bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: eggV2
2,483,276 sequences; 915,453,621 total letters
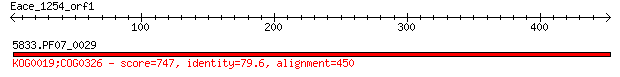
Query= Eace_1254_orf1
Length=452
Score E
Sequences producing significant alignments: (Bits) Value
5833.PF07_0029 747 0.0
> 5833.PF07_0029
Length=745
Score = 747 bits (1929), Expect = 0.0, Method: Compositional matrix adjust.
Identities = 358/450 (79%), Positives = 400/450 (88%), Gaps = 3/450 (0%)
Query 3 VKEVTREWEQLNKQKPLWMRKPEEVTEEEYASFYKSLSNDWEEHLAVKHFSVEGQLEFKA 62
+ V EWE+LNKQKPLWMRKPEEVT EEYASFYKSL+NDWE+HLAVKHFSVEGQLEFKA
Sbjct 299 IHTVEHEWEELNKQKPLWMRKPEEVTNEEYASFYKSLTNDWEDHLAVKHFSVEGQLEFKA 358
Query 63 LLFVPKRAPFDLFETRKKRNNIKLYVRRVFIMDDCEDIIPEWLNFVKGVVDSEDLPLNIS 122
LLF+PKRAPFD+FE RKKRNNIKLYVRRVFIMDDCE+IIPEWLNFVKGVVDSEDLPLNIS
Sbjct 359 LLFIPKRAPFDMFENRKKRNNIKLYVRRVFIMDDCEEIIPEWLNFVKGVVDSEDLPLNIS 418
Query 123 RESLQQNKILKVIRKNLVKKCLEMFAEIEEKKENYTKFYEQFSKNLKLGIHEDSANRAKI 182
RESLQQNKILKVI+KNL+KKCL+MF+E+ E KENY KFYEQFSKNLKLGIHED+ANR KI
Sbjct 419 RESLQQNKILKVIKKNLIKKCLDMFSELAENKENYKKFYEQFSKNLKLGIHEDNANRTKI 478
Query 183 AELLRFHSSKSGEDMVSFKEYVDRMKEGQKDIYYITGESRQTVANSPFLEKLTKKGYEVL 242
ELLRF +SKSG++M+ KEYVDRMKE QKDIYYITGES V+NSPFLE LTKKG+EV+
Sbjct 479 TELLRFQTSKSGDEMIGLKEYVDRMKENQKDIYYITGESINAVSNSPFLEALTKKGFEVI 538
Query 243 YMTDPIDEYAVQQLKEFDNHKLRCCTKEGLEIDETEEEKKKFEELKAEFEPLLKLIKEVL 302
YM DPIDEYAVQQLK+FD KL+CCTKEGL+ID++EE KK FE LKAE+E L K+IK+VL
Sbjct 539 YMVDPIDEYAVQQLKDFDGKKLKCCTKEGLDIDDSEEAKKDFETLKAEYEGLCKVIKDVL 598
Query 303 HDKVDKVVLSNRITDSPCVLVTTEFGWSANMERIMKAQALRDNSMTSYMVSKKTMEVNGH 362
H+KV+KVV+ RITDSPCVLVT+EFGWSANMERIMKAQALRDNSMTSYM+SKK ME+N
Sbjct 599 HEKVEKVVVGQRITDSPCVLVTSEFGWSANMERIMKAQALRDNSMTSYMLSKKIMEINAR 658
Query 363 HSIMVEIKNKAAVDKSDKTVKDLIWLLYDTALLTSGFSLEEPTQFAARIHRMIKLGLSID 422
H I+ +K KA DKSDKTVKDLIWLL+DT+LLTSGF+LEEPT F+ RIHRMIKLGLSI
Sbjct 659 HPIISALKQKADADKSDKTVKDLIWLLFDTSLLTSGFALEEPTTFSKRIHRMIKLGLSI- 717
Query 423 DDEEAKEDDLPPLEEVEGAADEASKMEEVD 452
D+EE + DLPPLEE A D SKMEEVD
Sbjct 718 DEEENNDIDLPPLEETVDATD--SKMEEVD 745
Lambda K H
0.315 0.132 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 188051580480
Database: eggV2
Posted date: Dec 15, 2009 4:47 PM
Number of letters in database: 915,453,621
Number of sequences in database: 2,483,276
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40