits (72), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 17/48 (35%), Positives = 26/48 (54%), Gaps = 3/48 (6%)
Query 194 TNAVSELIRCWNDYDK---GLYSTALLRHVFRAMRRIVSDAHVDELLK 238
T++ EL + +DK G S A LRHV + ++D VDE++K
Sbjct 80 TDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIK 127
> At2g27030
Length=149
Score = 32.3 bits (72), Expect = 1.4, Method: Compositional matrix adjust.
Identities = 17/48 (35%), Positives = 26/48 (54%), Gaps = 3/48 (6%)
Query 194 TNAVSELIRCWNDYDK---GLYSTALLRHVFRAMRRIVSDAHVDELLK 238
T++ EL + +DK G S A LRHV + ++D VDE++K
Sbjct 80 TDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIK 127
> YDR127w
Length=1588
Score = 32.0 bits (71), Expect = 1.8, Method: Compositional matrix adjust.
Identities = 17/55 (30%), Positives = 28/55 (50%), Gaps = 4/55 (7%)
Query 82 YGNMALEIFATSPVNYKALQQQDFVKMELSIMKSVKDEEIVLYSVKALAAGCKNP 136
YG A+E+ NY A DFV +LSI++ D ++++V+ + G P
Sbjct 1098 YGCEAVEVRVDHLANYSA----DFVSKQLSILRKATDSIPIIFTVRTMKQGGNFP 1148
> YKL213c
Length=715
Score = 32.0 bits (71), Expect = 2.0, Method: Compositional matrix adjust.
Identities = 36/142 (25%), Positives = 55/142 (38%), Gaps = 17/142 (11%)
Query 181 VERTAVNRNLYNKTNAVSELIRCWNDYDKG---LYSTALLRHVFRAMRRIVSDAHVDE-- 235
+E N+N+ V L+ C+N+ + G L S + + +F + S A +
Sbjct 557 IEEGLGNKNITLTMLTVRILVNCFNNENWGVKLLESNQVYKSIFETIDTEFSQASAKQSQ 616
Query 236 ---LLKANVLQRLIAIVKNLNSDIALLPDVLFLLGSLAVVPEIKTKIGELDGVAACNELL 292
+ + ++ A+V NSD+ LLP V I TK G L+ C E
Sbjct 617 NLAIAVSTLIFNYSALVTKGNSDLELLP---------IVADAINTKYGPLEEYQECEEAA 667
Query 293 QRSLANPQATAVVTNTCLAFAN 314
R A V T FAN
Sbjct 668 YRLTVAYGNLATVEPTLRQFAN 689
> ECU08g0160
Length=200
Score = 30.8 bits (68), Expect = 4.2, Method: Compositional matrix adjust.
Identities = 14/40 (35%), Positives = 23/40 (57%), Gaps = 0/40 (0%)
Query 161 KDTTLSNERKEEHLCSRLLLVERTAVNRNLYNKTNAVSEL 200
K+TT+ +ER+ EHL R ++R + YN+ N + E
Sbjct 18 KNTTVFDEREIEHLYERFQFLDRESRGYLTYNELNNIPEF 57
> 7304377
Length=1465
Score = 30.0 bits (66), Expect = 6.0, Method: Compositional matrix adjust.
Identities = 30/111 (27%), Positives = 49/111 (44%), Gaps = 6/111 (5%)
Query 81 QYGNM----ALEIFATSPVNYKALQQQDFVKMELSIMKSVKDEEIVLYSVKALAAGCKNP 136
Q GN+ ALE SP N K Q VK +S ++ +EE+ + + A +N
Sbjct 984 QSGNVVNHTALEELDQSPQNLKGSHNQKIVKSLVSDIQQNHNEELNSHQHQVKQANKQNL 1043
Query 137 ACAQDAARTVGCPNIVTEAVSRINKDTTLSNERKEEHLCSRLLLVERTAVN 187
Q+AA++ P + R + T ++EE +L +R A+N
Sbjct 1044 NTKQNAAQSALKP--INNENDRKREQTEEKKRQREERKRQQLEDDKRRALN 1092
> 7296005
Length=1638
Score = 30.0 bits (66), Expect = 7.2, Method: Compositional matrix adjust.
Identities = 21/68 (30%), Positives = 34/68 (50%), Gaps = 6/68 (8%)
Query 212 YSTALLRHVFRAMRRIVSDAHVDEL----LKANVLQRLIAIVKNLNSDIALLPDVLFLLG 267
Y T L+ + R MR++V D VD L + Q I+ +L D +LP +L+L
Sbjct 504 YDTVLMYKLIRIMRKLVVDMGVDSLNGPPPNSEAEQHYYDIMSSL--DACILPSLLYLDN 561
Query 268 SLAVVPEI 275
+ ++ EI
Sbjct 562 NCSMSEEI 569
> CE18563
Length=419
Score = 29.6 bits (65), Expect = 8.9, Method: Compositional matrix adjust.
Identities = 21/60 (35%), Positives = 31/60 (51%), Gaps = 6/60 (10%)
Query 227 IVSDAHVDELL-KANVLQRLIAIVKNLNSDIALLPDVLFLLGSLAVVPEIKTKIGELDGV 285
IVSD L+ + N L L + +KN + AL+PD +L ++ I KIG + GV
Sbjct 53 IVSDEERSRLVQRVNQLHSLTSGIKNTSPSCALMPD-----RNLEIIVAIIDKIGSVPGV 107
Lambda K H
0.319 0.133 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 7308161082
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40