bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: kyva
112,920 sequences; 47,500,486 total letters
Query= Eace_0131_orf4
Length=54
Score E
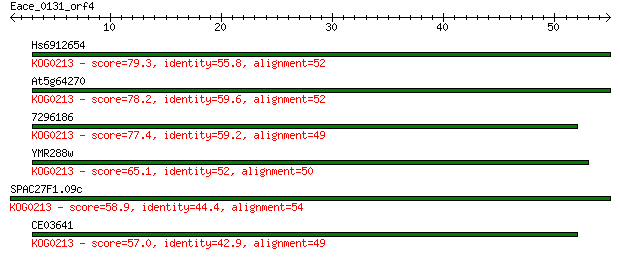
Sequences producing significant alignments: (Bits) Value
Hs6912654 79.3 2e-15
At5g64270 78.2 4e-15
7296186 77.4 5e-15
YMR288w 65.1 3e-11
SPAC27F1.09c 58.9 2e-09
CE03641 57.0 9e-09
> Hs6912654
Length=1304
Score = 79.3 bits (194), Expect = 2e-15, Method: Composition-based stats.
Identities = 29/52 (55%), Positives = 45/52 (86%), Gaps = 0/52 (0%)
Query 3 GLFHPARKVREVYWRVYNTVYIGHQDAMVAFYPKLPDDGKHSFARHELELFI 54
GLFHPARKVR+VYW++YN++YIG QDA++A YP++ +D K+++ R+EL+ +
Sbjct 1253 GLFHPARKVRDVYWKIYNSIYIGSQDALIAHYPRIYNDDKNTYIRYELDYIL 1304
> At5g64270
Length=1269
Score = 78.2 bits (191), Expect = 4e-15, Method: Composition-based stats.
Identities = 31/52 (59%), Positives = 42/52 (80%), Gaps = 0/52 (0%)
Query 3 GLFHPARKVREVYWRVYNTVYIGHQDAMVAFYPKLPDDGKHSFARHELELFI 54
GLFHPARKVREVYW++YN++YIG QD +VA YP L D+ + ++R EL +F+
Sbjct 1218 GLFHPARKVREVYWKIYNSLYIGAQDTLVAAYPVLEDEQNNVYSRPELTMFV 1269
> 7296186
Length=1349
Score = 77.4 bits (189), Expect = 5e-15, Method: Composition-based stats.
Identities = 29/49 (59%), Positives = 43/49 (87%), Gaps = 0/49 (0%)
Query 3 GLFHPARKVREVYWRVYNTVYIGHQDAMVAFYPKLPDDGKHSFARHELE 51
GLFHPARKVR+VYW++YN++YIG QDA++A YP++ +D K+ + R+EL+
Sbjct 1298 GLFHPARKVRDVYWKIYNSLYIGGQDALIAGYPRITNDPKNQYERYELD 1346
> YMR288w
Length=971
Score = 65.1 bits (157), Expect = 3e-11, Method: Composition-based stats.
Identities = 26/50 (52%), Positives = 35/50 (70%), Gaps = 0/50 (0%)
Query 3 GLFHPARKVREVYWRVYNTVYIGHQDAMVAFYPKLPDDGKHSFARHELEL 52
GLFHPA+ VR+ +WRVYN +Y+ +QDAMV FYP PD+ + +L L
Sbjct 922 GLFHPAKNVRKAFWRVYNNMYVMYQDAMVPFYPVTPDNNEEYIEELDLVL 971
> SPAC27F1.09c
Length=1188
Score = 58.9 bits (141), Expect = 2e-09, Method: Composition-based stats.
Identities = 24/54 (44%), Positives = 34/54 (62%), Gaps = 0/54 (0%)
Query 1 ILGLFHPARKVREVYWRVYNTVYIGHQDAMVAFYPKLPDDGKHSFARHELELFI 54
+ GLFHP+RKVR YW YN+ Y+ DAMV +YP + DD +++ L + I
Sbjct 1135 VQGLFHPSRKVRNTYWTSYNSAYVQSADAMVPYYPHVDDDQFNNYDMKTLHICI 1188
> CE03641
Length=1322
Score = 57.0 bits (136), Expect = 9e-09, Method: Composition-based stats.
Identities = 21/49 (42%), Positives = 35/49 (71%), Gaps = 0/49 (0%)
Query 3 GLFHPARKVREVYWRVYNTVYIGHQDAMVAFYPKLPDDGKHSFARHELE 51
L+HPARKVRE W+V+N + +G DA++A YP++ + + + R+EL+
Sbjct 1271 ALWHPARKVREPVWKVFNNLILGSADALIAAYPRIENTPTNQYVRYELD 1319
Lambda K H
0.329 0.146 0.476
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 1200194442
Database: kyva
Posted date: Jul 3, 2009 9:03 AM
Number of letters in database: 47,500,486
Number of sequences in database: 112,920
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40