bitscore colors: <40, 40-50 , 50-80, 80-200, >200
BLASTP 2.2.24+
Reference: Stephen F. Altschul, Thomas L. Madden, Alejandro A.
Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J.
Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of
protein database search programs", Nucleic Acids Res. 25:3389-3402.
Reference for composition-based statistics: Alejandro A. Schaffer,
L. Aravind, Thomas L. Madden, Sergei Shavirin, John L. Spouge, Yuri
I. Wolf, Eugene V. Koonin, and Stephen F. Altschul (2001),
"Improving the accuracy of PSI-BLAST protein database searches with
composition-based statistics and other refinements", Nucleic Acids
Res. 29:2994-3005.
Database: eggV2
2,483,276 sequences; 915,453,621 total letters
Query= Eace_0335_orf1
Length=520
Score E
Sequences producing significant alignments: (Bits) Value
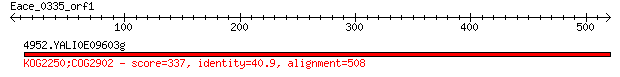
4952.YALI0E09603g 337 7e-91
> 4952.YALI0E09603g
Length=987
Score = 337 bits (864), Expect = 7e-91, Method: Compositional matrix adjust.
Identities = 208/545 (38%), Positives = 302/545 (55%), Gaps = 60/545 (11%)
Query 13 TEEFFSRFGDCLTMWGFYSQRKYVEPLANGSSIITTF-VELLPE---------GILVFPS 62
T +FS D +G S RKYVE +NG +II+ + V P+ + +P
Sbjct 206 TPGYFSALSDLYHFYGLTSTRKYVEQFSNGVTIISMYLVPAFPQTANTADEIKAMKRYP- 264
Query 63 MPIQERIDNLLSAVRLQFVMAKSKYAD-FARDRLLTLHEAAYAMNASHFALHFSGSVGHS 121
PI+ I ++ L F + + + FAR + +L E+ YA A F HF +G
Sbjct 265 -PIENSIHQIVKEASLLFCLPNNAFKHHFARGDM-SLQESIYAHCAFIFVQHFLNRLGSE 322
Query 122 FPRIEEIVRQFGGKSLTMQEIYE-LRTRLKTCPFSEDLIFKVVGENANIVKELYKEFASL 180
+ +++++ G EI E L+ RL+ F+ D +F+++ ++VK++Y +FA +
Sbjct 323 YTTLQQML----GTGKEHVEILEKLKRRLRQETFTRDYLFELINNQLDVVKQMYLQFADV 378
Query 181 HC----------------PRVKRVLFKETNELKERILRLESSEAVAILLK-FREFTKSIL 223
H R++ T+ELK+ I + +++ A++++ F F +L
Sbjct 379 HYIQSKSEGDSFLPTLSYQRLQTQSVLSTDELKKLIRKKAANDHEAMVMEAFLTFNTHVL 438
Query 224 CTNFYMPFKQGFAFRIRGSTLPSSDFPSTPCPIFFQIGGLAVGLHIRFAEVSRGGVRLVF 283
TNFY P K +FR+ LP S++P +F +G G H+RFA+++RGG+R+V
Sbjct 439 KTNFYTPTKVALSFRLSPDFLPESEYPQPLYGMFLVVGQEFRGFHLRFADIARGGIRIVK 498
Query 284 SVGTAAHETNRRSLLDEAYKLAFTQQFKNKDISEGGSKGIILLNKTQTLAEAKRQAPLAF 343
S A+ N RS+ DE Y LA TQQ KNKDI EGGSKG+ILLN E + +A +AF
Sbjct 499 SRNREAYSINARSMFDENYNLANTQQRKNKDIPEGGSKGVILLNN-----EHQDKAEIAF 553
Query 344 KAYIDNMLDLLLP------HHDVDDGLGISEVWFLGPDENTGTGGLLDWAAQRAKERGSV 397
YID+++DLLL + D G E+ F+GPDEN T GL++WA AK+RG+
Sbjct 554 HKYIDSVIDLLLKGDTPGIKEPIVDLHGSPEILFMGPDEN--TAGLVNWATMHAKQRGAP 611
Query 398 WWKAFTTGKLIQHGGIPHDRFGMTTASVEAYVKDAGIYNKLGLKEEEMTRIQTGGPDGDL 457
WWK+F TGK GGIPHD +GMT+ SV YVK GIY KL +++ + R QTGGPDGDL
Sbjct 612 WWKSFFTGKSPSLGGIPHDEYGMTSLSVREYVK--GIYRKLEIEQPTVRRQQTGGPDGDL 669
Query 458 GCNALLQTKSKTIAVVDGSGVLYDPNGLDVGELHRLCSLRFEGKPTNAML--YDSSLLSP 515
G N +L + K V+DG+GVLYDPNGLD EL L R M+ YD+S LSP
Sbjct 670 GSNEILLSAEKYTTVIDGAGVLYDPNGLDREELLSLAKRR-------VMISEYDASKLSP 722
Query 516 LGFKV 520
G++V
Sbjct 723 EGYRV 727
Lambda K H
0.322 0.139 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Effective search space used: 226463852331
Database: eggV2
Posted date: Dec 15, 2009 4:47 PM
Number of letters in database: 915,453,621
Number of sequences in database: 2,483,276
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Neighboring words threshold: 11
Window for multiple hits: 40